GC-MS-Based Metabolomics Study of Single- and Dual-Species Biofilms of Candida albicans and Klebsiella pneumoniae
Abstract
1. Introduction
2. Results and Discussion
3. Materials and Methods
3.1. Strains Cultivation
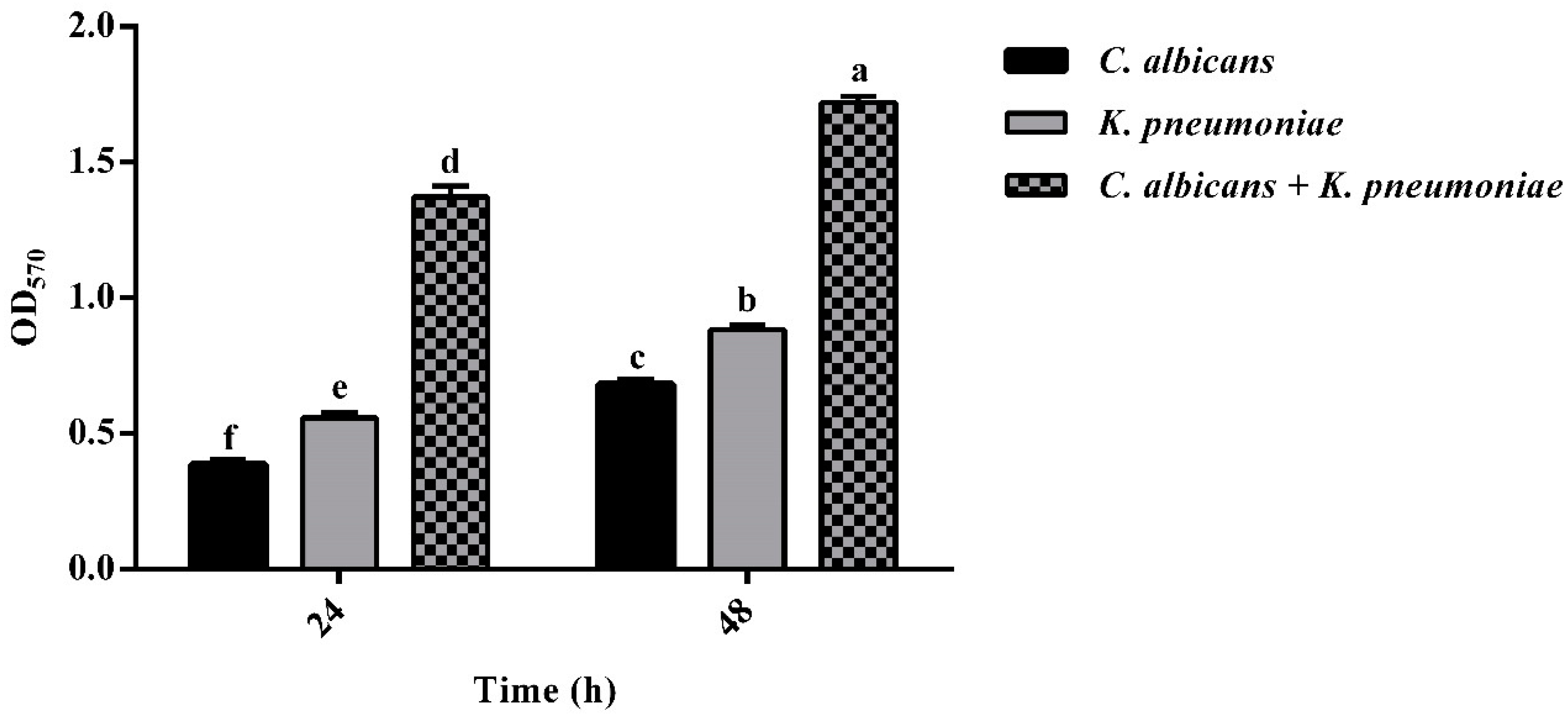
3.2. Development and Quantification of Mono- and Polymicrobial Biofilms
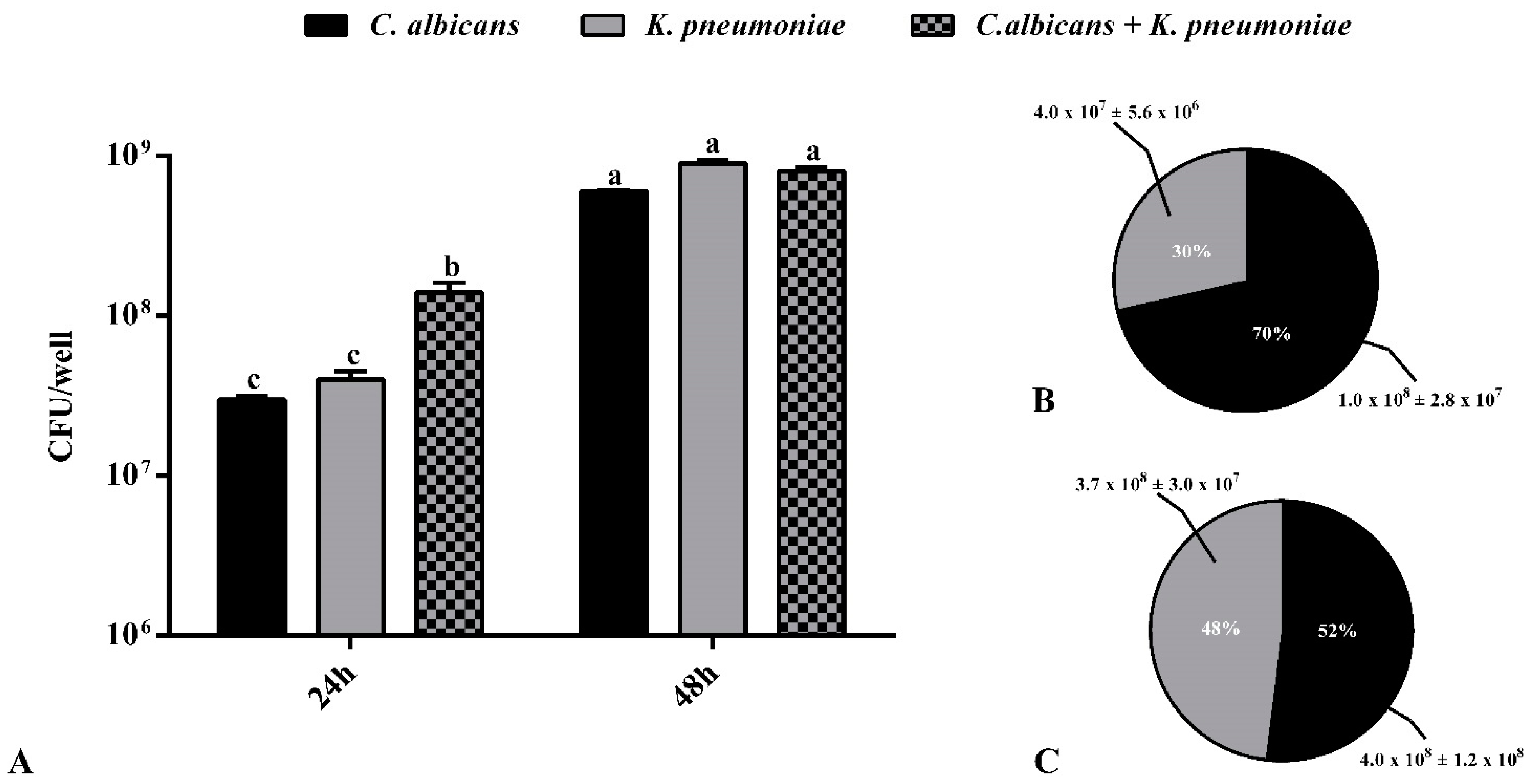
3.3. Quantification of Viable Cells in the Biofilms
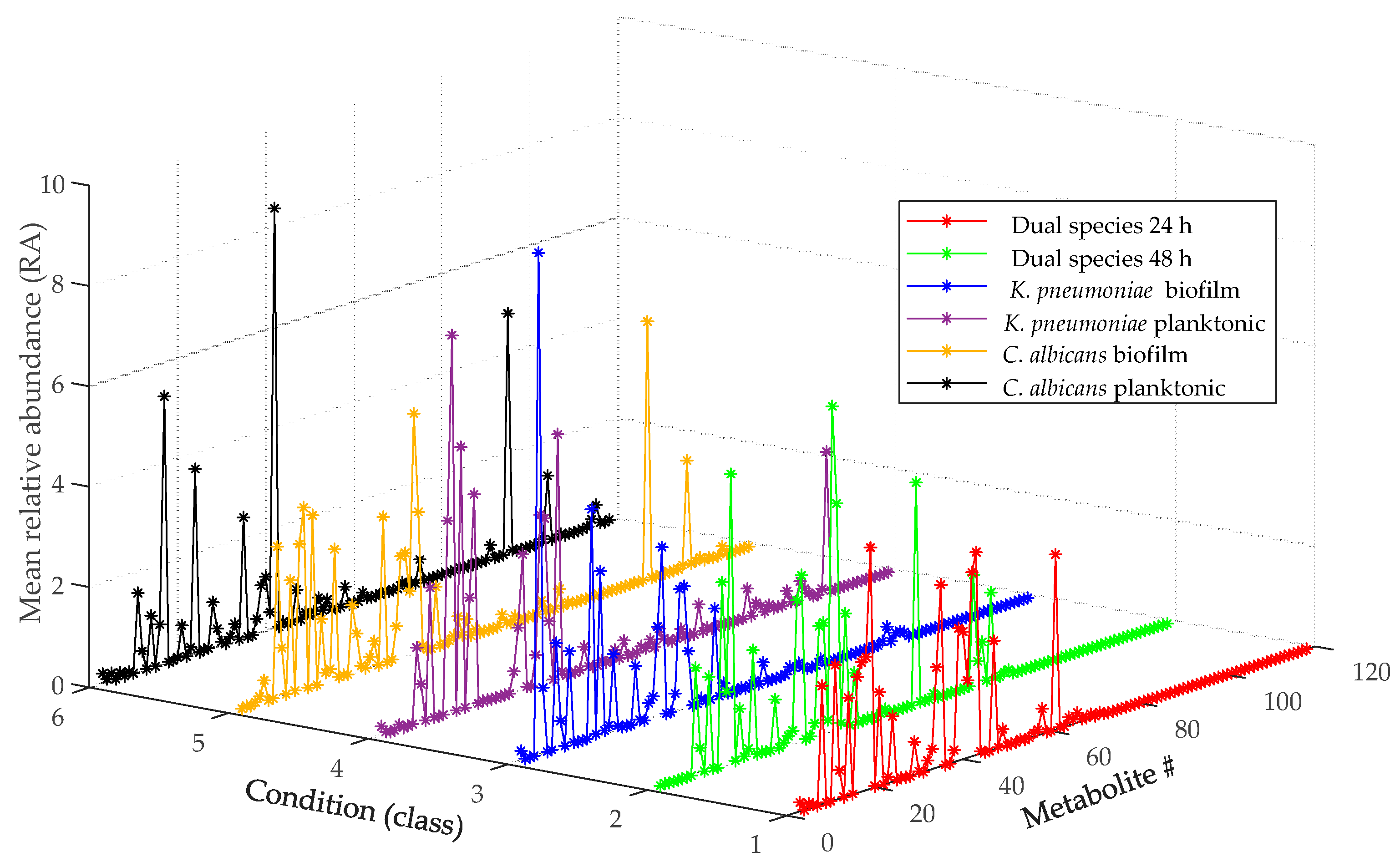
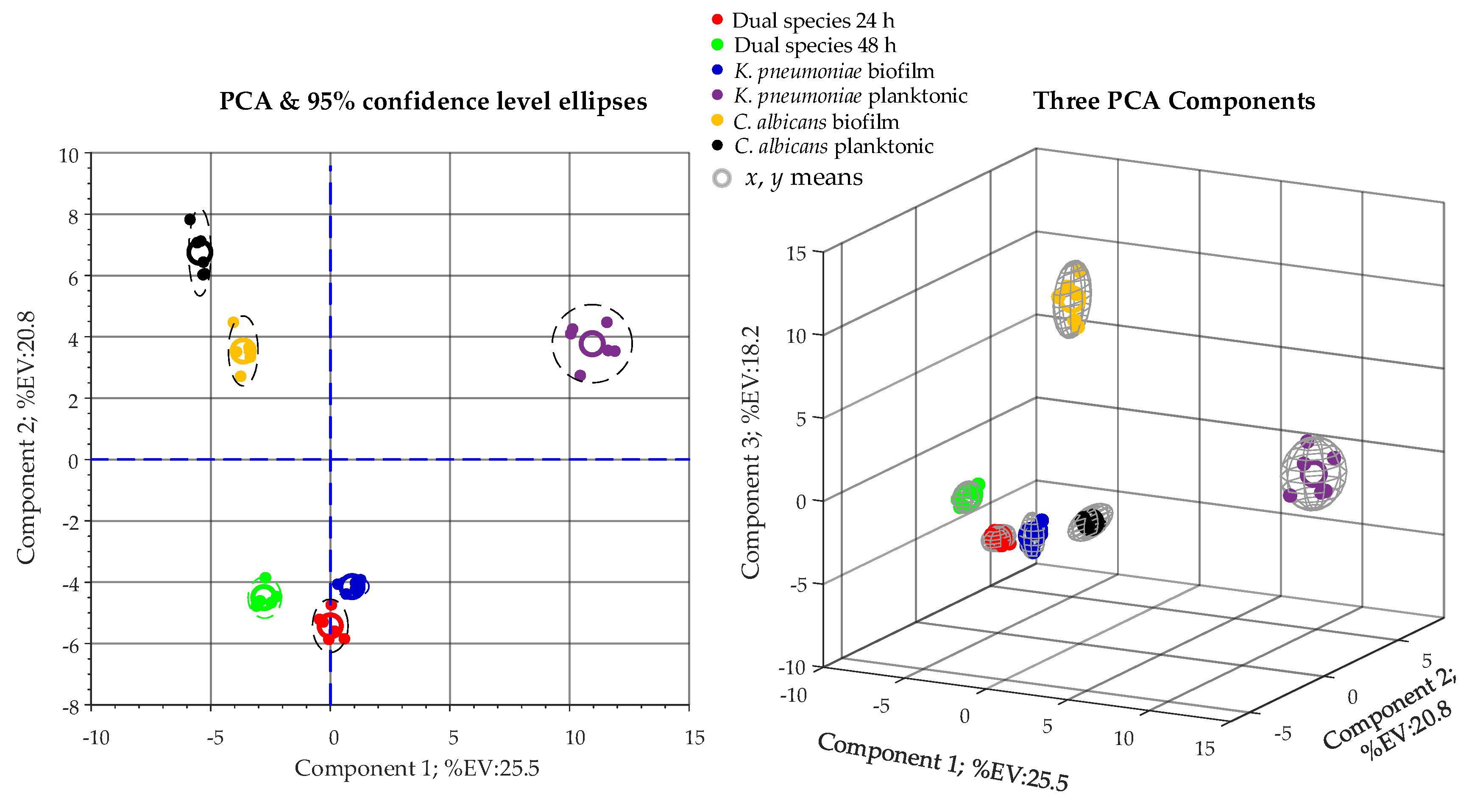
3.4. Metabolomic Analysis
3.4.1. Sample Preparation
3.4.2. GC-MS Analysis
3.4.3. Data Processing and Statistical Analysis
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Pereira, R.; dos Santos Fontenelle, R.O.; de Brito, E.H.S.; de Morais, S.M. Biofilm of Candida albicans: Formation, regulation and resistance. J. Appl. Microbiol. 2020, 1–12. [Google Scholar] [CrossRef]
- Cavalheiro, M.; Teixeira, M.C. Candida Biofilms: Threats, challenges, and promising strategies. Front. Med. 2018, 5, 1–15. [Google Scholar] [CrossRef]
- Rodrigues, M.E.; Gomes, F.; Rodrigues, C.F. Candida spp./bacteria mixed biofilms. J. Fungi 2020, 6, 5. [Google Scholar] [CrossRef]
- Peters, B.M.; Jabra-Rizk, M.A.; O’May, G.A.; William Costerton, J.; Shirtliff, M.E. Polymicrobial interactions: Impact on pathogenesis and human disease. Clin. Microbiol. Rev. 2012, 25, 193–213. [Google Scholar] [CrossRef]
