Sec20 Is Required for Autophagic and Endocytic Degradation Independent of Golgi-ER Retrograde Transport
Abstract
:1. Introduction
2. Materials and Methods
2.1. Fly Stocks
2.2. Larval Stages
2.3. Immunohistochemistry
2.4. LysoTracker Staining, Uptake Assays and Salivary Gland Dissection
2.5. Fluorescent Imaging
2.6. Electron Microscopic Analysis
2.7. Statistics
3. Results
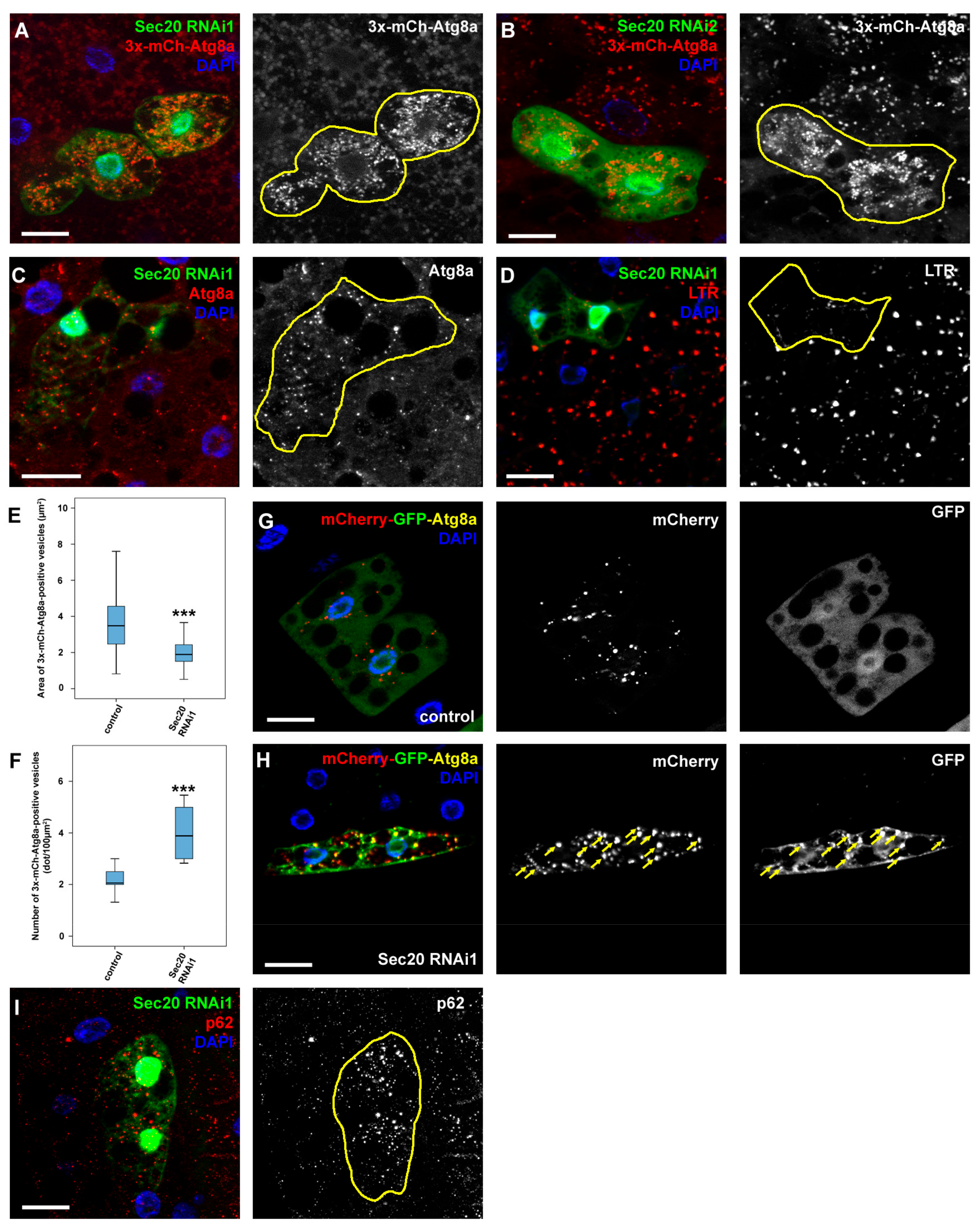
3.1. Sec20 Depletion Causes Defects during Endocytosis and Autophagy
3.2. Ultrastructural Analysis Reveals Non-Functional Lysosomes in Fat Cells and Nephrocytes
3.3. Sec20 Promotes Lysosomal Biogenesis and Degradation
3.4. Sec20 Acts Independently of Sec22 and Use1 in Lysosomal Regulation but It May Cooperate with Syx18
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Mukherjee, S.; Ghosh, R.N.; Maxfield, F.R. Endocytosis. Physiol. Rev. 1997, 77, 759–803. [Google Scholar] [CrossRef] [PubMed]
- Lamb, C.A.; Dooley, H.C.; Tooze, S.A. Endocytosis and autophagy: Shared machinery for degradation. Bioessays 2013, 35, 34–45. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Vicente, M.; Cuervo, A.M. Autophagy and neurodegeneration: When the cleaning crew goes on strike. Lancet Neurol. 2007, 6, 352–361. [Google Scholar] [CrossRef]
- Kundu, M.; Thompson, C.B. Autophagy: Basic principles and relevance to disease. Annu. Rev. Pathmechdis. Mech. Dis. 2008, 3, 427–455. [Google Scholar] [CrossRef] [PubMed]
- Doherty, G.J.; McMahon, H.T. Mechanisms of endocytosis. Annu. Rev. Biochem. 2009, 78, 857–902. [Google Scholar] [CrossRef] [PubMed]
- Wickner, W.; Schekman, R. Membrane fusion. Nat. Struct. Mol. Biol. 2008, 15, 658. [Google Scholar] [CrossRef] [PubMed]
- Kloepper, T.H.; Kienle, C.N.; Fasshauer, D. An elaborate classification of SNARE proteins sheds light on the conservation of the eukaryotic endomembrane system. Mol. Biol. Cell 2007, 18, 3463–3471. [Google Scholar] [CrossRef]
- Subramanian, T.; Chinnadurai, G. Interaction of cellular apoptosis regulating proteins with adenovirus anti-apoptosis protein E1B-19K. In Two-Hybrid Systems; Springer: Berlin, Germany, 2001; pp. 211–218. [Google Scholar]
- Novick, P.; Field, C.; Schekman, R. Identification of 23 complementation groups required for post-translational events in the yeast secretory pathway. Cell 1980, 21, 205–215. [Google Scholar] [CrossRef]
