Journal Description
Microorganisms
Microorganisms
is a scientific, peer-reviewed, open access journal of microbiology, published monthly online by MDPI. The Hellenic Society Mikrobiokosmos (MBK), the Spanish Society for Nitrogen Fixation (SEFIN) and the Society for Microbial Ecology and Disease (SOMED) are affiliated with the Microorganisms, and their members receive a discount on the article processing charges.
- Open Access— free for readers, with article processing charges (APC) paid by authors or their institutions.
- High Visibility: indexed within Scopus, SCIE (Web of Science), PubMed, PMC, PubAg, CAPlus / SciFinder, AGRIS, and other databases.
- Journal Rank: JCR - Q2 (Microbiology) / CiteScore - Q2 (Microbiology (medical))
- Rapid Publication: manuscripts are peer-reviewed and a first decision is provided to authors approximately 15.1 days after submission; acceptance to publication is undertaken in 2.8 days (median values for papers published in this journal in the second half of 2023).
- Recognition of Reviewers: reviewers who provide timely, thorough peer-review reports receive vouchers entitling them to a discount on the APC of their next publication in any MDPI journal, in appreciation of the work done.
- Testimonials: See what our editors and authors say about the Microorganisms.
- Companion journal: Applied Microbiology.
Impact Factor:
4.5 (2022);
5-Year Impact Factor:
4.8 (2022)
Latest Articles
Bloodstream Infection Caused by Erysipelothrix rhusiopathiae in an Immunocompetent Patient
Microorganisms 2024, 12(5), 942; https://doi.org/10.3390/microorganisms12050942 (registering DOI) - 07 May 2024
Abstract
Erysipelothrix rhusiopathiae is a facultative anaerobe Gram-positive bacillus, which is considered a zoonotic pathogen. E. rhusiopathiae causes erysipeloid, mainly in occupational groups such as veterinarians, slaughterhouse workers, farmers, and fishermen. Two cutaneous forms (localised and generalised) and a septicaemic form have been described. Here,
[...] Read more.
Erysipelothrix rhusiopathiae is a facultative anaerobe Gram-positive bacillus, which is considered a zoonotic pathogen. E. rhusiopathiae causes erysipeloid, mainly in occupational groups such as veterinarians, slaughterhouse workers, farmers, and fishermen. Two cutaneous forms (localised and generalised) and a septicaemic form have been described. Here, we report the isolation of a strain of E. rhusiopathiae from a 56-year-old immunocompetent obese male admitted to Fondazione IRCCS Policlinico San Matteo Pavia (Italy). Blood cultures were collected and Gram-positive bacilli were observed. E. rhusiopathiae grew and was identified. Antimicrobial susceptibility tests were performed and interpreted with EUCAST breakpoints (PK-PD). The strain was susceptible to all the antibiotics tested, while it was intrinsically resistant to vancomycin. The clinical diagnosis of E. rhusiopathiae can be challenging, due to the broad spectrum of symptoms and potential side effects, including serious systemic infections such as heart diseases. In the case described, bacteraemia caused by E. rhusiopathiae was detected in a immunocompetent patient. Bacteraemia caused by E. rhusiopathiae is rare in immunocompetent people and blood cultures were proven to be essential for the diagnosis and underdiagnosis of this pathogen, which is possible due to its resemblance to other clinical manifestations.
Full article
(This article belongs to the Section Public Health Microbiology)
►
Show Figures
Open AccessArticle
Canine Visceral Leishmaniasis: A Histological and Immunohistochemical Study of Fibropoiesis in Chronic Interstitial Pneumonitis
by
Frederico C. Gonçalves, Ramon de Alencar Pereira, Adriano Francisco Alves, Aldair Pinto Woyames Junio, Ricardo T. Fujiwara, David M. Mosser, Helida Monteiro Andrade, Geovanni D. Cassali, Enio Ferreira and Wagner Luiz Tafuri
Microorganisms 2024, 12(5), 941; https://doi.org/10.3390/microorganisms12050941 (registering DOI) - 07 May 2024
Abstract
We studied some fibrotic aspects of chronic interstitial pneumonitis in the lungs of dogs infected with Leishmania infantum. The lungs of eleven naturally infected dogs, twelve experimentally infected with two distinct strains of L. infantum (BH401 and BH46), and six uninfected (controls)
[...] Read more.
We studied some fibrotic aspects of chronic interstitial pneumonitis in the lungs of dogs infected with Leishmania infantum. The lungs of eleven naturally infected dogs, twelve experimentally infected with two distinct strains of L. infantum (BH401 and BH46), and six uninfected (controls) dogs, were analyzed by histological, parasitological, and immunohistochemical studies. Conventional histology (HE), collagen deposition (Gomori’s silver staining for reticulin collagen fibers), and immunohistochemistry for myofibroblast characterization were carried out based on the cellular expression of alpha-smooth muscle actin, vimentin, cytokeratin, E-cadherin, snail antigen homologue 1 (SNAI1) (Snail), and the cytokine expression of transforming growth factor-beta (TGF-β). Parasitological screening was carried out using conventional polymerase chain reaction (PCR) and the immunohistochemical reaction of streptavidin–peroxidase for visualizing Leishmania amastigotes. Dogs naturally infected with L. infantum and experimentally infected with L. infantum BH401 strains showed intense interstitial pneumonitis characterized by thickening of the alveolar septa as a consequence of an intense diffuse and focal (plaques) chronic exudate of mononuclear cells associated with fibrogenesis. The expression of alpha-actin, vimentin, and TGF-β was higher in the lung interstitium of all infected dogs than in the other two groups (BH46 strain and controls). Moreover, in both the naturally and experimentally infected dog (BH401 strain) groups, the expression of Snail was moderate to intense in contrast to the other groups. Based on these immunohistochemical results, we concluded that mesenchymal cells are active in promoting changes in the extracellular matrix in the lungs of dogs naturally and experimentally infected with L. infantum, but it depends on the virulence of the parasite.
