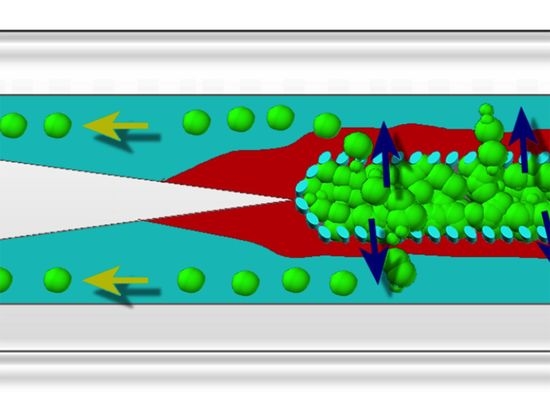
A 3D Microfluidic Model to Recapitulate Cancer Cell Migration and Invasion
Abstract
:1. Introduction
2. Materials and Methods
3. Results and Discussion
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Chaffer, C.L.; Weinberg, R.A. A Perspective on Cancer Cell Metastasis. Science 2011, 331, 1559–1564. [Google Scholar] [CrossRef] [PubMed]
- Duffy, M.J.; McGowan, P.M.; Gallagher, W.M. Cancer invasion and metastasis: Changing views. J. Pathol. 2008, 214, 283–293. [Google Scholar] [CrossRef] [PubMed]
- Zijlstra, A.; Lewis, J.; DeGryse, B.; Stuhlmann, H.; Quigley, J.P. The Inhibition of Tumor Cell Intravasation and Subsequent Metastasis via Regulation of In Vivo Tumor Cell Motility by the Tetraspanin CD151. Cancer Cell 2008, 13, 221–234. [Google Scholar] [CrossRef] [PubMed]
- Eckhardt, B.L.; Francis, P.A.; Parker, B.S.; Anderson, R.L. Strategies for the discovery and development of therapies for metastatic breast cancer. Nat. Rev. Drug Discov. 2012, 11, 479–497. [Google Scholar] [CrossRef] [PubMed]
- Gupta, G.P.; Massagué, J. Cancer metastasis: Building a framework. Cell 2006, 127, 679–695. [Google Scholar] [CrossRef] [PubMed]
- Paweletz, C.P.; Charboneau, L.; Liotta, L.A. Overview of Metastasis Assays. Curr. Protoc. Cell Biol. 2001, 12, 19.1.1–19.1.9. [Google Scholar] [CrossRef]
- Deryugina, E.; Quigley, J. Chick embryo chorioallantoic membrane model systems to study and visualize human tumor cell metastasis. Histochem. Cell Biol. 2008, 130, 1119–1130. [Google Scholar] [CrossRef] [PubMed]
- Sahai, E. Mechanisms of cancer cell invasion. Curr. Opin. Genet. Dev. 2005, 15, 87–96. [Google Scholar] [CrossRef] [PubMed]
- Kenny, H.A.; Krausz, T.; Yamada, S.D.; Lengyel, E. Use of a novel 3D culture model to elucidate the role of mesothelial cells, fibroblasts and extra-cellular matrices on adhesion and invasion of ovarian cancer cells to the omentum. Int. J. Cancer 2007, 121, 1463–1472. [Google Scholar] [CrossRef] [PubMed]
- Chien, W.; O’Kelly, J.; Lu, D.; Leiter, A.; Sohn, J.; Yin, D.; Karlan, B.; Vadgama, J.; Lyons, K.M.; Koeffler, H.P. Expression of connective tissue growth factor (CTGF/CCN2) in breast cancer cells is associated with increased migration and angiogenesis. Int. J. Oncol. 2011, 38, 1741–1747. [Google Scholar] [CrossRef] [PubMed]
- Yamauchi, A.; Yamamura, M.; Katase, N.; Itadani, M.; Okada, N.; Kobiki, K.; Nakamura, M.; Yamaguchi, Y.; Kuribayashi, F. Evaluation of pancreatic cancer cell migration with multiple parameters in vitro by using an optical real-time cell mobility assay device. BMC Cancer 2017, 17, 234. [Google Scholar] [CrossRef] [PubMed]
- Yarrow, J.C.; Perlman, Z.E.; Westwood, N.J.; Mitchison, T.J. A high-throughput cell migration assay using scratch wound healing, a comparison of image-based readout methods. BMC Biotechnol. 2004, 4, 21. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xia, M.; Yao, L.; Zhang, Q.; Wang, F.; Mei, H.; Guo, X.; Huang, W. Long noncoding RNA HOTAIR promotes metastasis of renal cell carcinoma by up-regulating histone H3K27 demethylase JMJD3. Oncotarget 2017, 8, 19795–19802. [Google Scholar] [CrossRef] [PubMed]
- Hulkower, K.I.; Herber, R.L. Cell Migration and Invasion Assays as Tools for Drug Discovery. Pharmaceutics 2011, 3, 107–124. [Google Scholar] [CrossRef] [PubMed]
- Nie, F.-Q.; Yamada, M.; Kobayashi, J.; Yamato, M.; Kikuchi, A.; Okano, T. On-chip cell migration assay using microfluidic channels. Biomaterials 2007, 28, 4017–4022. [Google Scholar] [CrossRef] [PubMed]