- Meiringer, C.T.; Rethmeier, R.; Auffarth, K.; Wilson, J.; Perz, A.; Barlowe, C.; Schmitt, H.D.; Ungermann, C. The Dsl1 protein tethering complex is a resident endoplasmic reticulum complex, which interacts with five soluble NSF (N-ethylmaleimide-sensitive factor) attachment protein receptors (SNAREs): Implications for fusion and fusion regulation. J. Biol. Chem. 2011, 286, 25039–25046. [Google Scholar] [CrossRef]
- Tripathi, A.; Ren, Y.; Jeffrey, P.D.; Hughson, F.M. Structural characterization of Tip20p and Dsl1p, subunits of the Dsl1p vesicle tethering complex. Nat. Struct. Mol. Biol. 2009, 16, 114. [Google Scholar] [CrossRef]
- Reilly, B.A.; Kraynack, B.A.; VanRheenen, S.M.; Waters, M.G. Golgi-to-endoplasmic reticulum (ER) retrograde traffic in yeast requires Dsl1p, a component of the ER target site that interacts with a COPI coat subunit. Mol. Biol. Cell 2001, 12, 3783–3796. [Google Scholar] [CrossRef] [PubMed]
- Nakajima, K.I.; Hirose, H.; Taniguchi, M.; Kurashina, H.; Arasaki, K.; Nagahama, M.; Tani, K.; Yamamoto, A.; Tagaya, M. Involvement of BNIP1 in apoptosis and endoplasmic reticulum membrane fusion. EMBO J. 2004, 23, 3216–3226. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yasuda, M.; Chinnadurai, G. Functional identification of the apoptosis effector BH3 domain in cellular protein BNIP1. Oncogene 2000, 19, 2363. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Cheung, P.; Yanagawa, B.; McManus, B.; Yang, D. BNips: A group of pro-apoptotic proteins in the Bcl-2 family. Apoptosis 2003, 8, 229–236. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Heim, J.; Meyhack, B. Novel BNIP1 variants and their interaction with BCL2 family members. FEBS Lett. 1999, 448, 23–27. [Google Scholar] [CrossRef]
- Wang, P.; Wu, Y.; Li, Y.; Zheng, J.; Tang, J. A novel RING finger E3 ligase RNF186 regulate ER stress-mediated apoptosis through interaction with BNip1. Cell. Signal. 2013, 25, 2320–2333. [Google Scholar] [CrossRef] [PubMed]
- Nishiwaki, Y.; Yoshizawa, A.; Kojima, Y.; Oguri, E.; Nakamura, S.; Suzuki, S.; Yuasa-Kawada, J.; Kinoshita-Kawada, M.; Mochizuki, T.; Masai, I. The BH3-only SNARE BNip1 mediates photoreceptor apoptosis in response to vesicular fusion defects. Dev. Cell 2013, 25, 374–387. [Google Scholar] [CrossRef]
- Tang, F.; Wang, B.; Li, N.; Wu, Y.; Jia, J.; Suo, T.; Chen, Q.; Liu, Y.-J.; Tang, J. RNF185, a novel mitochondrial ubiquitin E3 ligase, regulates autophagy through interaction with BNIP1. PLoS ONE 2011, 6, e24367. [Google Scholar] [CrossRef]
