3. Results
First, we sought to identify a toxic phenylalanine analog and work backward to design toxic phenylalanine precursors. Two promising toxic phenylalanine analogs were identified from the literature:
l-3-thienylalanine (L3T), consisting of phenylalanine with the benzene ring substituted for thiophene, and fluorophenylalanine (FPA), consisting of phenylalanine with a fluorine attached to the benzene ring (
Figure 2,
Table 1). A 2.6 mg L
−1 concentration of L-3-thienylalanine has been shown to inhibit growth of
S. cerevisiae by 50% [
9]. First, the toxicity of L3-thienylalanine was tested by plating wild-type strains S288c and AW077 to medium with L3-thienylalanine at 0 or 300 mg L
−1 and determining the relative viability. L3T killed all AW077 cells and nearly all of S288c cells (
Figure 3a). Next the toxicity of FPA was tested by plating strains S288c and AW077 onto medium containing FPA at 0 or 300 mg L
−1 and determining the relative viability (
Figure 3b). FPA at both of these concentrations killed almost all cells of S288c and AW077. Thus, we reconfirmed that L3T and FPA are toxic to
S. cerevisiae.Next, we sought to design a molecule that may be enzymatically converted to a toxic phenylalanine analog. By identifying a toxic phenylalanine precursor, we could use this compound to select for strains that have mutations in the phenylalanine biosynthetic pathway. We considered several precursors of phenylalanine; chorismate (this precursor is used to synthesize not only phenylalanine, but also tryptophan, tyrosine, folate, and ubiquinone). Thus, a strain with a chorismate deletion would have numerous other auxotrophies besides phenylalanine. Next, prephenate—however, this is not a good candidate for this particular intervention as adding fluorine or sulfur on the cyclohexadiene ring could interfere with transformation to a benzene ring. Additionally, there seems to be at least two mechanisms to convert prephenate to phenylalanine, as we found that deletion of
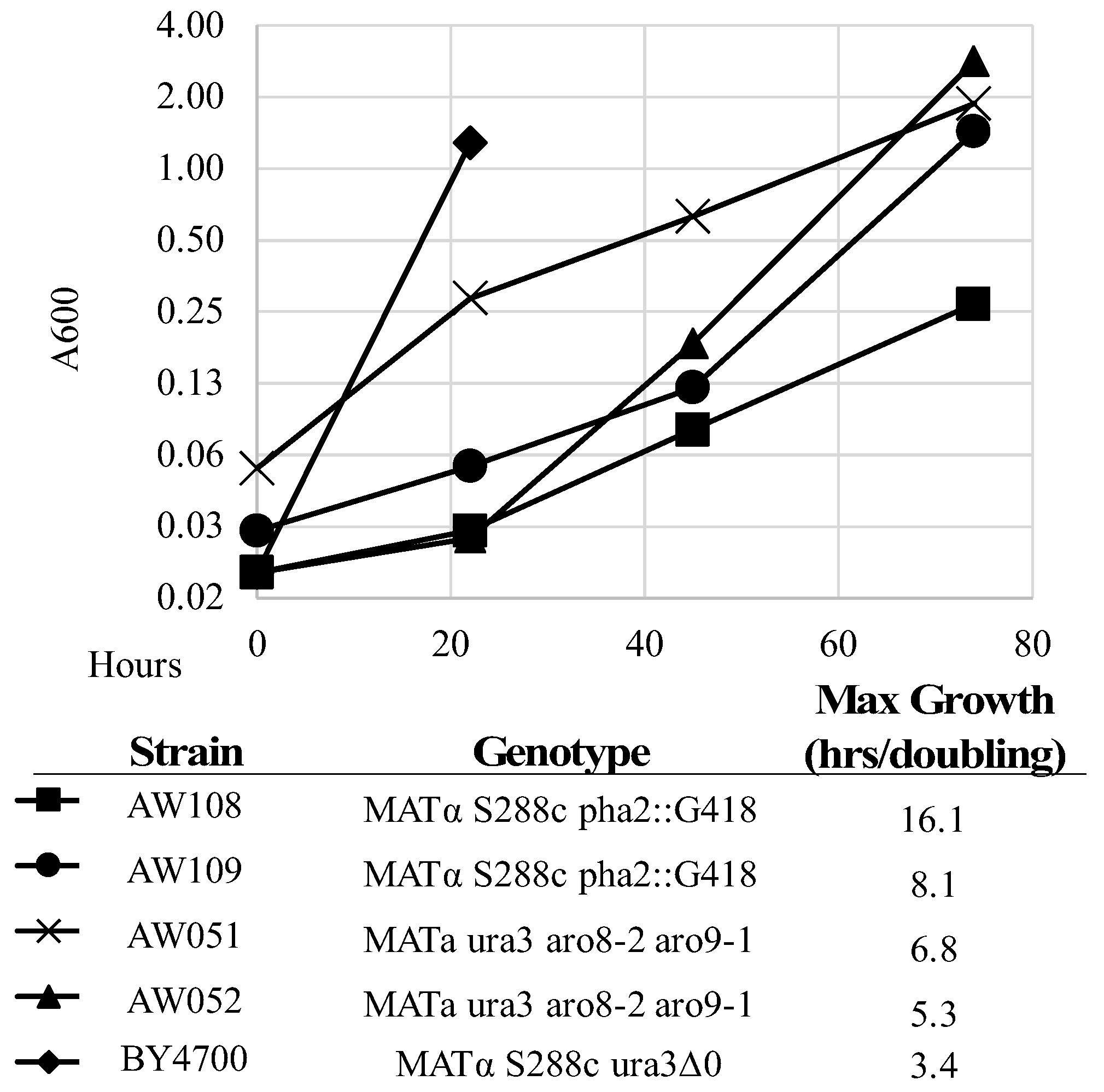
PHA2 does not confer complete phenylalanine requirement. To test this, strains lacking
PHA2 were inoculated alongside a strain with functioning
PHA2 in minimal medium with or without phenylalanine added (
Figure 4,
Table 3). The A600 was measured over four days, and maximum growth rate (lowest doubling time) was calculated. Strains AW108 and AW109 grew much better with phenylalanine added (
Table 3), showing the dependency on phenylalanine; however, they still grew over time. This is in contrast to previously described observations of strains lacking
PHA2. However, it is likely that the extremely slow growth rate explains this conclusion (
Figure 4) [