- Ponde, N.O.; Lortal, L.; Ramage, G.; Naglik, J.R.; Jonathan, P. Critical reviews in microbiology Candida albicans biofilms and polymicrobial interactions. Crit. Rev. Microbiol. 2021, 1–21. [Google Scholar] [CrossRef]
- Zago, C.E.; Silva, S.; Sanitá, P.V.; Barbugli, P.A.; Dias, C.M.I.; Lordello, V.B.; Vergani, C.E. Dynamics of biofilm formation and the Interaction between Candida albicans and methicillin-susceptible (MSSA) and -resistant Staphylococcus aureus (MRSA). PLoS ONE 2015, 10, e0123206. [Google Scholar] [CrossRef]
- Huffines, J.T.; Scoffield, J.A. Disruption of Streptococcus mutans and Candida albicans synergy by a commensal streptococcus. Sci. Rep. 2020, 10, 1–10. [Google Scholar] [CrossRef]
- Tang, M.; Kong, X.; Hao, J.; Liu, J. Epidemiological characteristics and formation mechanisms of multidrug-resistant hypervirulent Klebsiella pneumoniae. Front. Microbiol. 2020, 11, 1–10. [Google Scholar] [CrossRef]
- Tacconelli, E.; Carrara, E.; Savoldi, A.; Harbarth, S.; Mendelson, M.; Monnet, D.L.; Pulcini, C.; Kahlmeter, G.; Kluytmans, J.; Carmeli, Y.; et al. Discovery, research, and development of new antibiotics: The WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 2018, 18, 318–327. [Google Scholar] [CrossRef]
- Souza, L.C.D.; Mota, V.B.R.D.; Carvalho, A.V.D.S.Z.D.; Corrêa, R.D.G.C.F.; Libério, S.A.; Lopes, F.F. Association between pathogens from tracheal aspirate and oral biofilm of patients on mechanical ventilation. Braz. Oral Res. 2017, 31, e38. [Google Scholar] [CrossRef]