Full article
(This article belongs to the Special Issue New Advancements in the Field of Leishmaniasis)
►▼
Show Figures

Figure 1
Open AccessReview
Structure, Regulation, and Significance of Cyanobacterial and Chloroplast Adenosine Triphosphate Synthase in the Adaptability of Oxygenic Photosynthetic Organisms
by
Siyan Yi, Xin Guo, Wenjing Lou, Shaoming Mao, Guodong Luan and Xuefeng Lu
Microorganisms 2024, 12(5), 940; https://doi.org/10.3390/microorganisms12050940 - 06 May 2024
Abstract
In cyanobacteria and chloroplasts (in algae and plants), ATP synthase plays a pivotal role as a photosynthetic membrane complex responsible for producing ATP from adenosine diphosphate and inorganic phosphate, utilizing a proton motive force gradient induced by photosynthesis. These two ATP synthases exhibit
[...] Read more.
In cyanobacteria and chloroplasts (in algae and plants), ATP synthase plays a pivotal role as a photosynthetic membrane complex responsible for producing ATP from adenosine diphosphate and inorganic phosphate, utilizing a proton motive force gradient induced by photosynthesis. These two ATP synthases exhibit similarities in gene organization, amino acid sequences of subunits, structure, and functional mechanisms, suggesting that cyanobacterial ATP synthase is probably the evolutionary precursor to chloroplast ATP synthase. In this review, we explore the precise synthesis and assembly of ATP synthase subunits to address the uneven stoichiometry within the complex during transcription, translation, and assembly processes. We also compare the regulatory strategies governing ATP synthase activity to meet varying energy demands in cyanobacteria and chloroplasts amid fluctuating natural environments. Furthermore, we delve into the role of ATP synthase in stress tolerance and photosynthetic carbon fixation efficiency in oxygenic photosynthetic organisms (OPsOs), along with the current researches on modifying ATP synthase to enhance carbon fixation efficiency under stress conditions. This review aims to offer theoretical insights and serve as a reference for understanding the functional mechanisms of ATP synthase, sparking innovative ideas for enhancing photosynthetic carbon fixation efficiency by utilizing ATP synthase as an effective module in OPsOs.
Full article
(This article belongs to the Section Environmental Microbiology)
Open AccessEditorial
Clinical and Environmental Surveillance for the Prevention of Legionellosis
by
Maria Anna Coniglio and Mohamed H. Yassin
Microorganisms 2024, 12(5), 939; https://doi.org/10.3390/microorganisms12050939 - 06 May 2024
Abstract
Legionella is a Gram-negative bacterium whose natural hosts are aquatic protozoa, in which the microorganism replicates and is protected from adverse environmental conditions [...]
Full article
(This article belongs to the Special Issue Clinical and Environmental Surveillance for the Prevention of Legionellosis)
Open AccessArticle
Straw from Different Crop Species Recruits Different Communities of Lignocellulose-Degrading Microorganisms in Black Soil
by
Chunling Chang, Yue Guo, Kuanqiang Tang, Yunlong Hu, Weihui Xu, Wenjing Chen, Neil McLaughlin and Zhigang Wang
Microorganisms 2024, 12(5), 938; https://doi.org/10.3390/microorganisms12050938 - 05 May 2024
Abstract
The biological degradation of plant residues in the soil or on the soil surface is an integral part of the natural life cycle of annual plants and does not have adverse effects on the environment. Crop straw is characterized by a complex structure
[...] Read more.
The biological degradation of plant residues in the soil or on the soil surface is an integral part of the natural life cycle of annual plants and does not have adverse effects on the environment. Crop straw is characterized by a complex structure and exhibits stability and resistance to rapid microbial decomposition. In this study, we conducted a microcosm experiment to investigate the dynamic succession of the soil microbial community and the functional characteristics associated with lignocellulose-degrading pathways. Additionally, we aimed to identify lignocellulose-degrading microorganisms from the straw of three crop species prevalent in Northeast China: soybean (Glycine max Merr.), rice (Oryza sativa L.), and maize (Zea mays L.). Our findings revealed that both the type of straw and the degradation time influenced the bacterial and fungal community structure and composition. Metagenome sequencing results demonstrated that during degradation, different straw types assembled carbohydrate-active enzymes (CAZymes) and KEGG pathways in distinct manners, contributing to lignocellulose and hemicellulose degradation. Furthermore, isolation of lignocellulose-degrading microbes yielded 59 bacterial and 14 fungal strains contributing to straw degradation, with fungi generally exhibiting superior lignocellulose-degrading enzyme production compared to bacteria. Experiments were conducted to assess the potential synergistic effects of synthetic microbial communities (SynComs) comprising both fungi and bacteria. These SynComs resulted in a straw weight loss of 42% at 15 days post-inoculation, representing a 22% increase compared to conditions without any SynComs. In summary, our study provides novel ecological insights into crop straw degradation by microbes.
Full article
(This article belongs to the Special Issue Biotechnology for Environmental Remediation)
Open AccessArticle
Addressing the Concern of Orange-Yellow Fungus Growth on Palm Kernel Cake: Safeguarding Dairy Cattle Diets for Mycotoxin-Producing Fungi
by
Carlos Bastidas-Caldes, David Vasco-Julio, Maria Huilca-Ibarra, Salomé Guerrero-Freire, Yanua Ledesma-Bravo and Jacobus H. de Waard
Microorganisms 2024, 12(5), 937; https://doi.org/10.3390/microorganisms12050937 - 05 May 2024
Abstract
Palm kernel cake (PKC), a byproduct of palm oil extraction, serves an important role in Ecuador’s animal feed industry. The emergence of yellow-orange fungal growth in PKC on some cattle farms in Ecuador sparked concerns within the cattle industry regarding a potential mycotoxin-producing
[...] Read more.
Palm kernel cake (PKC), a byproduct of palm oil extraction, serves an important role in Ecuador’s animal feed industry. The emergence of yellow-orange fungal growth in PKC on some cattle farms in Ecuador sparked concerns within the cattle industry regarding a potential mycotoxin-producing fungus on this substrate. Due to the limited availability of analytical chemistry techniques in Ecuador for mycotoxin detection, we chose to isolate and identify the fungus to determine its association with mycotoxin-producing genera. Through molecular identification via ITS region sequencing, we identified the yellow-orange fungus as the yeast Candida ethanolica. Furthermore, we isolated two other fungi—the yeast Pichia kudriavzevii, and the fungus Geotrichum candidum. Molecular identification confirmed that all three species are not classified as mycotoxin-producing fungi but in contrast, the literature indicates that all three have demonstrated antifungal activity against Aspergillus and Penicillium species, genera associated with mycotoxin production. This suggests their potential use in biocontrol to counter the colonization of harmful fungi. We discuss preventive measures against the fungal invasion of PKC and emphasize the importance of promptly identifying fungi on this substrate. Rapid recognition of mycotoxin-producing and pathogenic genera holds the promise of mitigating cattle intoxication and the dissemination of mycotoxins throughout the food chain.