10]. The next candidate is phenylpyruvate, which does not require modification of its benzene ring prior to becoming phenylalanine, and it is the substrate for only two essentials molecules: phenylalanine and tyrosine. Use of phenylpyruvate has one drawback: there are two primary enzymes that can complete the transamination reaction converting phenylpyruvate to phenylalanine (
ARO8 and
ARO9) and several others that are predicted to be capable of performing this reaction (Bat1, Bat2, His5) [
11].
This presents a problem for selection, as strains would need to have deletions in two genes, rather than just one, in the case of uracil (URA3), lysine (LYS2), or arginine (CAN1) [
6,
12,
13]. To test if
ARO8 and
ARO9 were required for phenylalanine production, strains lacking
ARO8 and
ARO9 (AW051, AW052) were constructed and inoculated alongside a wild-type strain in minimal medium with or without phenylalanine added (
Figure 4,
Table 3).
The A600 was measured over four days, and maximum growth rate (lowest doubling time) was calculated. Strains AW051 and AW052 grew much better with phenylalanine added (
Table 3), showing their dependency on phenylalanine. However, they still grew over time. These strains do grow faster than the
PHA2 mutants but are not completely rescued solely by phenylalanine, as they also require tyrosine, hence the lower A600 on the second day of growth with phenylalanine added.
Furthermore, to verify there were no residual nutrients in the minimal medium, strains with various individual auxotrophies were inoculated to minimal medium and were found to not grow (
Table 4).
Nonetheless, multiple enzymes producing phenylalanine could be viewed as a boon, as cells are less likely to develop spontaneous resistance to a toxic phenylpyruvate analog, as they would have to delete multiple processing enzymes. Thus we set out to design phenylpyruvate-like derivatives that may be catalytically converted into toxic phenylalanine analogs so that these can be used for counter-selection of phenylalanine auxotrophies.
The first two phenylpyruvate-like derivatives of
l-3-thienylalanine tested were 3-thiolproprionate (3TP) and 2-thienylpyruvate (2TP). Wild-type yeast strains S288c and AW077 were pinned to plates containing these analogs at 500 mg L
−1 and 900 mg L
−1, respectively, and relative viability was calculated (
Figure 3a). Despite higher concentrations of the phenylpyruvate derivatives compared to L3T, S288c and AW077 strains exhibited no decrease in viability on 3TP and 2TP containing medium. Thus, while the phenylalanine analog was toxic to these strains, the phenylpyruvate analogs were not. Differences in toxicity could be due to adsorption to the cell wall, decreased uptake, or incompatibility with enzymes.