- Singhai, M.; Mohd, S.; Malik, A.; Malik, A.; Rawat, V. Colonization of peripheral intravascular catheters with biofilm producing microbes: Evaluation of risk factors. Niger. Med. J. 2012, 53, 37. [Google Scholar] [CrossRef]
- Vuotto, C.; Longo, F.; Balice, M.P.; Donelli, G.; Varaldo, P.E. Antibiotic resistance related to biofilm formation in Klebsiella pneumoniae. Pathogens 2014, 3, 743–758. [Google Scholar] [CrossRef]
- Galdiero, E.; Di Onofrio, V.; Maione, A.; Gambino, E.; Gesuele, R.; Menale, B.; Ciaravolo, M.; Carraturo, F.; Guida, M. Allium ursinum and allium oschaninii against Klebsiella pneumoniae and Candida albicans mono-and polymicrobic biofilms in in vitro static and dynamic models. Microorganisms 2020, 8, 336. [Google Scholar] [CrossRef] [PubMed]
- Maione, A.; de Alteriis, E.; Carraturo, F.; Galdiero, S.; Falanga, A.; Guida, M.; Di Cosmo, A.; Maselli, V.; Galdiero, E. The membranotropic peptide gH625 to combat mixed Candida albicans/Klebsiella pneumoniae biofilm: Correlation between in vitro anti-biofilm activity and in vivo antimicrobial protection. J. Fungi 2021, 7, 26. [Google Scholar] [CrossRef]
- Galdiero, E.; Salvatore, M.M.; Maione, A.; Carraturo, F.; Galdiero, S.; Andolfi, A.; Salvatore, F.; Guida, M. Impact of the peptide WMR-K on dual-Species biofilm Candida albicans/Klebsiella pneumoniae and on the untargeted metabolomic profile. Pathogens 2021, 10, 214. [Google Scholar] [PubMed]