Full article
(This article belongs to the Special Issue Food Microbiota and Food Safety)
►▼
Show Figures

Figure 1
Open AccessArticle
Prevalence and Risk Factors for Colonization by Multidrug-Resistant Microorganisms among Long-Term Travelers and Recently Arrived Migrants
by
Víctor Monsálvez, Paula Bierge, María Luisa Machado, Oscar Q. Pich, Elisa Nuez-Zaragoza, Carme Roca, Ana I. Jiménez-Lozano, Ángela Martínez-Perez, Aina Gomila-Grange, Isabel Vera-Garcia, Ana Requena-Méndez, Silvia Capilla and Oriol Gasch
Microorganisms 2024, 12(5), 936; https://doi.org/10.3390/microorganisms12050936 - 04 May 2024
Abstract
Multidrug-resistant (MDR) bacteria have become one of the most important health problems. We aimed to assess whether international travel may facilitate their spread through the colonization of asymptomatic travelers. A cross-sectional study was conducted (November 2018 to February 2022). Pharyngeal and rectal swabs
[...] Read more.
Multidrug-resistant (MDR) bacteria have become one of the most important health problems. We aimed to assess whether international travel may facilitate their spread through the colonization of asymptomatic travelers. A cross-sectional study was conducted (November 2018 to February 2022). Pharyngeal and rectal swabs were obtained from long-term travelers and recently arrived migrants from non-European countries, and an epidemiological survey was performed. Colonization by Gram-negative bacteria and methicillin-resistant Staphylococcus aureus (MRSA) was determined by chromogenic media and MALDI-TOF-MS. Resistance mechanisms were determined by the biochip-based molecular biology technique. Risk factors for colonization were assessed by logistic regression. In total, 122 participants were included: 59 (48.4%) recently arrived migrants and 63 (51.6%) long-term travelers. After their trip, 14 (11.5%) participants—5 (8.5%) migrants and 9 (14.3%) travelers—had rectal colonization by one MDR bacterium. Escherichia coli carrying the extended-spectrum beta-lactamase (ESBL) CTX-M-15 was the most frequent. No participants were colonized by MRSA or carbapenemase-producing Enterobacteriaceae. The only risk factor independently associated with MDR bacterial colonization was previous hospital attention [OR, 95% CI: 10.16 (2.06–50.06)]. The risk of colonization by MDR bacteria among recently arrived migrants and long-term travelers is similar in both groups and independently associated with previous hospital attention.
Full article
(This article belongs to the Section Antimicrobial Agents and Resistance)
Open AccessArticle
Improving Bacterial Metagenomic Research through Long-Read Sequencing
by
Noah Greenman, Sayf Al-Deen Hassouneh, Latifa S. Abdelli, Catherine Johnston and Taj Azarian
Microorganisms 2024, 12(5), 935; https://doi.org/10.3390/microorganisms12050935 - 04 May 2024
Abstract
Metagenomic sequencing analysis is central to investigating microbial communities in clinical and environmental studies. Short-read sequencing remains the primary approach for metagenomic research; however, long-read sequencing may offer advantages of improved metagenomic assembly and resolved taxonomic identification. To compare the relative performance for
[...] Read more.
Metagenomic sequencing analysis is central to investigating microbial communities in clinical and environmental studies. Short-read sequencing remains the primary approach for metagenomic research; however, long-read sequencing may offer advantages of improved metagenomic assembly and resolved taxonomic identification. To compare the relative performance for metagenomic studies, we simulated short- and long-read datasets using increasingly complex metagenomes comprising 10, 20, and 50 microbial taxa. Additionally, we used an empirical dataset of paired short- and long-read data generated from mouse fecal pellets to assess real-world performance. We compared metagenomic assembly quality, taxonomic classification, and metagenome-assembled genome (MAG) recovery rates. We show that long-read sequencing data significantly improve taxonomic classification and assembly quality. Metagenomic assemblies using simulated long reads were more complete and more contiguous with higher rates of MAG recovery. This resulted in more precise taxonomic classifications. Principal component analysis of empirical data demonstrated that sequencing technology affects compositional results as samples clustered by sequence type, not sample type. Overall, we highlight strengths of long-read metagenomic sequencing for microbiome studies, including improving the accuracy of classification and relative abundance estimates. These results will aid researchers when considering which sequencing approaches to use for metagenomic projects.
Full article
(This article belongs to the Section Microbiomes)
►▼
Show Figures

Figure 1
Open AccessArticle
A Highly Homogeneous Airborne Fungal Community around a Copper Open Pit Mine Reveals the Poor Contribution Made by the Local Aerosolization of Particles
by
Sebastián Fuentes-Alburquenque, Victoria Olivencia Suez, Omayra Aguilera, Blanca Águila, Luis Rojas Araya and Dinka Mandakovic
Microorganisms 2024, 12(5), 934; https://doi.org/10.3390/microorganisms12050934 - 04 May 2024
Abstract
Fungi are ubiquitous and metabolically versatile. Their dispersion has important scientific, environmental, health, and economic implications. They can be dispersed through the air by the aerosolization of near surfaces or transported from distant sources. Here, we tested the contribution of local (scale of
[...] Read more.
Fungi are ubiquitous and metabolically versatile. Their dispersion has important scientific, environmental, health, and economic implications. They can be dispersed through the air by the aerosolization of near surfaces or transported from distant sources. Here, we tested the contribution of local (scale of meters) versus regional (kilometers) sources by analyzing an airborne fungal community by ITS sequencing around a copper mine in the North of Chile. The mine was the regional source, whereas the soil and vegetal detritus were the local sources at each point. The airborne community was highly homogeneous at ca. 2000 km2, impeding the detection of regional or local contributions. Ascomycota was the dominant phylum in the three communities. Soil and vegetal detritus communities had lower alpha diversity, but some taxa had abundance patterns related to the distance from the mine and altitude. On the contrary, the air was compositionally even and unrelated to environmental or spatial factors, except for altitude. The presence of plant pathogens in the air suggests that other distant sources contribute to this region’s airborne fungal community and reinforces the complexity of tracking the sources of air microbial communities in a real world where several natural and human activities coexist.