The toxicity of the FPA derivative fluorophenylpyruvate (FPP) was next examined. Strains AW077 and S288c were serially diluted in water, then pinned to plates containing 0, 300 mg L
−1, or 500 mg L
−1 FPP and relative viability was calculated (
Figure 3b). At 300 mg L
−1 FPP strain S288c showed mild growth inhibition but no lethality, while strain AW077 was more growth-inhibited. At 500 mg L
−1, no growth was detected from S288c and AW077 strains (
Figure 3b). These results indicate that the phenylpyruvate analog FPP is toxic to S288c and AW077 strains.
Phenylpyruvate is converted to phenylalanine primarily by the proteins Aro8p and Aro9p. However, there are other proteins that can catalyze this reaction. To test if the mechanism of FPP toxicity was due to its conversion to FPA, the toxicity of FPP was tested on strains lacking genes
ARO8 and
ARO9 (
Figure 3b). Strains lacking
ARO8 and
ARO9 (AW051, AW052) are able to grow on this medium, showing approximately equal growth on YPD. However, these strains are hypersensitive to FPA, which may reflect the cells increased import of phenylalanine, due to a phenylalanine auxotrophy (
Figure 3c). This result suggests that
ARO8 and
ARO9 are primarily responsible for affecting the toxicity of FPP.
For FPP to be of use as an auxotrophic marker, the natural resistance for it should be low. For example, the frequency of spontaneous resistance to 5-fluoroanthranilic acid or 5-fluoroorotic acid, for selection of tryptophan or uracil auxotrophs, respectively, was 6 × 10
−7 [
5]. To test the natural resistance frequency of S288c and AW077 strains to FPP, we plated 5 × 10