- Ryu, S.W.; Choi, K.; Yoon, J.; Kim, S.; Choi, C. Endoplasmic reticulum-specific BH3-only protein BNIP1 induces mitochondrial fragmentation in a Bcl-2-and Drp1-dependent manner. J. Cell. Physiol. 2012, 227, 3027–3035. [Google Scholar] [CrossRef]
- Cho, B.; Choi, S.Y.; Park, O.-H.; Sun, W.; Geum, D. Differential expression of BNIP family members of BH3-only proteins during the development and after axotomy in the rat. Mol. Cells 2012, 33, 605–610. [Google Scholar] [CrossRef]
- Korolchuk, V.I.; Saiki, S.; Lichtenberg, M.; Siddiqi, F.H.; Roberts, E.A.; Imarisio, S.; Jahreiss, L.; Sarkar, S.; Futter, M.; Menzies, F.M. Lysosomal positioning coordinates cellular nutrient responses. Nat. Cell Biol. 2011, 13, 453. [Google Scholar] [CrossRef] [PubMed]
- Hegedűs, K.; Takáts, S.; Boda, A.; Jipa, A.; Nagy, P.; Varga, K.; Kovács, A.L.; Juhász, G. The Ccz1-Mon1-Rab7 module and Rab5 control distinct steps of autophagy. Mol. Biol. Cell 2016, 27, 3132–3142. [Google Scholar] [CrossRef] [Green Version]
- Pulipparacharuvil, S.; Akbar, M.A.; Ray, S.; Sevrioukov, E.A.; Haberman, A.S.; Rohrer, J.; Krämer, H. Drosophila Vps16A is required for trafficking to lysosomes and biogenesis of pigment granules. J. Cell Sci 2005, 118, 3663–3673. [Google Scholar] [CrossRef] [PubMed]
- Takáts, S.; Pircs, K.; Nagy, P.; Varga, Á.; Kárpáti, M.; Hegedűs, K.; Kramer, H.; Kovács, A.L.; Sass, M.; Juhász, G. Interaction of the HOPS complex with Syntaxin 17 mediates autophagosome clearance in Drosophila. Mol. Biol. Cell 2014, 25, 1338–1354. [Google Scholar] [CrossRef] [PubMed]
- Lorincz, P.; Kenez, L.A.; Toth, S.; Kiss, V.; Varga, A.; Csizmadia, T.; Simon-Vecsei, Z.; Juhasz, G. Vps8 overexpression inhibits HOPS-dependent trafficking routes by outcompeting Vps41/Lt. Cell Biol. 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Mauvezin, C.; Ayala, C.; Braden, C.R.; Kim, J.; Neufeld, T.P. Assays to monitor autophagy in Drosophila. Methods 2014, 68, 134–139. [Google Scholar] [CrossRef] [Green Version]
- Nagy, P.; Varga, Á.; Kovács, A.L.; Takáts, S.; Juhász, G. How and why to study autophagy in Drosophila: It’s more than just a garbage chute. Methods 2015, 75, 151–161. [Google Scholar] [CrossRef]
- Biyasheva, A.; Do, T.V.; Lu, Y.; Vaskova, M.; Andres, A.J. Glue secretion in the Drosophila salivary gland: A model for steroid-regulated exocytosis. Dev. Biol. 2001, 231, 234–251. [Google Scholar] [CrossRef]
- Csizmadia, T.; Lőrincz, P.; Hegedűs, K.; Széplaki, S.; Lőw, P.; Juhász, G. Molecular mechanisms of developmentally programmed crinophagy in Drosophila. J. Cell Biol. 2018, 217, 361–374. [Google Scholar] [CrossRef]