Full article
(This article belongs to the Special Issue Airborne Microbial Communities)
►▼
Show Figures
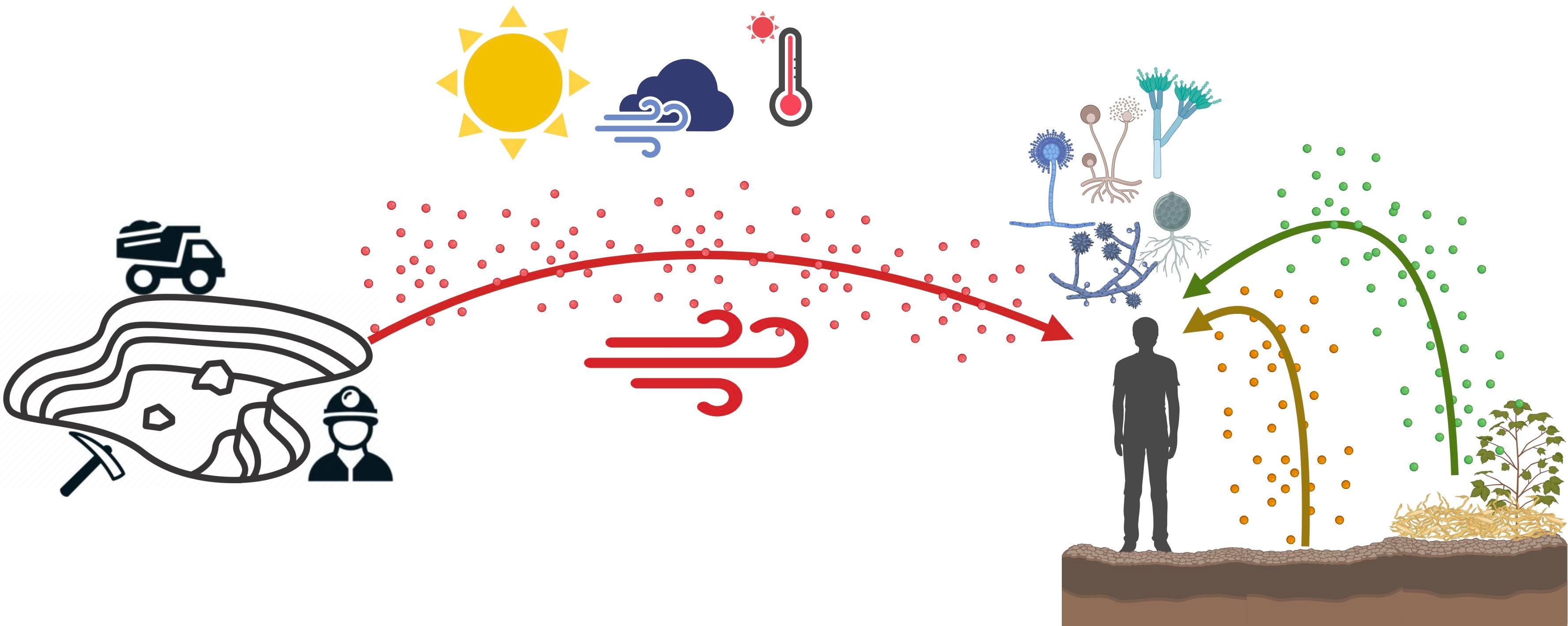
Graphical abstract
Open AccessArticle
Multiple Cofactor Engineering Strategies to Enhance Pyridoxine Production in Escherichia coli
by
Lijuan Wu, Jinlong Li, Yahui Zhang, Zhizhong Tian, Zhaoxia Jin, Linxia Liu and Dawei Zhang
Microorganisms 2024, 12(5), 933; https://doi.org/10.3390/microorganisms12050933 - 03 May 2024
Abstract
Pyridoxine, also known as vitamin B6, is an essential cofactor in numerous cellular processes. Its importance in various applications has led to a growing interest in optimizing its production through microbial biosynthesis. However, an imbalance in the net production of NADH
[...] Read more.
Pyridoxine, also known as vitamin B6, is an essential cofactor in numerous cellular processes. Its importance in various applications has led to a growing interest in optimizing its production through microbial biosynthesis. However, an imbalance in the net production of NADH disrupts intracellular cofactor levels, thereby limiting the efficient synthesis of pyridoxine. In our study, we focused on multiple cofactor engineering strategies, including the enzyme design involved in NAD+-dependent enzymes and NAD+ regeneration through the introduction of heterologous NADH oxidase (Nox) coupled with the reduction in NADH production during glycolysis. Finally, the engineered E. coli achieved a pyridoxine titer of 676 mg/L in a shake flask within 48 h by enhancing the driving force. Overall, the multiple cofactor engineering strategies utilized in this study serve as a reference for enhancing the efficient biosynthesis of other target products.
Full article
(This article belongs to the Section Microbial Biotechnology)
Open AccessArticle
Designing a Multiplex PCR-xMAP Assay for the Detection and Differentiation of African Horse Sickness Virus, Serotypes 1–9
by
Martin Ashby, Rebecca Moore, Simon King, Kerry Newbrook, John Flannery and Carrie Batten
Microorganisms 2024, 12(5), 932; https://doi.org/10.3390/microorganisms12050932 - 03 May 2024
Abstract
African horse sickness is a severe and often fatal disease affecting all species of equids. The aetiological agent, African horse sickness virus (AHSV), can be differentiated into nine serotypes. The identification of AHSV serotypes is vital for disease management, as this can influence
[...] Read more.
African horse sickness is a severe and often fatal disease affecting all species of equids. The aetiological agent, African horse sickness virus (AHSV), can be differentiated into nine serotypes. The identification of AHSV serotypes is vital for disease management, as this can influence vaccine selection and help trace disease incursion routes. In this study, we report the development and optimisation of a novel, molecular-based assay that utilises multiplex PCR and microsphere-based technology to expedite detection and differentiation of multiple AHSV serotypes in one assay. We demonstrated the ability of this assay to identify all nine AHSV serotypes, with detection limits ranging from 1 to 277 genome copies/µL depending on the AHSV serotype. An evaluation of diagnostic sensitivity and specificity revealed a sensitivity of 88% and specificity of 100%. This method can serotype up to 42 samples per run and can be completed in approximately 4–6 h. It provides a powerful tool to enhance the rapidity and efficiency of AHSV serotype detection, thereby facilitating the generation of epidemiological data that can help understand and control the incidence of AHSV worldwide.
