Assessment of Genetic Diversity in Differently Colored Raspberry Cultivars Using SSR Markers Located in Flavonoid Biosynthesis Genes
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. Simple Sequence Repeat (SSR) Marker and Polymerase Chain Reaction (PCR) Primer Development
2.3. DNA Isolation, PCR Amplification and Fragment Analysis
2.4. Genetic Data Analysis
3. Results
3.1. Polymorphism and Genetic Diversity Analysis
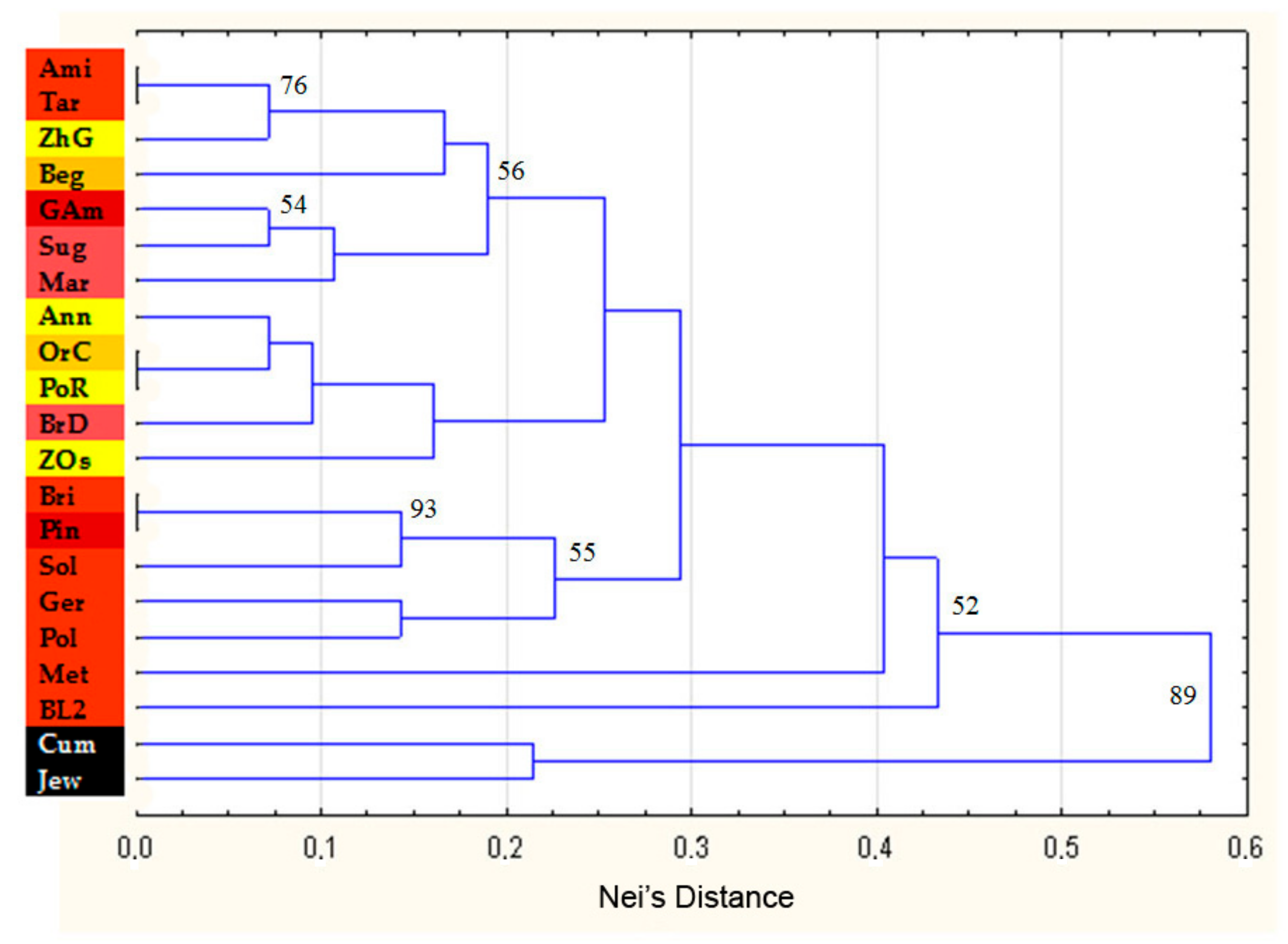
3.2. Cluster Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Thompson, M.M. Chromosome numbers of Rubus species at the National Clonal Germplasm Repository. HortScience 1995, 30, 1447–1452. [Google Scholar] [CrossRef]
- Skrovankova, S.; Sumczynski, D.; Mlcek, J.; Jurikova, T.; Sochor, J. Bioactive compounds and antioxidant activity in different types of berries. Int. J. Mol. Sci. 2015, 16, 24673–24706. [Google Scholar] [CrossRef]
- Halvorsen, B.L.; Myhrstad, M.C.W.; Wold, A.B.; Jacobs, D.R.; Haffner, K.; Holte, K.; Andersen, L.F.; Baugerod, H.; Barikmo, I.; Hvattum, E.; et al. A systematic screening of total antioxidants in dietary plants. J. Nutr. 2002, 132, 461–471. [Google Scholar] [CrossRef]
- Olas, B. Berry phenolic antioxidants—Implications for human health? Front. Pharmacol. 2018, 9, 78. [Google Scholar] [CrossRef]
- FAOSTAT. 2019. Available online: http://www.fao.org/faostat/ (accessed on 29 July 2019).