6 cells on plates containing 500 mg L
−1 FPP. We found no resistant cells grew on this medium, indicating natural resistance of wild-type strains to FPP is below a frequency of 2 × 10
−7. As a comparison to this, we also plated these strains to medium containing 500 mg L
−1 FPA. Strains S288c and AW077 formed 240 and 86 colonies giving a frequency of resistance of 3 × 10
−5 and 5 × 10
−5, respectively. This indicates that the natural frequency of resistance to FPP is quite low, while that of FPA is higher. This indicates that resistance to FPP toxicity requires either more mutations or more specific mutations, as compared to FPA, and that perhaps these molecules do not induce toxicity via the same mechanism.
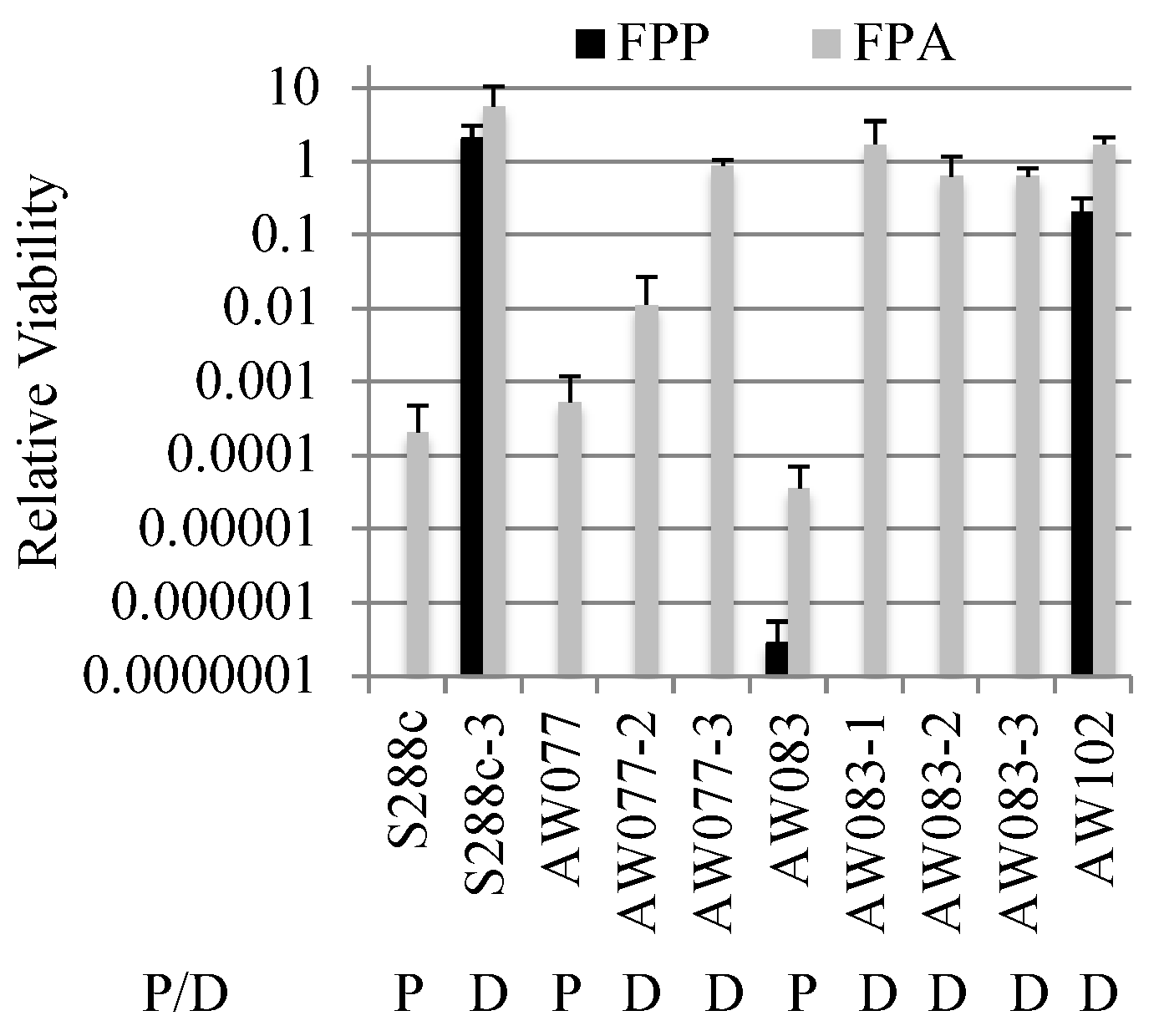
To further investigate the difference between FPP and FPA resistance, strains that were FPA resistant were tested for FPP resistance. Isolates that were resistant to FPA were plated onto medium with or without FPP and relative viability was calculated (
Figure 5). Three isolates derived from strain AW083 and one from AW077 showed resistance to FPA, but no resistance to FPP. This indicates that FPA resistance is not sufficient for FPP resistance. However, an isolate from a wild-type yeast wine strain (AW102) and an S288c mutant isolate (S288c-3) were recovered that showed resistance to FPP and FPA. When the strain S288c-3 was mated to a non-resistant strain (AW081) and sporulated, the resulting haploid segregants generally showed a pattern of 1:3 resistance to FPP, while resistance to FPA showed a pattern of 2:2 (