- Patejko, M.; Jacyna, J.; Markuszewski, M.J. Sample preparation procedures utilized in microbial metabolomics: An overview. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2017, 1043, 150–157. [Google Scholar]
- Peng, C.; Viana, T.; Petersen, M.A.; Larsen, F.H.; Arneborg, N. Metabolic footprint analysis of metabolites that discriminate single and mixed yeast cultures at two key time-points during mixed culture alcoholic fermentations. Metabolomics 2018, 14, 93. [Google Scholar] [CrossRef]
- de Alteriis, E.; Lombardi, L.; Falanga, A.; Napolano, M.; Galdiero, S.; Siciliano, A.; Carotenuto, R.; Guida, M.; Galdiero, E. Polymicrobial antibiofilm activity of the membranotropic peptide gH625 and its analogue. Microb. Pathog. 2018, 125, 189–195. [Google Scholar] [CrossRef]
- Fox, E.P.; Cowley, E.S.; Nobile, C.J.; Hartooni, N.; Newman, D.K.; Johnson, A.D. Anaerobic bacteria grow within Candida albicans biofilms and induce biofilm formation in suspension cultures. Curr. Biol. 2014, 24, 2411–2416. [Google Scholar] [CrossRef]
- AMDIS. Available online: www.amdis.net (accessed on 9 February 2021).
- Styczynski, M.P.; Moxley, J.F.; Tong, L.V.; Walther, J.L.; Jensen, K.L.; Stephanopoulos, G.N. Systematic identification of conserved metabolites in GC/MS data for metabolomics and biomarker discovery. Anal. Chem. 2007, 79, 966–973. [Google Scholar] [CrossRef]
- MATLAB. Available online: www.mathworks.com (accessed on 1 December 2020).