- Graham, J.; Jennings, S.N. Raspberry breeding. In Breeding Tree Crops; Jain, S.M., Priyadarshan, M., Eds.; IBH & Science Publication: Oxford, UK, 2009; pp. 233–248. [Google Scholar]
- Bushakra, J.M.; Lewers, K.S.; Staton, M.E.; Zhebentyayeva, T.; Saski, C.A. Developing expressed sequence tag libraries and the discovery of simple sequence repeat markers for two species of raspberry (Rubus L.). BMC Plant Biol. 2015, 15, 258. [Google Scholar] [CrossRef]
- Castillo, N.R.F.; Reed, B.M.; Graham, J.; Fernandez-Fernandez, F.; Bassil, N.V. Microsatellite markers for raspberry and blackberry. J. Am. Soc. Hortic. Sci. 2010, 135, 271–278. [Google Scholar] [CrossRef]
- Graham, J.; Smith, K.; MacKenzie, K.; Jorgenson, L.; Hackett, C.; Powell, W. The construction of a genetic linkage map of red raspberry (Rubus idaeus subsp. idaeus) based on AFLPs, genomic-SSR and EST-SSR markers. Theor. Appl. Genet. 2004, 109, 740–749. [Google Scholar] [CrossRef]
- Sargent, D.J.; Fernández-Fernández, F.; Rys, A.; Knight, V.H.; Simpson, D.W.; Tobutt, K.R. Mapping of A1 conferring resistance to the aphid Amphorophora idaei and dw (dwarfing habit) in red raspberry (Rubus idaeus L.) using AFLP and microsatellite markers. BMC Plant Biol. 2007, 7, 15. [Google Scholar] [CrossRef]
- Pattison, J.A.; Samuelian, S.K.; Weber, C.A. Inheritance of Phytophthora root rot resistance in red raspberry determined by generation means and molecular linkage analysis. Theor. Appl. Genet. 2007, 115, 225–236. [Google Scholar] [CrossRef]
- Woodhead, M.; McCallum, S.; Smith, K.; Cardle, L.; Mazzitelli, L.; Graham, J. Identification, characterisation and mapping of simple sequence repeat (SSR) markers from raspberry root and bud ESTs. Mol. Breed. 2008, 22, 555–563. [Google Scholar] [CrossRef]
- Bushakra, J.M.; Stephens, M.J.; Atmadjaja, A.N.; Lewers, K.S.; Symonds, V.V.; Udall, J.A.; Chagne, D.; Buck, E.J.; Gardiner, S.E. Construction of black (Rubus occidentalis) and red (R. idaeus) raspberry linkage maps and their comparison to the genomes of strawberry, apple, and peach. Theor. Appl. Genet. 2012, 125, 311–327. [Google Scholar] [CrossRef]
- Castro, P.; Stafne, E.T.; Clark, J.R.; Lewers, K.S. Genetic map of the primocane-fruiting and thornless traits of tetraploid blackberry. Theor. Appl. Genet. 2013, 126, 2521–2532. [Google Scholar] [CrossRef]
- Graham, J.; Smith, K.; Tierney, I.; MacKenzie, K.; Hackett, C.A. Mapping gene H controlling cane pubescence in raspberry and its association with resistance to cane botrytis and spur blight, rust and cane spot. Theor. Appl. Genet. 2006, 112, 818–831. [Google Scholar] [CrossRef]
- Kassim, A.; Poette, J.; Paterson, A.; Zait, D.; McCallum, S.; Woodhead, M.; Smith, K.; Hackett, C.; Graham, J. Environmental and seasonal influences on red raspberry anthocyanin antioxidant contents and identification of quantitative traits loci (QTL). Mol. Nutr. Food Res. 2009, 53, 625–634. [Google Scholar] [CrossRef]
- Graham, J.; Hackett, C.A.; Smith, K.; Woodhead, M.; Hein, I.; McCallum, S. Mapping QTL for developmental traits in raspberry from bud break to ripe fruit. Theor. Appl. Genet. 2009, 118, 1143–1155. [Google Scholar] [CrossRef]