Table 5).
A predominance of 1:3 segregation (tetratype) indicates that two genes are segregating independently, following the pattern of 1:1:4 PD:NPD:TT, while a 2:2 pattern indicates a single gene. All haploid segregants that were resistant to FPP were also resistant to FPA. The haploid segregants that were only resistant to FPA generally showed slow growth on FPA. One interpretation of these results is that detoxification of FPP requires the activity of two genes, either of which will suffice to detoxify FPA.
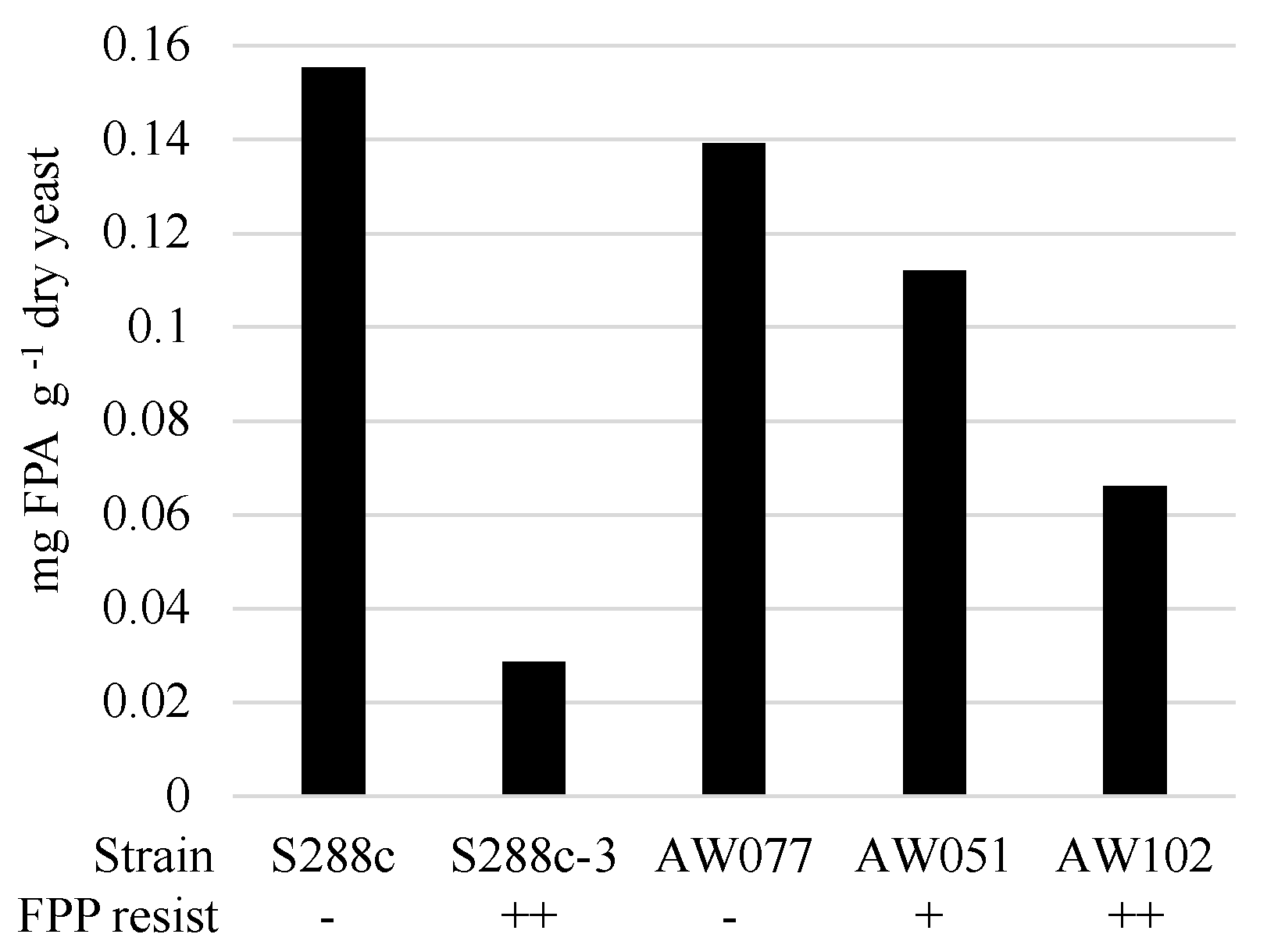
To determine if cells converted FPP into FPA or some other compound, we evaluated the presence of FPA in cells treated with FPP. Strains were inoculated to medium containing FPP and allowed to incubate for > 12 h, enzymatically digested to recover individual amino acids, and then tested for the presence of FPA using LC-MS/MS (
Figure 6). Strains sensitive to FPP, AW077, and S288c were observed to accumulate the highest amounts of FPA, while the less sensitive strain AW051 accumulated less. The modest decrease in FPP conversion of AW051 is not too surprising, as there are other enzymes capable of converting phenylpyruvate to phenylalanine (Bat1, Bat2, His5), and this strain is capable of growth without phenylalanine supplementation. Strains S288c-3 and AW102, both of which are resistant to FPP, had the lowest levels of FPA. This suggests that their mechanism of resistance involves decreased conversion of FPP to FPA. Thus cells can convert FPP into FPA, and this may be the cause of some of the toxicity of FPP.