- Costantino, B.F.; Bricker, D.K.; Alexandre, K.; Shen, K.; Merriam, J.R.; Antoniewski, C.; Callender, J.L.; Henrich, V.C.; Presente, A.; Andres, A.J. A novel ecdysone receptor mediates steroid-regulated developmental events during the mid-third instar of Drosophila. PLoS Genet. 2008, 4, e1000102. [Google Scholar] [CrossRef]
- Riedel, F.; Gillingham, A.K.; Rosa-Ferreira, C.; Galindo, A.; Munro, S. An antibody toolkit for the study of membrane traffic in Drosophila melanogaster. Biol. Open 2016, 5, 987–992. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, T.; Nakamura, A. The endocytic pathway acts downstream of Oskar in Drosophila germ plasm assembly. Development 2008, 135, 1107–1117. [Google Scholar] [CrossRef] [PubMed]
- Takats, S.; Nagy, P.; Varga, A.; Pircs, K.; Karpati, M.; Varga, K.; Kovacs, A.L.; Hegedus, K.; Juhasz, G. Autophagosomal Syntaxin17-dependent lysosomal degradation maintains neuronal function in Drosophila. J. Cell Biol. 2013, 201, 531–539. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pircs, K.; Nagy, P.; Varga, A.; Venkei, Z.; Erdi, B.; Hegedus, K.; Juhasz, G. Advantages and limitations of different p62-based assays for estimating autophagic activity in Drosophila. PLoS ONE 2012, 7, e44214. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.-Y.; Neufeld, T.P. Autophagy takes flight in Drosophila. FEBS Lett. 2010, 584, 1342–1349. [Google Scholar] [CrossRef]
- Na, J.; Cagan, R. The Drosophila nephrocyte: Back on stage. Am. Soc. Nephrol. 2013, 24, 161–163. [Google Scholar] [CrossRef] [PubMed]
- Weavers, H.; Prieto-Sánchez, S.; Grawe, F.; Garcia-López, A.; Artero, R.; Wilsch-Bräuninger, M.; Ruiz-Gómez, M.; Skaer, H.; Denholm, B. The insect nephrocyte is a podocyte-like cell with a filtration slit diaphragm. Nature 2009, 457, 322. [Google Scholar] [CrossRef] [PubMed]
- Lőrincz, P.; Lakatos, Z.; Varga, Á.; Maruzs, T.; Simon-Vecsei, Z.; Darula, Z.; Benkő, P.; Csordás, G.; Lippai, M.; Ando, I.; et al. MiniCORVET is a Vps8-containing early endosomal tether in Drosophila. eLife 2016, 5, e14226. [Google Scholar] [CrossRef]
- Jiang, P.; Nishimura, T.; Sakamaki, Y.; Itakura, E.; Hatta, T.; Natsume, T.; Mizushima, N. The HOPS complex mediates autophagosome–lysosome fusion through interaction with syntaxin 17. Mol. Biol. Cell 2014, 25, 1327–1337. [Google Scholar] [CrossRef]