Full article
(This article belongs to the Section Virology)
►▼
Show Figures

Figure 1
Open AccessArticle
Innovative Biomarkers for Obesity and Type 1 Diabetes Based on Bifidobacterium and Metabolomic Profiling
by
Angelica Nobili, Marco Pane, Mariya Skvortsova, Meryam Ben Salem, Stephan Morgenthaler, Emily Jamieson, Marina Di Stefano, Eirini Bathrellou, Eirini Mamalaki, Victoria Ramos-Garcia, Julia Kuligowski, Miltiadis Vasileiadis, Panagiotis Georgiadis, Marika Falcone and Paulo Refinetti
Microorganisms 2024, 12(5), 931; https://doi.org/10.3390/microorganisms12050931 - 03 May 2024
Abstract
The role of Bifidobacterium species and microbial metabolites such as short-chain fatty acids (SCFAs) and human milk oligosaccharides in controlling intestinal inflammation and the pathogenesis of obesity and type 1 diabetes (T1D) has been largely studied in recent years. This paper discusses the
[...] Read more.
The role of Bifidobacterium species and microbial metabolites such as short-chain fatty acids (SCFAs) and human milk oligosaccharides in controlling intestinal inflammation and the pathogenesis of obesity and type 1 diabetes (T1D) has been largely studied in recent years. This paper discusses the discovery of signature biomarkers for obesity and T1D based on data from a novel test for profiling several Bifidobacterium species, combined with metabolomic analysis. Through the NUTRISHIELD clinical study, a total of 98 children were recruited: 40 healthy controls, 40 type 1 diabetics, and 18 obese children. Bifidobacterium profiles were assessed in stool samples through an innovative test allowing high taxonomic resolution and precise quantification, while SCFAs and branched amino acids were measured in urine samples through gas chromatography–mass spectrometry (GC-MS). KIDMED questionnaires were used to evaluate the children’s dietary habits and correlate them with the Bifidobacterium and metabolomic profiles. We found that B. longum subs. infantis and B. breve were higher in individuals with obesity, while B. bifidum and B. longum subs. longum were lower compared to healthy individuals. In individuals with T1D, alterations were found at the metabolic level, with an overall increase in the level of the most measured metabolites. The high taxonomic resolution of the Bifidobacterium test used meant strong correlations between the concentrations of valine and isoleucine, and the relative abundance of some Bifidobacterium species such as B. longum subs. infantis, B. breve, and B. bifidum could be observed.
Full article
(This article belongs to the Special Issue Novel Strategies in the Study of the Human Gut Microbiota 2.0)
Open AccessArticle
Role of Volatile Organic Compounds Produced by Kosakonia cowanii Cp1 during Competitive Colonization Interaction against Pectobacterium aroidearum SM2
by
Mayra Paola Mena Navarro, Merle Ariadna Espinosa Bernal, Adriana Eunice Martinez-Avila, Leonela Sofia Aponte Pineda, Luis Alberto Montes Flores, Carlos Daniel Chan Ku, Yoali Fernanda Hernández Gómez, Jacqueline González Espinosa, Juan Ramiro Pacheco Aguilar, Miguel Ángel Ramos López, Jackeline Lizzeta Arvizu Gómez, Carlos Saldaña Gutierrez, José Alberto Rodríguez Morales, Aldo Amaro Reyes, José Luis Hernández Flores and Juan Campos Guillén
Microorganisms 2024, 12(5), 930; https://doi.org/10.3390/microorganisms12050930 - 03 May 2024
Abstract
The competitive colonization of bacteria on similar ecological niches has a significant impact during their establishment. The synthesis speeds of different chemical classes of molecules during early competitive colonization can reduce the number of competitors through metabolic effects. In this work, we demonstrate
[...] Read more.
The competitive colonization of bacteria on similar ecological niches has a significant impact during their establishment. The synthesis speeds of different chemical classes of molecules during early competitive colonization can reduce the number of competitors through metabolic effects. In this work, we demonstrate for the first time that Kosakonia cowanii Cp1 previously isolated from the seeds of Capsicum pubescens R. P. produced volatile organic compounds (VOCs) during competitive colonization against Pectobacterium aroidearum SM2, affecting soft rot symptoms in serrano chili (Capsicum annuum L.). The pathogen P. aroidearum SM2 was isolated from the fruits of C. annuum var. Serrano with soft rot symptoms. The genome of the SM2 strain carries a 5,037,920 bp chromosome with 51.46% G + C content and 4925 predicted protein-coding genes. It presents 12 genes encoding plant-cell-wall-degrading enzymes (PCDEWs), 139 genes involved in five types of secretion systems, and 16 genes related to invasion motility. Pathogenic essays showed soft rot symptoms in the fruits of C. annuum L., Solanum lycopersicum, and Physalis philadelphica and the tubers of Solanum tuberosum. During the growth phases of K. cowanii Cp1, a mix of VOCs was identified by means of HS-SPME-GC-MS. Of these compounds, 2,5-dimethyl-pyrazine showed bactericidal effects and synergy with acetoin during the competitive colonization of K. cowanii Cp1 to completely reduce soft rot symptoms. This work provides novel evidence grounding a better understanding of bacterial interactions during competitive colonization on plant tissue, where VOC synthesis is essential and has a high potential capacity to control pathogenic microorganisms in agricultural systems.
Full article
(This article belongs to the Special Issue Advanced Research on Biological Control of Plant Disease or Microbial Interactions 2.0)
►▼
Show Figures

Figure 1
Open AccessArticle
Plasmid-Borne Biosynthetic Gene Clusters within a Permanently Stratified Marine Water Column
by
Paraskevi Mara, David Geller-McGrath, Elizabeth Suter, Gordon T. Taylor, Maria G. Pachiadaki and Virginia P. Edgcomb
Microorganisms 2024, 12(5), 929; https://doi.org/10.3390/microorganisms12050929 - 02 May 2024
Abstract
Plasmids are mobile genetic elements known to carry secondary metabolic genes that affect the fitness and survival of microbes in the environment. Well-studied cases of plasmid-encoded secondary metabolic genes in marine habitats include toxin/antitoxin and antibiotic biosynthesis/resistance genes. Here, we examine metagenome-assembled genomes
[...] Read more.