- Salvatore, M.M.; Nicoletti, R.; Salvatore, F.; Naviglio, D.; Andolfi, A. GC–MS approaches for the screening of metabolites produced by marine-derived Aspergillus. Mar. Chem. 2018, 206, 19–33. [Google Scholar] [CrossRef]
- Kart, D.; Yabanoglu Ciftci, S.; Nemutlu, E. Altered metabolomic profile of dual-species biofilm: Interactions between Proteus mirabilis and Candida albicans. Microbiol. Res. 2020, 230, 126346. [Google Scholar] [CrossRef]
- Nett, J.E.; Lepak, A.J.; Marchillo, K.; Andes, D.R. Time course global gene expression analysis of an in vivo Candida biofilm. J. Infect. Dis. 2009, 200, 307–313. [Google Scholar] [CrossRef]
- Zhu, Z.; Wang, H.; Shang, Q.; Jiang, Y.; Cao, Y.; Chai, Y. Time course analysis of Candida albicans metabolites during biofilm development. J. Proteome Res. 2013, 12, 2375–2385. [Google Scholar] [CrossRef] [PubMed]
- Branda, S.S.; Vik, Å.; Friedman, L.; Kolter, R. Biofilms: The matrix revisited. Trends Microbiol. 2005, 13, 20–26. [Google Scholar] [CrossRef]
- Díaz-Pérez, A.L.; Díaz-Pérez, C.; Campos-García, J. Bacterial l-leucine catabolism as a source of secondary metabolites. Rev. Environ. Sci. Biotechnol. 2016, 15, 1–29. [Google Scholar] [CrossRef]
- Sánchez-Fresneda, R.; Guirao-Abad, J.P.; Argüelles, A.; González-Párraga, P.; Valentín, E.; Argüelles, J.C. Specific stress-induced storage of trehalose, glycerol and d-arabitol in response to oxidative and osmotic stress in Candida albicans. Biochem. Biophys. Res. Commun. 2013, 430, 1334–1339. [Google Scholar] [CrossRef] [PubMed]
- Kordowska-Wiater, M. Production of arabitol by yeasts: Current status and future prospects. J. Appl. Microbiol. 2015, 119, 303–314. [Google Scholar] [CrossRef] [PubMed]
- Domergue, R.; Castaño, I.; De Las Peñas, A.; Zupancic, M.; Lockatell, V.; Hebel, J.R.; Johnson, D.; Cormack, B.P. Nicotinic acid limitation regulates silencing of Candida adhesins during UTI. Science 2005, 308, 866–870. [Google Scholar] [CrossRef] [PubMed]
- Ma, B.; Pan, S.J.; Zupancic, M.L.; Cormack, B.P. Assimilation of NAD+ precursors in Candida glabrata. Mol. Microbiol. 2007, 66, 14–25. [Google Scholar] [CrossRef]
- Park, S.J.; Han, K.H.; Park, J.Y.; Choi, S.J.; Lee, K.H. Influence of bacterial presence on biofilm formation of Candida albicans. Yonsei Med. J. 2014, 55, 449–458. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.; Kim, N.J.; Kim, S.Y.; Kim, I.H.; Kim, K.S.; Lee, G.R. Cyclo(Phe-Pro) produced by the human pathogen Vibrio vulnificus inhibits host innate immune responses through the NF-κB pathway. Infect. Immun. 2015, 83, 1150–1161. [Google Scholar] [CrossRef] [PubMed]
- Salvatore, M.M.; Félix, C.; Lima, F.; Ferreira, V.; Duarte, A.S.; Salvatore, F.; Alves, A.; Esteves, A.C.; Andolfi, A. Effect of γ-aminobutyric acid (GABA) on the metabolome of two strains of Lasiodiplodia theobromae isolated from grapevine. Molecules 2020, 25, 3833. [Google Scholar] [CrossRef] [PubMed]
- Salvatore, M.M.; DellaGreca, M.; Nicoletti, R.; Salvatore, F.; Vinale, F.; Naviglio, D.; Andolfi, A. Talarodiolide, a new 12-membered macrodiolide, and GC/MS investigation of culture filtrate and mycelial extracts of Talaromyces pinophilus. Molecules 2018, 23, 950. [Google Scholar] [CrossRef]
- Mooney, S.; Leuendorf, J.E.; Hendrickson, C.; Hellmann, H. Vitamin B6: A long known compound of surprising complexity. Molecules 2009, 14, 329–351. [Google Scholar] [CrossRef] [PubMed]
- Richards, A.B.; Krakowka, S.; Dexter, L.B.; Schmid, H.; Wolterbeek, A.P.M.; Waalkens-Berendsen, D.H.; Shigoyuki, A.; Kurimoto, M. Trehalose: A review of properties, history of use and human tolerance, and results of multiple safety studies. Food Chem. Toxicol. 2002, 40, 871–898. [Google Scholar] [CrossRef]
- Alvarez-Peral, F.J.; Zaragoza, O.; Pedreño, Y.; Argüelles, J.C. Protective role of trehalose during severe oxidative stress caused by hydrogen peroxide and the adaptive oxidative stress response in Candida albicans. Microbiology 2002, 148, 2599–2606. [Google Scholar] [CrossRef]
- Cao, Y.; Zhu, Z.; Chen, X.; Yao, X.; Zhao, L.; Wang, H.; Yan, L.; Wu, H.; Chai, Y.; Jiang, Y. Effect of amphotericin B on the metabolic profiles of Candida albicans. J. Proteome Res. 2013, 12, 2921–2932. [Google Scholar] [CrossRef]
- Thevelein, J.M. Regulation of trehalose mobilization in fungi. Microbiol. Rev. 1984, 48, 42–59. [Google Scholar] [CrossRef]
- Ballén, V.; Ratia, C.; Cepas, V.; Soto, S.M. Enterococcus faecalis inhibits Klebsiella pneumoniae growth in polymicrobial biofilms in a glucose-enriched medium. Biofouling 2020, 36, 846–861. [Google Scholar] [CrossRef]
- Guida, M.; Salvatore, M.M.; Salvatore, F. A strategy for GC/MS quantification of polar compounds via their silylated surrogates: Silylation and quantification of biological amino acids. J. Anal. Bioanal. Tech. 2015, 6, 1. [Google Scholar]
- NIST 14. Available online: https://www.nist.gov/srd/nist-standard-reference-database-1a (accessed on 9 February 2021).
- Golm Metabolome Database. Available online: http://gmd.mpimp-golm.mpg.de/ (accessed on 9 February 2021).