- Lőrincz, P.; Tóth, S.; Benkő, P.; Lakatos, Z.; Boda, A.; Glatz, G.; Zobel, M.; Bisi, S.; Hegedűs, K.; Takáts, S.; et al. Rab2 promotes autophagic and endocytic lysosomal degradation. J. Cell Biol. 2017, 216, 1937–1947. [Google Scholar] [CrossRef] [Green Version]
- Marwaha, R.; Sharma, M. DQ-Red BSA Trafficking Assay in Cultured Cells to Assess Cargo Delivery to Lysosomes. Bio. Protoc. 2017, 7. [Google Scholar] [CrossRef] [PubMed]
- Lőrincz, P.; Mauvezin, C.; Juhász, G. Exploring autophagy in Drosophila. Cells 2017, 6, 22. [Google Scholar] [CrossRef] [PubMed]
- Nezis, I.P.; Simonsen, A.; Sagona, A.P.; Finley, K.; Gaumer, S.; Contamine, D.; Rusten, T.E.; Stenmark, H.; Brech, A. Ref (2) P, the Drosophila melanogaster homologue of mammalian p62, is required for the formation of protein aggregates in adult brain. J. Cell Biol. 2008, 180, 1065–1071. [Google Scholar] [CrossRef] [PubMed]
- Maruzs, T.; Lorincz, P.; Szatmari, Z.; Szeplaki, S.; Sandor, Z.; Lakatos, Z.; Puska, G.; Juhasz, G.; Sass, M. Retromer Ensures the Degradation of Autophagic Cargo by Maintaining Lysosome Function in Drosophila. Traffic 2015, 16, 1088–1107. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mauvezin, C.; Nagy, P.; Juhasz, G.; Neufeld, T.P. Autophagosome-lysosome fusion is independent of V-ATPase-mediated acidification. Nat. Commun. 2015, 6, 7007. [Google Scholar] [CrossRef] [PubMed]
- Lloyd, V.; Ramaswami, M.; Krämer, H. Not just pretty eyes: Drosophila eye-colour mutations and lysosomal delivery. Trends Cell Biol. 1998, 8, 257–259. [Google Scholar] [CrossRef]
- Chyb, S.; Gompel, N. Atlas of Drosophila Morphology: Wild-Type and Classical Mutants; Academic Press: Elsevier: Amsterdam, The Netherlands, 2013. [Google Scholar]
- Warner, T.; Sinclair, D.; Fitzpatrick, K.; Singh, M.; Devlin, R.; Honda, B. The light gene of Drosophila melanogaster encodes a homologue of VPS41, a yeast gene involved in cellular-protein trafficking. Genome 1998, 41, 236–243. [Google Scholar] [CrossRef] [PubMed]
- Mathew, R.; Karantza-Wadsworth, V.; White, E. Role of autophagy in cancer. Nat. Rev. Cancer 2007, 7, 961–967. [Google Scholar] [CrossRef]
- Sriburi, R.; Jackowski, S.; Mori, K.; Brewer, J.W. XBP1: A link between the unfolded protein response, lipid biosynthesis, and biogenesis of the endoplasmic reticulum. J. Cell Biol. 2004, 167, 35–41. [Google Scholar] [CrossRef]
- Sriram, V.; Krishnan, K.S.; Mayor, S. Deep-orange and carnation define distinct stages in late endosomal biogenesis in Drosophila melanogaster. J. Cell. Biol. 2003, 161, 593–607. [Google Scholar] [CrossRef]