Plasmids are mobile genetic elements known to carry secondary metabolic genes that affect the fitness and survival of microbes in the environment. Well-studied cases of plasmid-encoded secondary metabolic genes in marine habitats include toxin/antitoxin and antibiotic biosynthesis/resistance genes. Here, we examine metagenome-assembled genomes (MAGs) from the permanently-stratified water column of the Cariaco Basin for integrated plasmids that encode biosynthetic gene clusters of secondary metabolites (smBGCs). We identify 16 plasmid-borne smBGCs in MAGs associated primarily with Planctomycetota and Pseudomonadota that encode terpene-synthesizing genes, and genes for production of ribosomal and non-ribosomal peptides. These identified genes encode for secondary metabolites that are mainly antimicrobial agents, and hence, their uptake via plasmids may increase the competitive advantage of those host taxa that acquire them. The ecological and evolutionary significance of smBGCs carried by prokaryotes in oxygen-depleted water columns is yet to be fully elucidated.
Full article
(This article belongs to the Special Issue Marine Microorganisms in a Changing Ocean: From Single Species to Community Responses)
►▼
Show Figures

Figure 1
Open AccessArticle
Influence of Compost Amendments on Soil and Human Gastrointestinal Bacterial Communities during a Single Gardening Season
by
Sihan Bu, Alyssa W. Beavers, Kameron Y. Sugino, Sarah F. Keller, Katherine Alaimo and Sarah S. Comstock
Microorganisms 2024, 12(5), 928; https://doi.org/10.3390/microorganisms12050928 - 01 May 2024
Abstract
To measure associations between gardening with different compost amendments and the human gut microbiota composition, gardeners (n = 25) were provided with one of three types of compost: chicken manure (CM), dairy manure and plant material (DMP), or plant-based (P). Stool samples were
[...] Read more.
To measure associations between gardening with different compost amendments and the human gut microbiota composition, gardeners (n = 25) were provided with one of three types of compost: chicken manure (CM), dairy manure and plant material (DMP), or plant-based (P). Stool samples were collected before gardening (T1), after compost amendment (T2), and at peak garden harvest (T3). Compost and soil samples were collected. DNA was extracted, 16S rRNA libraries were established, and libraries were sequenced by Illumina MiSeq. Sequences were processed using mothur, and data were analyzed in R software version 4.2.2. Fast expectation-maximization microbial source tracking analysis was used to determine stool bacteria sources. At T2/T3, the gut microbiotas of P participants had the lowest Shannon alpha diversity, which was also the trend at T1. In stool from T2, Ruminococcus 1 were less abundant in the microbiotas of those using P compost as compared to those using CM or DMP. At T2, Prevotella 9 had the highest abundance in the microbiotas of those using CM compost. In participants who used CM compost to amend their gardening plots, a larger proportion of the human stool bacteria were sourced from CM compared to soil. Soil exposure through gardening was associated with a small but detectable change in the gardeners’ gut microbiota composition. These results suggest that human interactions with soil through gardening could potentially impact health through alterations to the gut microbiota.
Full article
(This article belongs to the Section Gut Microbiota)
►▼
Show Figures
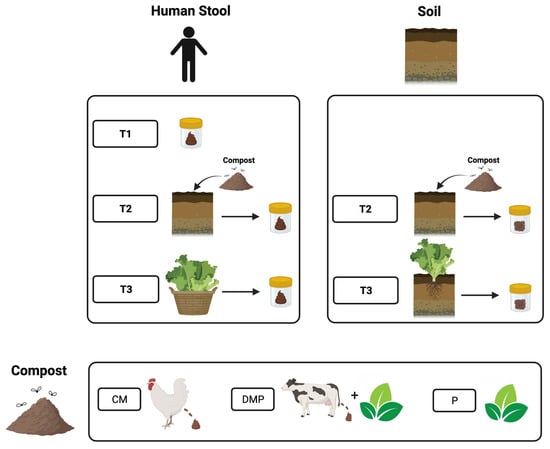
Figure 1
Open AccessReview
Candida auris Outbreaks: Current Status and Future Perspectives
by
Silvia De Gaetano, Angelina Midiri, Giuseppe Mancuso, Maria Giovanna Avola and Carmelo Biondo
Microorganisms 2024, 12(5), 927; https://doi.org/10.3390/microorganisms12050927 - 01 May 2024
Abstract
Candida auris has been identified by the World Health Organization (WHO) as a critical priority pathogen on its latest list of fungi. C. auris infections are reported in the bloodstream and less commonly in the cerebrospinal fluid and abdomen, with mortality rates that
[...] Read more.
Candida auris has been identified by the World Health Organization (WHO) as a critical priority pathogen on its latest list of fungi. C. auris infections are reported in the bloodstream and less commonly in the cerebrospinal fluid and abdomen, with mortality rates that range between 30% and 72%. However, no large-scale epidemiology studies have been reported until now. The diagnosis of C. auris infections can be challenging, particularly when employing conventional techniques. This can impede the early detection of outbreaks and the implementation of appropriate control measures. The yeast can easily spread between patients and in healthcare settings through contaminated environments or equipment, where it can survive for extended periods. Therefore, it would be desirable to screen patients for C. auris colonisation. This would allow facilities to identify patients with the disease and take appropriate prevention and control measures. It is frequently unsusceptible to drugs, with varying patterns of resistance observed among clades and geographical regions. This review provides updates on C. auris, including epidemiology, clinical characteristics, genomic analysis, evolution, colonisation, infection, identification, resistance profiles, therapeutic options, prevention, and control.
Full article
(This article belongs to the Special Issue Host Versus Pathogen: Candida Infections, Immune Response and Therapy Perspectives)
Open AccessBrief Report
Malian Children’s Core Gut Mycobiome
by
Abdourahim Abdillah, Aly Kodio and Stéphane Ranque
Microorganisms 2024, 12(5), 926; https://doi.org/10.3390/microorganisms12050926 - 01 May 2024
Abstract
Because data on the fungal gut community structure of African children are scarce, we aimed to describe it by reanalysing rRNA ITS1 and ITS2 metabarcoding data from a study designed to assess the influence of microbiota in malaria susceptibility in Malian children from
[...] Read more.