- McCallum, S.; Smith, K.; Woodhead, M.; Hackett, C.; Paterson, A.; Graham, J. Developing molecular markers for quality traits in red raspberry. Theor. Appl. Genet. 2010, 121, 611–627. [Google Scholar] [CrossRef]
- Jennings, S.N.; Graham, J.; Ferguson, L.; Young, V. New developments in raspberry breeding in Scotland. Acta Hortic. 2016, 1133, 23–28. [Google Scholar] [CrossRef]
- Bushakra, J.M.; Bryant, D.B.; Dossett, M.; Vining, K.J.; VanBuren, R.; Gilmore, B.S.; Lee, J.; Mockler, T.C.; Finn, C.E.; Bassil, N.V. A genetic linkage map of black raspberry (Rubus occidentalis) and the mapping of Ag4 conferring resistance to the aphid Amphorophora agathonica. Theor. Appl. Genet. 2015, 128, 1631–1646. [Google Scholar] [CrossRef]
- Powell, W.; Machray, G.C.; Provan, J. Polymorphism revealed by simple sequence repeats. Trends Plant Sci. 1996, 1, 215–222. [Google Scholar] [CrossRef]
- Graham, J.; Smith, K.; Woodhead, M.; Russell, J. Development and use of simple sequence repeat SSR markers in Rubus species. Mol. Ecol. Notes 2002, 2, 250–252. [Google Scholar] [CrossRef]
- Gezan, S.; Osorio, L.; Verma, S.; Whitaker, V. An experimental validation of genomic selection in octoploid strawberry. Hortic. Res. 2017, 4, 16070. [Google Scholar] [CrossRef] [Green Version]
- Bobinaite, R.; Viskelis, P.; Venskutonis, P.R. Variation of total phenolics, anthocyanins, ellagic acid and radical scavenging capacity in various raspberry (Rubus spp.) cultivars. Food Chem. 2012, 132, 1495–1501. [Google Scholar] [CrossRef]
- Lee, S.S.; Lee, E.M.; An, B.C.; Barampuram, S.; Kim, J.-S.; Cho, J.Y.; Lee, I.-C.; Chung, B.Y. Molecular cloning and characterization of a flavanone-3-hydroxylase gene from Rubus occidentalis L. J. Radiat. Ind. 2008, 2, 121–128. [Google Scholar]
- Chen, Q.; Yu, H.W.; Wang, X.R.; Xie, X.L.; Yue, X.Y.; Tang, H.R. An alternative cetyltrimethylammonium bromide-based protocol for RNA isolation from blackberry (Rubus L.). Genet. Mol. Res. 2012, 11, 1773–1782. [Google Scholar] [CrossRef]
- Li, S. Transcriptional control of flavonoid biosynthesis: Fine-tuning of the MYB-bHLH-WD40 (MBW) complex. Plant Signal. Behav. 2014, 9, e27522. [Google Scholar] [CrossRef]
- Dossett, M.; Bassil, N.V.; Lewers, K.S.; Finn, C.E. Genetic diversity in wild and cultivated black raspberry (Rubus occidentalis L.) evaluated by simple sequence repeat markers. Genet. Resour. Crop Evol. 2012, 59, 1849–1865. [Google Scholar] [CrossRef]
- Lee, G.-A.; Song, J.Y.; Choi, H.-R.; Chung, J.-W.; Jeon, Y.-A.; Lee, J.-R.; Ma, K.-H.; Lee, M.-C. Novel microsatellite markers acquired from Rubus coreanus Miq. and cross-amplification in other Rubus species. Molecules 2015, 20, 6432–6442. [Google Scholar] [CrossRef]
- Martins, W.S.; Lucas, D.C.S.; Neves, K.F.S.; Bertioli, D.J. WebSat—A web software for microsatellite marker development. Bioinformation 2009, 3, 282–283. [Google Scholar] [CrossRef]
- Nunes, C.F.; Ferreira, J.L.; Nunes-Fernandes, M.C.; de Souza Breves, S.; Generoso, A.L.; Fontes-Soares, B.D.; Carvalho-Dias, M.S.; Pasqual, M.; Borem, A.; de Almeida Cancado, G.M. An improved method for genomic DNA extraction from strawberry leaves. Ciência Rural 2011, 41, 1383–1389. [Google Scholar] [CrossRef] [Green Version]