For FPP to be an effective selective agent, it should exert its toxic effect amongst diverse genre of yeast. To test the evolutionary conservation of FPP toxicity, strains from diverse species such as
Candida tropicalis, Candida albicans, Saccharomyces bayanus, and
Saccharomyces paradoxus were exposed to 0 or 500 mg L
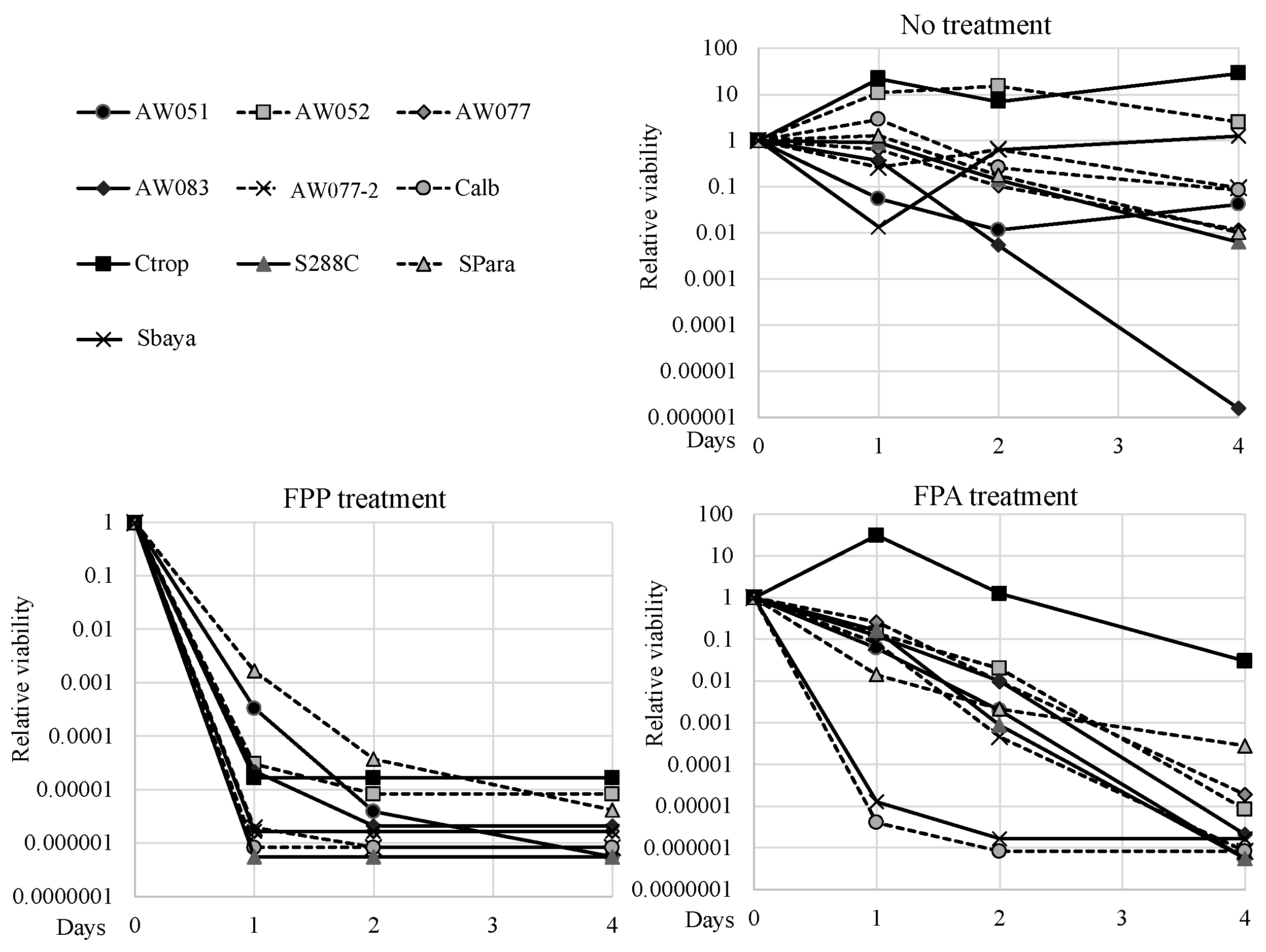
−1 FPP or FPA and their viability was measured over several days, and relative viabilities were calculated (
Figure 7). To test these strains, the FPP and FPA sensitivity test was performed by inoculating cells into media with said compounds and evaluating culture viability using the Tadpoling method [
8]. All the strains tested retained viability or slightly increased in cell number when inoculated into media without FPP or FPA, except for AW083. When inoculated to media containing FPA,
C. tropicalis exhibited high resistance while most of the other strains lost viability over the course of four days. Both
C. albicans and
S. bayanus exhibited enhanced sensitivity to FPA, as these lost almost all culture viability within one day. Surprisingly, in media containing FPP, all strains showed greater sensitivity than in FPA and lost at least four orders of magnitude culture viability within two days. This result would suggest that FPP may be transported more effectively than FPA, or that FPP can exert its toxicity through a mechanism distinct from that of FPA.
S.paradoxus and AW051 (
aro8-2 aro9-1) showed the least sensitivity to FPP of all the strains tested, but even these cultures succumbed after four days. This data shows that FPP can act through diverse yeast genre to exert its toxic effect and may be a more effective fungicide than FPA.