- Hummel, J.; Strehmel, N.; Selbig, J.; Walther, D.; Kopka, J. Decision tree supported substructure prediction of metabolites from GC-MS profiles. Metabolomics 2010, 6, 322–333. [Google Scholar] [CrossRef]
- Koo, I.; Shi, X.; Kim, S.; Zhang, X. IMatch2: Compound identification using retention index for analysis of gas chromatography-mass spectrometry data. J. Chromatogr. A 2014, 1337, 202–210. [Google Scholar] [CrossRef]

| Pairwise Comparisons | Univariate Statistical Test | Multivariate Analysis | R2X | R2Y | Q2Y |
|---|---|---|---|---|---|
| (C. albicans planktonic) vs. (C. albicans biofilm) | Student’s t-test | PLS-DA | 0.7287 | 0.9996 | 0.9876 |
| (K. pneumoniae planktonic) vs. (K. pneumoniae biofilm) | Student’s t-test | PLS-DA | 0.6749 | 0.9983 | 0.9663 |
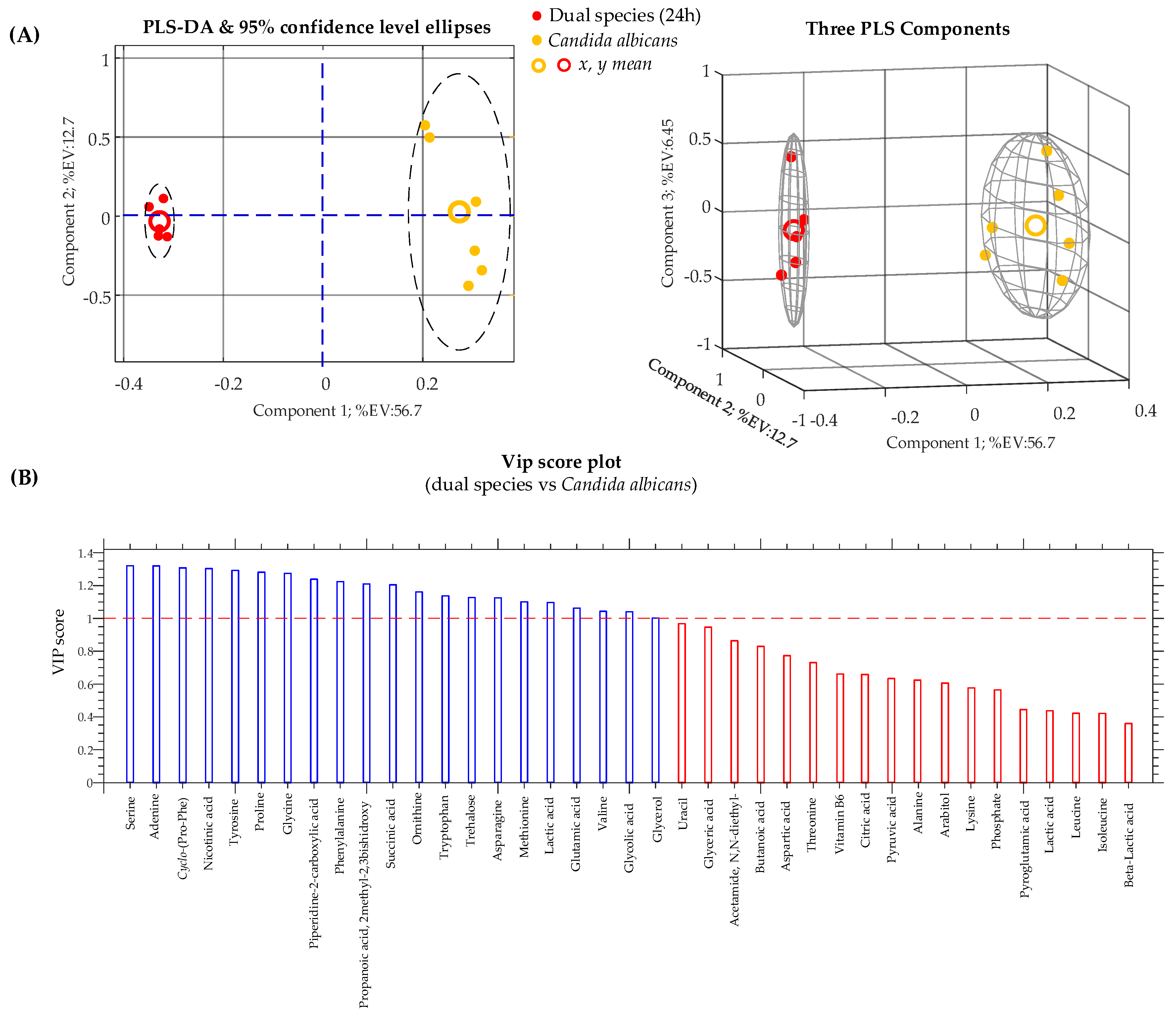
| (C. albicans/K. pneumoniae biofilm 24 h) vs. (C. albicans biofilm 24 h) | Student’s t-test | PLS-DA | 0.7348 | 0.9994 | 0.9803 |
| (C. albicans/K. pneumoniae biofilm 24 h) vs. (K. pneumoniae biofilm 24 h) | Student’s t-test | PLS-DA | 0.6368 | 0.9952 | 0.9261 |
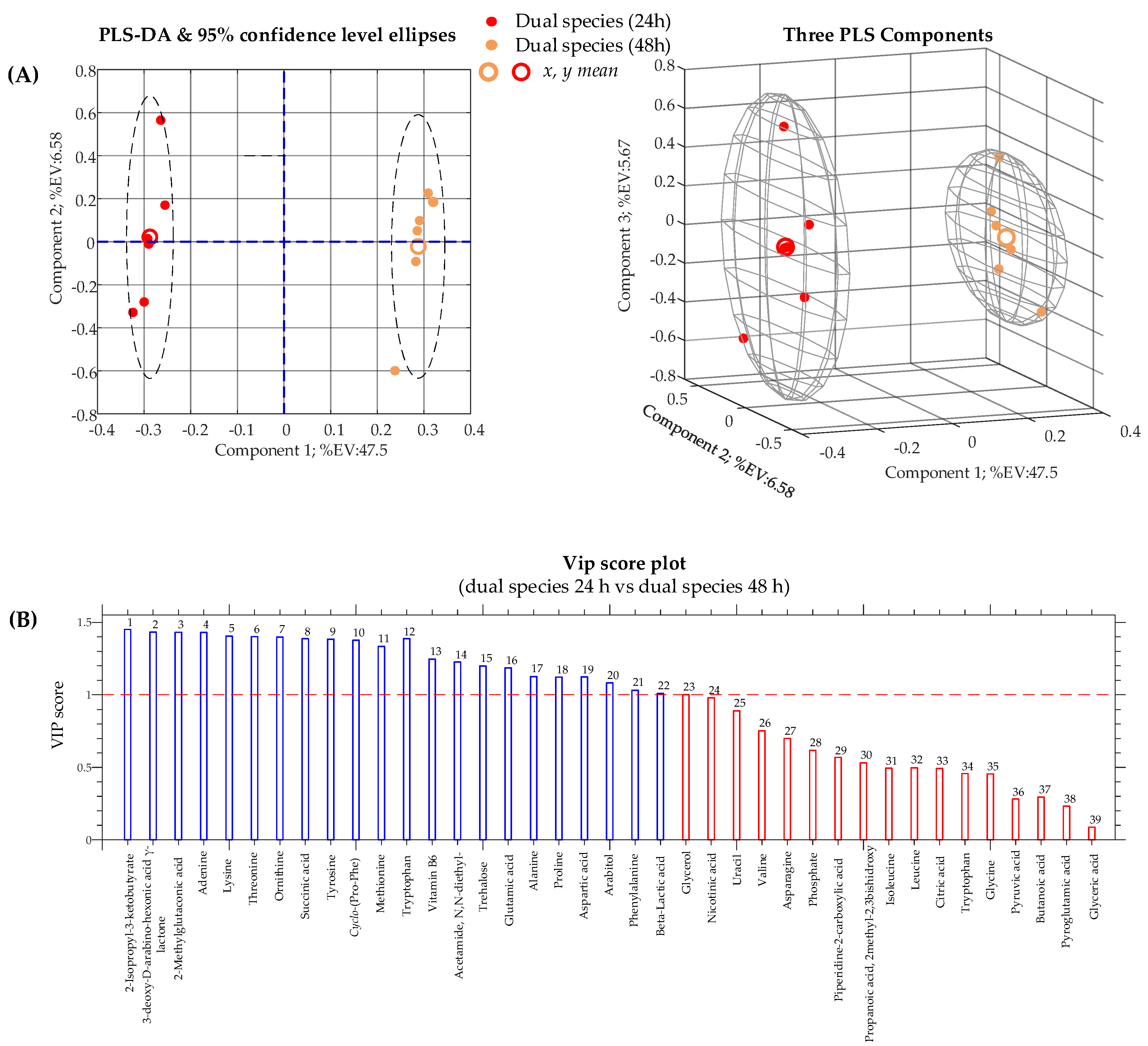
| (C. albicans/K. pneumoniae biofilm 24 h) vs. (C. albicans/K. pneumoniae biofilm 48 h) | Student’s t-test | PLS-DA | 0.5975 | 0.9988 | 0.9988 |
| Dual Species vs. C. albicans | Dual Species vs. K. pneumoniae | |||||
|---|---|---|---|---|---|---|
| Name | VIP Score | t-Test p-Value | Fold Change | VIP Score | t-Test p-Value | Fold Change |
| Acetamide, N,N-diethyl- (RI: 1045) | 0.864 | 3.10 × 10−2 | =2.24 | 0.264 | 6.70 × 10−2 | =1.13 |
| Adenine, TMS (RI: 1890) | 1.320 | 2.86 × 10−9 | ↑ 25.63 | 1.561 | 3.12 × 10−9 | ↑ 19.85 |
| Alanine, 2TMS (RI: 1124) | 0.623 | 1.76 × 10−1 | =1.15 | 1.076 | 2.16 × 10−2 | ↑ 1.24 |
| Arabitol, 5TMS (RI: 1750) | 0.606 | 1.78 × 10−1 | =1.28 | 1.573 | 4.22 × 10−13 | ↑ 883.44 |
| Asparagine, 3TMS (RI: 1687) | 1.125 | 2.56 × 10−3 | ↓ 1.24 | 1.490 | 1.26 × 10−7 | ↓ 1.73 |
| Aspartic acid, 3TMS (RI: 1540) | 0.773 | 6.12 × 10−2 | =1.59 | 0.454 | 3.94 × 10−1 | =1.16 |
| Beta-lactic acid, 2TMS (RI: 1156) | 0.359 | 4.38 × 10−1 | =1.17 | 1.278 | 2.32 × 10−3 | ↑ 1.76 |