- Liu, K.; Muse, S.V. PowerMarker: An integrated analysis environment for genetic marker analysis. Bioinformatics 2005, 21, 2128–2129. [Google Scholar] [CrossRef]
- Ahmad, A.; Wang, J.-D.; Pan, Y.-B.; Rahat Sharif, R.; Gao, S.-J. Development and use of simple sequence repeats (SSRs) markers for sugarcane breeding and genetic studies. Agronomy 2018, 8, 260. [Google Scholar] [CrossRef]
- Fernandez-Fernandez, F.; Antanaviciute, L.; Govan, C.L.; Sargent, D.J. Development of a multiplexed microsatellite set for fingerprinting red raspberry (Rubus idaeus) germplasm and its transferability to other Rubus species. J. Berry Res. 2011, 1, 177–187. [Google Scholar] [CrossRef]
- Han, Y.; Huang, K.; Liu, Y.; Jiao, T.; Ma, G.; Qian, Y.; Wang, P.; Dai, X.; Gao, L.; Xia, T. Functional analysis of two flavanone-3-hydroxylase genes from Camellia sinensis: A critical role in flavonoid accumulation. Genes 2017, 8, 300. [Google Scholar] [CrossRef]
- Tian, J.; Han, Z.; Zhang, J.; Hu, Y.; Song, T.; Yao, Y. The balance of expression of dihydroflavonol 4-reductase and flavonol synthase regulates flavonoid biosynthesis and red foliage coloration in crabapples. Sci. Rep. 2015, 5, 12228. [Google Scholar] [CrossRef]
- Saito, K.; Yonekura-Sakakibara, K.; Nakabayashi, R.; Higashi, Y.; Yamazaki, M.; Tohge, T.; Fernie, A.R. The flavonoid biosynthetic pathway in Arabidopsis: Structural and genetic diversity. Plant Physiol. Biochem. 2013, 72, 21–34. [Google Scholar] [CrossRef]
- Petrussa, E.; Braidot, E.; Zancani, M.; Peresson, C.; Bertolini, A.; Patui, S.; Vianello, A. Plant Flavonoids—Biosynthesis, transport and involvement in stress responses. Int. J. Mol. Sci. 2013, 14, 14950–14973. [Google Scholar] [CrossRef]
- Zheng, D.; Schröder, G.; Schröder, J.; Hrazdina, G. Molecular and biochemical characterization of three aromatic polyketide synthase genes from Rubus idaeus. Plant Mol. Biol. 2001, 46, 1–15. [Google Scholar] [CrossRef]
- García-Gómez, B.; Razi, M.; Salazar, J.A.; Prudencio, A.S.; Ruiz, D.; Dondini, L.; Martínez-Gómez, P. Comparative analysis of SSR markers developed in exon, intron, and intergenic regions and distributed in regions controlling fruit quality traits in Prunus species: Genetic diversity and association studies. Plant Mol. Biol. Rep. 2018, 36, 23–35. [Google Scholar] [CrossRef]
- Holland, J.B.; Helland, S.J.; Sharopova, N.; Rhyne, D.C. Polymorphism of PCR-based markers targeting exons, introns, promoter regions, and SSRs in maize and introns and repeat sequences in oat. Genome 2001, 44, 1065–1076. [Google Scholar] [CrossRef]
- Krutovsky, K.V.; Neale, D.B. Nucleotide diversity and linkage disequilibrium in cold-hardiness and wood quality-related candidate genes in Douglas-fir. Genetics 2005, 171, 2029–2041. [Google Scholar] [CrossRef]
- Cai, C.; Wu, S.; Niu, E.; Cheng, C.; Guo, W. Identification of genes related to salt stress tolerance using intron-length polymorphic markers, association mapping and virus-induced gene silencing in cotton. Sci. Rep. 2017, 7, 528. [Google Scholar] [CrossRef] [Green Version]