One potential mechanism of FPP toxicity is to affect the mitochondria. Phenylpyruvate acts as an inhibitor of pyruvate transport into the mitochondria where pyruvate would normally be decarboxylated to acetate [
14]. Indeed, accumulation of phenylpyruvate in phenylketonuria patients becomes toxic as it may disrupt oxidative metabolism by preventing normal pyruvate movement into the mitochondria. To test if phenylpyruvate alone may exert a toxic effect on yeast, exponential phase cells were inoculated to 500 mg L
−1 phenylpyruvate and culture viability was measured over several days using the Tadpoling method. The ratio of CFUs of cells exposed to phenylpyruvate with control cells was calculated over the course of several days (
Figure 8). There were no significant (
p < 0.05) deviations from the ideal 1 ratio for all strains tested, indicating that phenylpyruvate does not significantly affect culture viability. Nonetheless, there were notable deviations in decreased viability of S288c cells exposed to phenylpyruvate and an increase in viability of AW052 cells exposed to phenylpyruvate. Thus, it is likely that fluorine is required for fluorophenylalanine toxicity.
In
S. cerevisiae, one method of increasing the production of the valuable chemical phenylethanol is to select for resistance to FPA [
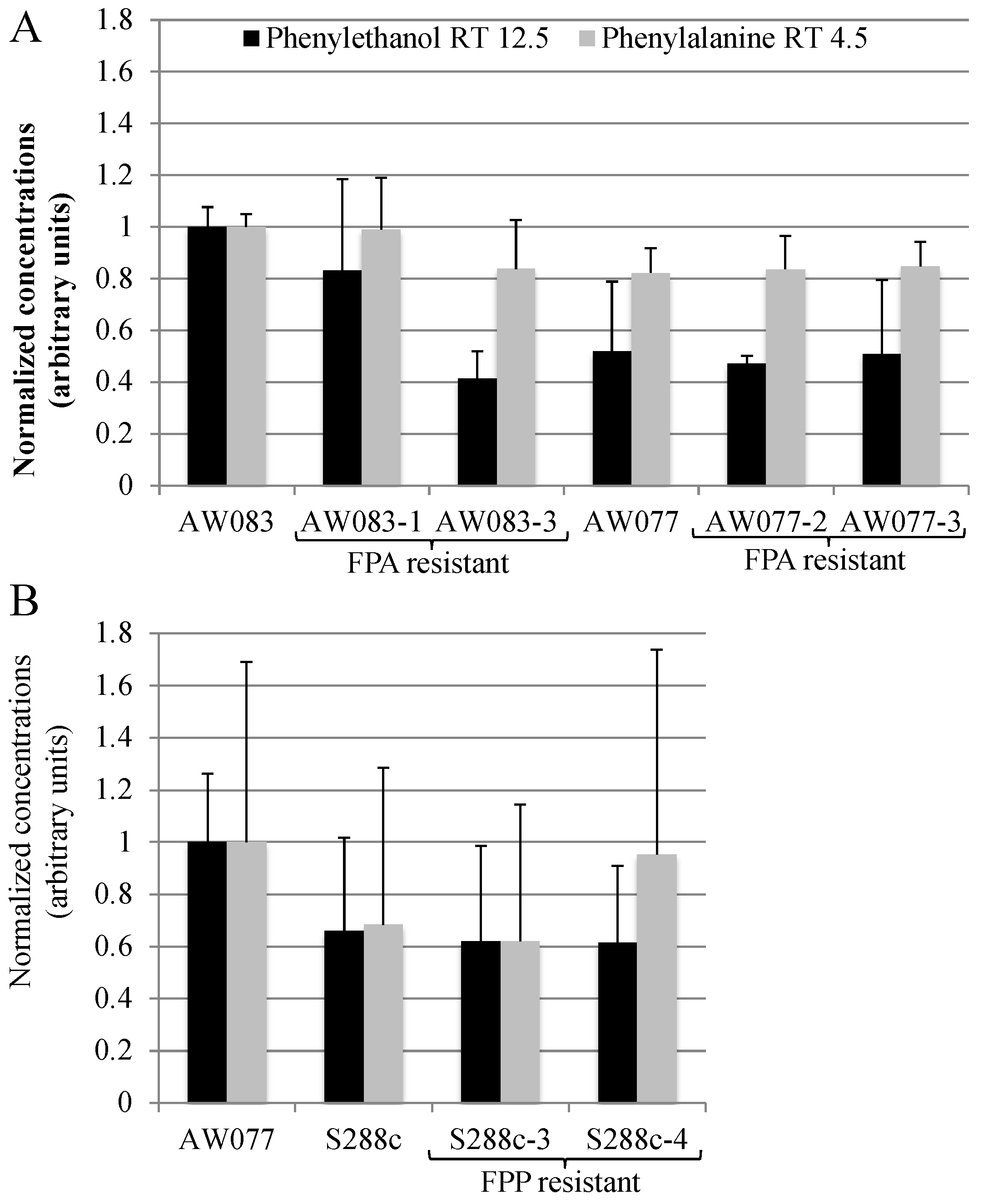
2]. We wanted to test whether FPA resistant strains obtained herein would produce high amounts of phenylethanol. To do this, we inoculated strains into medium containing phenylalanine as the source of nitrogen and measured the phenylethanol and phenylalanine concentration in the medium after three days of growth using GC-MS (
Figure 9A).