- Swetha, M.G.; Sriram, V.; Krishnan, K.S.; Oorschot, V.M.; ten Brink, C.; Klumperman, J.; Mayor, S. Lysosomal membrane protein composition, acidic pH and sterol content are regulated via a light-dependent pathway in metazoan cells. Traffic 2011, 12, 1037–1055. [Google Scholar] [CrossRef] [PubMed]
- Lund, V.K.; Madsen, K.L.; Kjaerulff, O. Drosophila Rab2 controls endosome-lysosome fusion and LAMP delivery to late endosomes. 2018, 14, 1520–1542. Autophagy 2018, 14, 1520–1542. [Google Scholar] [CrossRef] [PubMed]
- Ayala, C.I.; Kim, J.; Neufeld, T.P. Rab6 promotes insulin receptor and cathepsin trafficking to regulate autophagy induction and activity in Drosophila. J. Cell Sci. 2018, 131. [Google Scholar] [CrossRef] [PubMed]
- Koike, M.; Nakanishi, H.; Saftig, P.; Ezaki, J.; Isahara, K.; Ohsawa, Y.; Schulz-Schaeffer, W.; Watanabe, T.; Waguri, S.; Kametaka, S.; et al. Cathepsin D deficiency induces lysosomal storage with ceroid lipofuscin in mouse CNS neurons. J. Neurosci. 2000, 20, 6898–6906. [Google Scholar] [CrossRef] [PubMed]
- Arasaki, K.; Taniguchi, M.; Tani, K.; Tagaya, M. RINT-1 regulates the localization and entry of ZW10 to the syntaxin 18 complex. Mol. Biol. Cell 2006, 17, 2780–2788. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Yang, H.; Liu, W.; Duan, X.; Shang, W.; Xia, D.; Tong, C. Sec22 regulates endoplasmic reticulum morphology but not autophagy and is required for eye development in Drosophila. J. Biol. Chem. 2015, 290, 7943–7951. [Google Scholar] [CrossRef] [PubMed]





© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lakatos, Z.; Lőrincz, P.; Szabó, Z.; Benkő, P.; Kenéz, L.A.; Csizmadia, T.; Juhász, G. Sec20 Is Required for Autophagic and Endocytic Degradation Independent of Golgi-ER Retrograde Transport. Cells 2019, 8, 768. https://doi.org/10.3390/cells8080768
Lakatos Z, Lőrincz P, Szabó Z, Benkő P, Kenéz LA, Csizmadia T, Juhász G. Sec20 Is Required for Autophagic and Endocytic Degradation Independent of Golgi-ER Retrograde Transport. Cells. 2019; 8(8):768. https://doi.org/10.3390/cells8080768
Chicago/Turabian StyleLakatos, Zsolt, Péter Lőrincz, Zoltán Szabó, Péter Benkő, Lili Anna Kenéz, Tamás Csizmadia, and Gábor Juhász. 2019. "Sec20 Is Required for Autophagic and Endocytic Degradation Independent of Golgi-ER Retrograde Transport" Cells 8, no. 8: 768. https://doi.org/10.3390/cells8080768