- Lewers, K.S.; Weber, C.A. The trouble with genetic mapping of raspberry. HortScience 2005, 40, 1108. [Google Scholar] [CrossRef]
- Anderson, J.R.; Lübberstedt, T. Functional markers in plants. Trends Plant Sci. 2003, 8, 554–560. [Google Scholar] [CrossRef]
- Rafique, M.Z.; Carvalho, E.; Stracke, R.; Palmieri, L.; Herrera, L.; Feller, A.; Malnoy, M.; Martens, S. Nonsense mutation inside anthocyanidin synthase gene controls pigmentation in yellow raspberry (Rubus idaeus L.). Front. Plant Sci. 2016, 7, 1892. [Google Scholar] [CrossRef]
- Michael, T.P.; VanBuren, R. Progress, challenges and the future of crop genomes. Curr. Opin. Plant Biol. 2015, 24, 71–81. [Google Scholar] [CrossRef]
- VanBuren, R.; Bryant, D.; Bushakra, J.M.; Vining, K.J.; Edger, P.P.; Rowley, E.R.; Priest, H.D.; Michael, T.P.; Lyons, E.; Filichkin, S.A.; et al. The genome of black raspberry (Rubus occidentalis). Plant J. 2016, 87, 535–547. [Google Scholar] [CrossRef]
- Wight, H.; Zhou, J.; Li, M.; Hannenhalli, S.; Mount, S.M.; Liu, Z. Draft genome assembly and annotation of red raspberry Rubus idaeus. BioRxiv 2019, 546135, 1–22. [Google Scholar] [CrossRef]
| Cultivar | Abbr. | Genetic Origin and Background | Fruit Color | Origin |
|---|---|---|---|---|
| R. idaeus (red raspberry) | ||||
| Amira | Ami | Polka × Tulameen | red | Italy |
| Anne | Ann | Amity × Glenn Garry | yellow | USA |
| Babye Leto II | BL2 | Autumn Bliss × Babye Leto | red | Russia (Bryansk) |
| Beglyanka | Beg | Kostinbrodskaya × Novost Kuzmina | orange | Russia (Bryansk) |
| Brilliantovaya | Bri | open pollination of interspecific hybrids | red | Russia (Bryansk) |
| Bryanskoe Divo | BrD | 47-18-4 (open pollination) | light-red | Russia (Bryansk) |
| Gerakl | Ger | Autumn Bliss × 14-205-4 | red | Russia (Bryansk) |
| Glen Ample | GAm | SCRI7326EI × SCRI7412H16 | dark red | UK |
| Marosejka | Mar | 7324/50 × 7331/3 | light-red | Russia (Moscow) |
| Meteor | Met | Kostinbrodskaya × Novost Kuzmina | red | Russia (Bryansk) |
| Oranzhevoe Chudo | OrC | Shapka Monomaha (open pollination) | orange | Russia (Bryansk) |
| Pingvin | Pin | interspecific hybrid | dark red | Russia (Bryansk) |
| Polka | Pol | P89141(open pollination) | red | Poland |
| Poranna Rosa | PoR | 83291 × ORUS 1098-1 | yellow | Poland |
| Solnyshko | Sol | Kostinbrodskaya × Novost Kuzmina | red | Russia (Bryansk) |
| Sugana | Sug | Autumn Bliss × Tulameen | light-red | Switzerland |
| Tarusa | Tar | Stolichnaya × Shtambovyj-1 | red | Russia (Moscow) |
| Zheltyj Gigant | ZhG | Marosejka × Ivanovskaya | yellow | Russia (Moscow) |
| Zolotaya Osen | ZOs | 13-39-11 (open pollination) | yellow | Russia (Bryansk) |
| R. occidentalis (black raspberry) | ||||
| Cumberland | Cum | Gregg selfed | blue-black | USA |
| Jewel | Jew | (Bristol × Dundee) × Dundee | black | USA |
| Locus | Gene | Species | NCBI GenBank Accession Number | Motif and Number of Repeats | Location in the Gene | PCR Primer Nucleotide Sequence | T *, °C | Allele Size, bp | |||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Expected | Observed | ||||||||||
| Forward | Reverse | Red Raspberry | Black Raspberry | ||||||||