Because data on the fungal gut community structure of African children are scarce, we aimed to describe it by reanalysing rRNA ITS1 and ITS2 metabarcoding data from a study designed to assess the influence of microbiota in malaria susceptibility in Malian children from the Dogon country. More specifically, we aimed to establish the core gut mycobiome and compare the gut fungal community structure of breastfed children, aged 0–2 years, with other age groups. Briefly, DNA was extracted from 296 children’s stool samples. Both rRNA ITS1 and ITS2 genomic barcodes were amplified and subjected to Illumina MiSeq sequencing. The ITS2 barcode generated 1,975,320 reads and 532 operational taxonomic units (OTUs), while the ITS1 barcode generated 647,816 reads and 532 OTUs. The alpha diversity was significantly higher by using the ITS1 compared to the ITS2 barcode (p < 0.05); but, regardless of the ITS barcode, we found no significant difference between breastfed children, aged 0–2 years, compared to the other age groups. The core gut mycobiome of the Malian children included Saccharomyces cerevisiae, Candida albicans, Pichia kudriavzevii, Malassezia restricta, Candida tropicalis and Aspergillus section Aspergillus, which were present in at least 50% of the 296 children. Further studies in other African countries are warranted to reach a global view of African children’s core gut mycobiome.
Full article
(This article belongs to the Special Issue Gut Microbiome and Children’s Health)
►▼
Show Figures
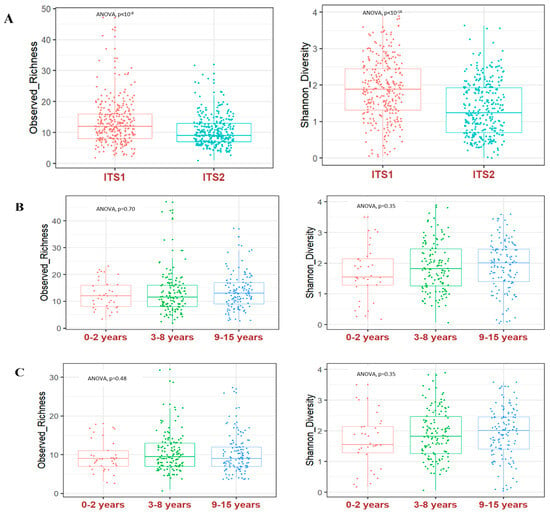
Figure 1
Open AccessArticle
Baseline Skin Microbiota of the Leatherback Sea Turtle
by
Samantha G. Kuschke, Jeanette Wyneken and Debra Miller
Microorganisms 2024, 12(5), 925; https://doi.org/10.3390/microorganisms12050925 - 01 May 2024
Abstract
The integumentary system of the leatherback sea turtle (Dermochelys coriacea) is the most visible and defining difference of the species, with its smooth and waxy carapace and finely scaled skin, distinguishing it from the other six sea turtle species. The skin
[...] Read more.
The integumentary system of the leatherback sea turtle (Dermochelys coriacea) is the most visible and defining difference of the species, with its smooth and waxy carapace and finely scaled skin, distinguishing it from the other six sea turtle species. The skin is the body’s largest organ and serves as a primary defense against the outside world and is thus essential to health. To date, we have begun to understand that the microorganisms located on the skin aid in these functions. However, many host–microbial interactions are not yet fully defined or understood. Prior to uncovering these crucial host–microbial interactions, we must first understand the communities of microorganisms present and how they differ through life-stage classes and across the body. Here, we present a comprehensive bacterial microbial profile on the skin of leatherbacks. Using next-generation sequencing (NGS), we identified the major groups of bacteria on the skin of neonates at emergence, neonates at 3–4 weeks of age (i.e., post-hatchlings), and nesting females. These data show that the predominant bacteria on the skin of the leatherback are different at each life-stage class sampled. This suggests that there is a shift in the microbial communities of the skin associated with life-stage class or even possibly age. We also found that different sample locations on the nesting female (i.e., carapace and front appendages = flipper) have significantly different communities of bacteria present. This is likely due to differences in the microhabitats of these anatomic locations and future studies should explore if this variation also holds true for neonates. These data define baseline skin microbiota on the leatherback and can serve as a foundation for additional work to broaden our understanding of the leatherbacks’ host–microbial interactions, the impacts of environmental changes or stressors over time, and even the pathogenicity of disease processes.
Full article
(This article belongs to the Special Issue Microbiota and Microbiomes in Plants, Animals and Environment: A One Heath Perspective)
►▼
Show Figures
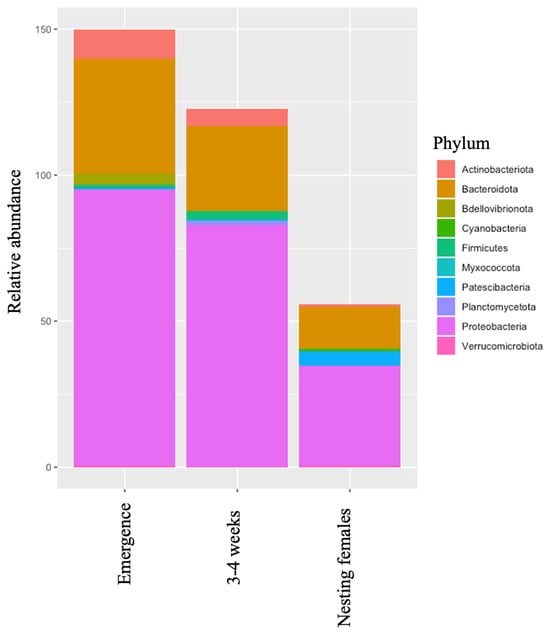
Figure 1
Open AccessReview
Management Strategies for Common Animal Bites in Pediatrics: A Narrative Review on the Latest Progress
by
Dragos Septelici, Giulia Carbone, Alessandro Cipri and Susanna Esposito
Microorganisms 2024, 12(5), 924; https://doi.org/10.3390/microorganisms12050924 - 01 May 2024
Abstract
Animal bites are a common reason for children to visit primary care and emergency departments. Dog bites are the most prevalent, followed by cat bites at 20–30%. Other animals such as bats, monkeys, snakes, and rats collectively contribute less than 1% of cases.
[...] Read more.