| Butanoic acid, 3TMS (RI: 1425) | 0.829 | 4.17 × 10−2 | =2.24 | 0.385 | 5.92 × 10−1 | =1.03 |
| Citric acid, 4TMS (RI: 1844) | 0.659 | 1.28 × 10−1 | =1.46 | 0.706 | 1.73 × 10−1 | =1.58 |
| Cyclo-(Phe-Pro) (RI: 2434) | 1.308 | 5.26 × 10−8 | ↓ 8.84 | 1.293 | 1.87 × 10−3 | ↓ 4.51 |
| 3-Deoxy-D-arabino-hexonic acid γ-lactone, 3TMS (RI: 1797) | – | 1.276 | 2.37 × 10−3 | ↓ 1.30 | ||
| Glutamic acid, 3TMS (RI: 1638) | 1.062 | 3.35 × 10−3 | ↑ 1.33 | 0.778 | 1.28 × 10−1 | =1.12 |
| Glyceric acid, 3TMS (RI: 1346) | 0.946 | 1.76 × 10−2 | =1.34 | 1.538 | 1.73 × 10−7 | ↓ 4.81 |
| Glycerol, 3TMS (RI: 1290) | 1.002 | 7.48 × 10−3 | ↓ 1.33 | 1.524 | 1.17 × 10−6 | ↑ 644.01 |
| Glycine, 2TMS (RI: 1136) | 1.274 | 3.46 × 10−6 | ↓ 1.83 | 0.223 | 9.37 × 10−1 | =1.01 |
| Glycolic acid (2TMS) (RI: 1100) | 1.040 | 4.75 × 10−3 | ↓ 2.15 | – | ||
| Isoleucine, 2TMS (RI: 1307) | 0.421 | 3.56 × 10−1 | =1.60 | 0.694 | 1.82 × 10−1 | =1.60 |
| Lactic acid, 2TMS (RI: 1083) | 0.437 | 3.42 × 10−1 | =10.26 | 0.971 | 4.57 × 10−2 | =7.38 |
| Leucine, 2TMS (RI: 1286) | 0.422 | 3.50 × 10−1 | =1.32 | 1.189 | 7.00 × 10−3 | ↑ 523.57 |
| Lysine, 3TMS (RI: 1722) | 0.576 | 1.98 × 10−1 | =1.23 | 1.212 | 6.35 × 10−3 | ↑ 1.41 |
| Methionine, 2TMS (RI: 1536) | 1.101 | 1.70 × 10−3 | ↓ 1.21 | 0.374 | 7.28 × 10−1 | =1.02 |
| Nicotinic acid, TMS (RI: 1304) | 1.304 | 1.22 × 10−7 | ↓ 2.51 | 1.231 | 4.39 × 10−3 | ↓ 1.36 |
| Ornithine, 3TMS (RI: 1632) | 1.161 | 4.61 × 10−4 | ↑ 4.34 | 0.583 | 2.86 × 10−1 | =1.20 |
| Phenylalanine, 2TMS (RI: 1647) | 1.224 | 6.09 × 10−5 | ↑ 1.26 | 0.884 | 8.34 × 10−2 | =1.07 |
| Phosphate, 3TMS (RI: 1297) | 0.564 | 2.12 × 10−1 | =1.48 | 0.242 | 7.02 × 10−1 | =1.11 |
| Piperidine-2-carboxylic acid, 2TMS (RI: 1624) | 1.239 | 3.40 × 10−5 | ↓ 2.69 | 1.299 | 1.78 × 10−3 | ↓ 1.38 |
| Proline, 2TMS (RI: 1314) | 1.282 | 1.88 × 10−6 | ↓ 11.44 | 1.127 | 1.34 × 10−2 | ↓ 3.60 |
| Propanoic acid, 2methyl-2,3bishidroxy, 3TMS (RI: 1336) | 1.210 | 1.08 × 10−4 | ↓ 5.16 | – | ||
| Pyroglutamic acid, 2TMS (RI: 1546) | 0.444 | 3.28 × 10−1 | =1.23 | 0.669 | 2.06 × 10−1 | =1.08 |
| Pyruvic acid, 2TMS (RI: 1108) | 0.633 | 1.67 × 10−1 | =2.91 | 0.367 | 5.47 × 10−1 | =1.19 |