| RiG001 | aromatic polyketide synthase (PKS3) | R. idaeus | AF292369 | (AT)6 | intron | TGTCCGATCCTTTTCTTTGG | CGCTTCTTGATCCTTGACTTGT | 55 | 345 | 349, 350, 351 | 0 |
| RcFH01 | flavanone-3-hydroxylase (F3H) | R. coreanus | EU255776 | (TATG)3 | intron | GGTCCAAGTGCATTCCATATTAC | GTTCTTGAATCTCCCGTTGCT | 60 | 262 | 255, 265, 271 | 255, 271 |
| FaFS01 | flavonol synthase (FLS) | Fragaria × ananassa | DQ834905 | (CT)12 | intron | CATCCCTAATGCCCTAGTCATC | TGTACTTCGGTGGATTCTCCTT | 60 | 304 | 323, 328 | 323 |
| FaFS02 | flavonol synthase (FLS) | F. ananassa | DQ834905 | (GGAAG)2 | exon | AAGCTCCTCAAACAAATCTTCG | GTAGTTAATGGCAGAAGGTGGC | 60 | 273 | 255, 271 | 271 |
| RiAS01 | anthocyanidin synthase (ANS) | R. idaeus | KX950789 | (ATCTC)2 | exon | TCAACAAGGAGAAGGTGAGGAT | CCGTTAGGAGAGATGAAAGCAG | 60 | 334 | 309, 333, 358 | 309 |
| FaAR01 ** | anthocyanidin reductase (ANR) | F. ananassa | DQ664193 | (TGCTG)2 (CATTT)2 | exon intron | AATCTGCTTCTGGTCGGTACAT | AGAGAGTATGGTCTTCGCCTTG | 60 | 244 | 250 | 250 |
| RhUF01 ** | UDP-glucose flavonoid 3-O-glycosyltransferase-like protein (UFGT) | R. hybrid | JF764808 | (GAG)7 (ACAAGC)2 | exon | AGGAGCTGAAGAAAAGACTCCA | AAAGTCCTCTAGGTTTCCCCTG | 60 | 275 | 267, 270 | 269, 270 |
| RiMY01 ** | transcription factor MYB10 | R. idaeus | EU155165 | (TAATA)2 (CT)7 (AT)15 | introns | GTTCCTCTCCAAGCAGGTTATT | TGCAAAGTCTCCTCTCTTGATG | 59 | 330 | 323, 325, 327, 329, 331, 333, 341, 342 | 358 |
| RiTT01 | transparent testa glabra 1 (TTG1) protein | R. idaeus | HM579852 | (CAC)5 | exon | ACTCCACACAAGAATCCCATCT | CTGTTGTTCAAGACCGAAATTG | 60 | 379 | 379 | 379 |
| Locus | Location in the Gene | Major Allele Frequency | Number of Alleles | Heterozygosity | Polymorphism Information Content (PIC) | |
|---|---|---|---|---|---|---|
| Expected (He) | Observed (Ho) | |||||
| RiG001 | intron | 0.81 | 4 | 0.33 | 0.19 | 0.31 |
| RcFH01 | intron | 0.74 | 3 | 0.41 | 0.52 | 0.35 |
| FaFS01 | intron | 0.76 | 2 | 0.36 | 0.48 | 0.30 |
| FaFS02 | exon | 0.98 | 2 | 0.05 | 0.05 | 0.05 |
| RiAS01 | exon | 0.79 | 3 | 0.36 | 0.19 | 0.33 |
| RhUF01 | exon | 0.90 | 3 | 0.18 | 0.00 | 0.17 |
| RiMY01 | introns | 0.29 | 9 | 0.84 | 0.57 | 0.82 |
| Mean | 0.75 | 3.71 | 0.36 | 0.29 | 0.33 | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lebedev, V.G.; Subbotina, N.M.; Maluchenko, O.P.; Krutovsky, K.V.; Shestibratov, K.A. Assessment of Genetic Diversity in Differently Colored Raspberry Cultivars Using SSR Markers Located in Flavonoid Biosynthesis Genes. Agronomy 2019, 9, 518. https://doi.org/10.3390/agronomy9090518
Lebedev VG, Subbotina NM, Maluchenko OP, Krutovsky KV, Shestibratov KA. Assessment of Genetic Diversity in Differently Colored Raspberry Cultivars Using SSR Markers Located in Flavonoid Biosynthesis Genes. Agronomy. 2019; 9(9):518. https://doi.org/10.3390/agronomy9090518
Chicago/Turabian StyleLebedev, Vadim G., Natalya M. Subbotina, Oleg P. Maluchenko, Konstantin V. Krutovsky, and Konstantin A. Shestibratov. 2019. "Assessment of Genetic Diversity in Differently Colored Raspberry Cultivars Using SSR Markers Located in Flavonoid Biosynthesis Genes" Agronomy 9, no. 9: 518. https://doi.org/10.3390/agronomy9090518