Animal bites are a common reason for children to visit primary care and emergency departments. Dog bites are the most prevalent, followed by cat bites at 20–30%. Other animals such as bats, monkeys, snakes, and rats collectively contribute less than 1% of cases. Hospitalization is necessary in only 4% of animal bite incidents. The main aim of this narrative review is to summarize the main protocols currently followed in pediatrics in cases involving the most common bites from different animal species. Analysis of the literature showed that the management of common animal bites in children presents a multifaceted challenge requiring a comprehensive understanding of the epidemiology, clinical presentation, and treatment modalities associated with each specific species. Effective wound management is paramount in reducing the risk of infection and promoting optimal healing outcomes. Additionally, tetanus vaccination status should be assessed and updated as necessary, and prophylactic antibiotics may be indicated in certain cases to prevent secondary infections. Furthermore, the role of rabies prophylaxis cannot be overstated, particularly in regions where rabies is endemic or following bites from high-risk animals. In addition to medical management, psychosocial support for both the child and their caregivers is integral to the overall care continuum. Future studies exploring the efficacy of novel treatment modalities, such as topical antimicrobial agents or advanced wound dressings, may offer new insights into optimizing wound healing and reducing the risk of complications.
Full article
(This article belongs to the Special Issue Emerging Infectious Diseases in Humans and Animals)
Open AccessArticle
Antiparasitic Activity of Enterocin M and Durancin-like from Beneficial Enterococci in Mice Experimentally Infected with Trichinella spiralis
by
Miroslava Petrová, Zuzana Hurníková, Andrea Lauková and Emília Dvorožňáková
Microorganisms 2024, 12(5), 923; https://doi.org/10.3390/microorganisms12050923 - 01 May 2024
Abstract
Beneficial/probiotic strains protect the host from pathogens by competitive displacement and production of antibacterial substances, i.e., bacteriocins. The antiparasitic potential of bacteriocins/enterocins and their producing strains in experimental murine trichinellosis were tested as a new therapeutic strategy. Enterocin M and Durancin-like and their
[...] Read more.
Beneficial/probiotic strains protect the host from pathogens by competitive displacement and production of antibacterial substances, i.e., bacteriocins. The antiparasitic potential of bacteriocins/enterocins and their producing strains in experimental murine trichinellosis were tested as a new therapeutic strategy. Enterocin M and Durancin-like and their producers Enterococcus faecium CCM8558 and Enterococcus durans ED26E/7 were administered daily to mice that were challenged with Trichinella spiralis. Our study confirmed the antiparasitic effect of enterocins/enterococci, which reduced the number of adults in the intestine (Enterocin M—43.8%, E. faecium CCM8558—54.5%, Durancin-like—16.4%, E. durans ED26E/7—35.7%), suppressed the Trichinella reproductive capacity ex vivo (Enterocin M—61%, E. faecium CCM8558—74%, Durancin-like—38%, E. durans ED26E/7—66%), and reduced the number of muscle larvae (Enterocin M—39.6%, E. faecium CCM8558—55.7%, Durancin-like—15%, E. durans ED26E/7—36.3%). The direct effect of enterocins on Trichinella fecundity was documented by an in vitro test in which Durancin-like showed a comparable reducing effect to Enterocin M (40–60%) in contrast to the ex vivo test. The reducing activity of T.spiralis infection induced by Enterocin M was comparable to its strain E. faecium CCM8558; Durancin-like showed lower antiparasitic activity than its producer E. durans ED26E/7.
Full article
(This article belongs to the Special Issue New Advances in Functional Ingredients’ Development to Improve Physiological and Functional Properties of Probiotics)
►▼
Show Figures

Figure 1

Journal Menu
► ▼ Journal Menu-
- Microorganisms Home
- Aims & Scope
- Editorial Board
- Reviewer Board
- Topical Advisory Panel
- Instructions for Authors
- Special Issues
- Topics
- Sections & Collections
- Article Processing Charge
- Indexing & Archiving
- Editor’s Choice Articles
- Most Cited & Viewed
- Journal Statistics
- Journal History
- Journal Awards
- Society Collaborations
- Editorial Office
Journal Browser
► ▼ Journal BrowserHighly Accessed Articles
Latest Books
E-Mail Alert
News
Topics
Topic in
Aquaculture Journal, Fishes, Microorganisms, Water
Women in Aquaculture Research
Topic Editors: Camino Ordás, Patrícia Díaz-RosalesDeadline: 30 June 2024
Topic in
Agronomy, Environments, Microorganisms, Pollutants, Sustainability, Water
Soil and Water Pollution Process and Remediation Technologies, 2nd Volume
Topic Editors: Hongbiao Cui, Ru Wang, Yu Shi, Haiying Lu, Lin ChenDeadline: 15 July 2024
Topic in
Antibiotics, Antioxidants, JoF, Microbiology Research, Microorganisms
Redox in Microorganisms, 2nd Edition
Topic Editors: Michal Letek, Volker BehrendsDeadline: 31 July 2024
Topic in
Molecules, Pharmaceutics, Antibiotics, Microorganisms, Biomolecules, Marine Drugs, Polymers, IJMS
Antimicrobial Agents and Nanomaterials
Topic Editors: Sandra Pinto, Vasco D. B. BonifácioDeadline: 30 September 2024

Conferences
Special Issues
Special Issue in
Microorganisms
Human Skin Microbiota 2.0
Guest Editors: Rolf Lood, Holger BrüggemannDeadline: 15 May 2024
Special Issue in
Microorganisms
The Urban Microbiome
Guest Editors: Haruo Suzuki, Soojin JangDeadline: 31 May 2024
Special Issue in
Microorganisms
Emerging Pathogens Causing Acute Hepatitis
Guest Editors: Sayed F. Abdelwahab, Ibrahim M. Sayed, Ahmed El-ShamyDeadline: 15 June 2024
Special Issue in
Microorganisms
Monkeypox—Current Knowledge and Future Perspectives
Guest Editors: Jakab Ferenc, Krisztián BányaiDeadline: 30 June 2024
Topical Collections
Topical Collection in
Microorganisms
Trends in Yeast Biochemistry and Biotechnology
Collection Editors: Seiji Shibasaki, Mitsuyoshi Ueda
Topical Collection in
Microorganisms
Epidemiology and Pathogenicity of Animal-Adapted Streptococci
Collection Editors: Peter Valentin-Weigand, Marcus Fulde
Topical Collection in
Microorganisms
Biodegradation and Environmental Microbiomes
Collection Editors: Shuangjiang Liu, Hongzhi Tang, Jiandong Jiang, Xiaolei Wu