Resistance to FPA did not induce higher phenylethanol production in the four strains that were tested. This is not surprising, as previously, the frequency of FPA-resistant mutants with high levels of phenylethanol production was approximately 20% [
15]. Thus, if we had tested five or more isolates we may have expected to obtain at least one. However, because of the low frequency of obtaining FPA-resistant strains we did not pursue this further. Because we had seen that FPP resistance was more difficult to attain than FPA, we predicted that these mutants may produce phenylalanine at a greater rate than FPA-resistant mutants and thus may have higher phenylethanol production. However, upon testing FPP-resistant strains, no increase in phenylethanol production was noted (
Figure 9B). Thus, FPA and FPP resistance is not sufficient to induce high levels of phenylethanol production.
4. Discussion
Here, we report that FPP can act as a toxic agent in diverse strains of yeast, and this effect is at least partially mediated by ARO8 and ARO9 conversion to FPA. Whether FPP exerts its toxic effect solely by conversion to FPA or via another route rquires further investigation. FPP exhibited higher toxicity than FPA, suggesting a separate mechanism of toxicity, however this could be due to enhanced adsorption to cell wall, decreased cell uptake, or other enzymatic activities that decrease the effective concentration of FPA.
The mechanism of FPP toxicity still remains somewhat of a mystery. It is possible that there is preferred transport of phenylpyruvate over phenylalanine although evidence for this is absent. Another hypothesis is that FPP is converted into some other compound that FPA is not, such as fluoro-phenylethanol. Phenylalanine degradation follows this pathway: phenylalanine > phenylpyruvate > phenylacetaldehyde > phenylethanol. Phenylpyruvate is one enzymatic step closer to phenylethanol, and it is possible the fluorine moiety decreases its ability to convert to fluoro-phenylalanine, but instead allows it to become fluoro-phenylethanol. Phenylethanol has been shown to be toxic to
C. albicans at 20 mg L
-l [
16].
We observed differences in the sensitivity of strains to FPP and FPA depending on the method of delivery—solid or liquid medium. Phenylpyruvate shows increased solubility relative to phenylalanine (112 g L
−1 versus 27 g L
−1) that may explain its increased toxicity in liquid medium [
17]. This observation may be explained by the increased exposure of cells to all toxic molecules in a liquid, as opposed to a solid medium in which cells are only exposed to toxins within close physical proximity. Previous groups have reported similar enhanced drug potency in liquid media [
18].
The effect of fluorinated phenylalanine metabolites has been observed previously in other fungal species. Fluorophenylalanine at concentrations as low as 2.5 mM can inhibit growth of fungal species
Cladosporium cucumerinum and
Colletotrichum lagenarium [
19]. We found that FPP was toxic to a variety of yeast species at 2.74 mM indicating that it is a more potent fungal inhibitor than FPA.
Interestingly, we also found that
PHA2 was not required for slow growth. This is in contrast to several other works that indicate that
PHA2 causes an absolute requirement for phenylalanine in the medium [
10,
20]. However, strain growth is typically assayed over a period of up to three days on solid medium, while in our studies, liquid medium was used which allows for faster growth.
PHA2 converts prephenate to phenylpyruvate, but an alternative method to converting prephenate to phenylalanine is via the arogenate dehydratase enzyme. This enzyme has not been reported in
S. cerevisiae. The closest species having a putative arogenate dehydratase is
Neurospora crassa, which shares the same phylum with
S. cerevisiae. Further investigation is needed to determine the mechanism of slow phenylalanine generation in strains lacking
PHA2.
These observations should be helpful in better understanding phenylalanine metabolism and may be of interest in identifying potential secondary targets of FPP.