| Serine, 3TMS (RI: 1375) | 1.320 | 1.49 × 10−9 | ↓ 307.99 | – | ||
| Shikimic acid, 4TMS (RI: 1951) | – | 1.476 | 1.71 × 10−5 | ↓ 2.98 | ||
| Succinic acid, 2TMS (RI: 1322) | 1.205 | 1.34 × 10−4 | ↓ 1.77 | 0.141 | 1 | =1.00 |
| Threonine, 3TMS (RI: 1400) | 0.730 | 9.76 × 10−2 | =1.14 | 1.540 | 1.58 × 10−7 | ↓ 1.71 |
| Trehalose, 8TMS (RI: 2781) | 1.127 | 1.06 × 10−3 | ↓ 10.10 | – | ||
| Tryptophan, 3TMS (RI: 2244) | 1.138 | 4.53 × 10−4 | ↑ 1.90 | 1.119 | 1.29 × 10−2 | ↑ 1.28 |
| Tyrosine, 3TMS (RI: 1962) | 1.293 | 3.24 × 10−6 | ↑ 1.81 | 1.323 | 3.28 × 10−4 | ↑ 1.21 |
| Uracil, 2TMS (RI: 1351) | 0.968 | 1.20 × 10−2 | =1.67 | 0.901 | 1.05 × 10−1 | =1.38 |
| Valine, 2TMS (RI: 1230) | 1.043 | 4.41 × 10−3 | ↓ 1.16 | 0.512 | 3.37 × 10−1 | =1.05 |
| Vitamin B6, 3TMS (RI: 1924) | 0.660 | 7.28 × 10−1 | =1.36 | 0.564 | 1.76 × 10−1 | =1.09 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Galdiero, E.; Salvatore, M.M.; Maione, A.; de Alteriis, E.; Andolfi, A.; Salvatore, F.; Guida, M. GC-MS-Based Metabolomics Study of Single- and Dual-Species Biofilms of Candida albicans and Klebsiella pneumoniae. Int. J. Mol. Sci. 2021, 22, 3496. https://doi.org/10.3390/ijms22073496
Galdiero E, Salvatore MM, Maione A, de Alteriis E, Andolfi A, Salvatore F, Guida M. GC-MS-Based Metabolomics Study of Single- and Dual-Species Biofilms of Candida albicans and Klebsiella pneumoniae. International Journal of Molecular Sciences. 2021; 22(7):3496. https://doi.org/10.3390/ijms22073496
Chicago/Turabian StyleGaldiero, Emilia, Maria Michela Salvatore, Angela Maione, Elisabetta de Alteriis, Anna Andolfi, Francesco Salvatore, and Marco Guida. 2021. "GC-MS-Based Metabolomics Study of Single- and Dual-Species Biofilms of Candida albicans and Klebsiella pneumoniae" International Journal of Molecular Sciences 22, no. 7: 3496. https://doi.org/10.3390/ijms22073496
APA StyleGaldiero, E., Salvatore, M. M., Maione, A., de Alteriis, E., Andolfi, A., Salvatore, F., & Guida, M. (2021). GC-MS-Based Metabolomics Study of Single- and Dual-Species Biofilms of Candida albicans and Klebsiella pneumoniae. International Journal of Molecular Sciences, 22(7), 3496. https://doi.org/10.3390/ijms22073496