- Saadi, W.; Wang, S.-J.; Lin, F.; Jeon, N.L. A parallel-gradient microfluidic chamber for quantitative analysis of breast cancer cell chemotaxis. Biomed. Microdevices 2006, 8, 109–118. [Google Scholar] [CrossRef] [PubMed]
- Jeon, N.L.; Baskaran, H.; Dertinger, S.K.W.; Whitesides, G.M.; van der Water, L.; Toner, M. Neutrophil chemotaxis in linear and complex gradients of interleukin-8 formed in a microfabricated device. Nat. Biotechnol. 2002, 20, 826–830. [Google Scholar] [CrossRef] [PubMed]
- Chaw, K.C.; Manimaran, M.; Tay, E.H.; Swaminathan, S. Multi-step microfluidic device for studying cancer metastasis. Lab Chip 2007, 7, 1041–1047. [Google Scholar] [CrossRef] [PubMed]
- Businaro, L.; de Ninno, A.; Schiavoni, G.; Lucarini, V.; Ciasca, G.; Gerardino, A.; Belardelli, F.; Gabriele, L.; Mattei, F. Cross talk between cancer and immune cells: Exploring complex dynamics in a microfluidic environment. Lab Chip 2013, 13, 229–239. [Google Scholar] [CrossRef] [PubMed]
- Mi, S.; Du, Z.; Xu, Y.; Wu, Z.; Qian, X.; Zhang, M.; Sun, W. Microfluidic co-culture system for cancer migratory analysis and anti-metastatic drugs screening. Sci. Rep. 2016, 6, 35544. [Google Scholar] [CrossRef] [PubMed]
- Bogenrieder, T.; Herlyn, M. Axis of evil: Molecular mechanisms of cancer metastasis. Oncogene 2003, 22, 6524–6536. [Google Scholar] [CrossRef] [PubMed]
- Erler, J.T.; Weaver, V.M. Three-dimensional context regulation of metastasis. Clin. Exp. Metastasis 2009, 26, 35–49. [Google Scholar] [CrossRef] [PubMed]
- Chung, S.; Sudo, R.; Mack, P.J.; Wan, C.-R.; Vickerman, V.; Kamm, R.D. Cell migration into scaffolds under co-culture conditions in a microfluidic platform. Lab Chip 2009, 9, 269–275. [Google Scholar] [CrossRef] [PubMed]
- Zervantonakis, I.K.; Hughes-Alford, S.K.; Charest, J.L.; Condeelis, J.S.; Gertler, F.B.; Kamm, R.D. Three-dimensional microfluidic model for tumor cell intravasation and endothelial barrier function. Proc. Natl. Acad. Sci. USA 2012, 109, 13515–13520. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.; Hyun, E.; Seo, J.; Blundell, C.; Kim, H.C.; Lee, E.; Lee, S.H.; Moon, A.; Moon, W.C.; Huh, D. A microengineered pathophysiological model of early-stage breast cancer. Lab Chip 2015, 15, 3350–3357. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Gao, D.; Liu, H.; Lin, S.; Jiang, Y. Drug cytotoxicity and signaling pathway analysis with three-dimensional tumor spheroids in a microwell-based microfluidic chip for drug screening. Anal. Chim. Acta 2015, 898, 85–92. [Google Scholar] [CrossRef] [PubMed]
- Toh, Y.C.; Zhang, C.; Zhang, J.; Khong, Y.M.; Chang, S.; Samper, V.D.; van Noort, D.; Hutmacher, D.W.; Yu, H. A novel 3D mammalian cell perfusion-culture system in microfluidic channels. Lab Chip 2007, 7, 302–309. [Google Scholar] [CrossRef] [PubMed]
- Kihara, T.; Ito, J.; Miyake, J. Measurement of Biomolecular Diffusion in Extracellular Matrix Condensed by Fibroblasts Using Fluorescence Correlation Spectroscopy. PLoS ONE 2013, 8, e82382. [Google Scholar] [CrossRef] [PubMed]
- Yamazaki, D.; Kurisu, S.; Takenawa, T. Regulation of cancer cell motility through actin reorganization. Cancer Sci. 2005, 96, 379–386. [Google Scholar] [CrossRef] [PubMed]
- Croft, D.R.; Olson, M.F. Regulating the conversion between rounded and elongated modes of cancer cell movement. Cancer Cell 2008, 14, 349–351. [Google Scholar] [CrossRef] [PubMed]
- Ong, S.M.; Zhang, C.; Toh, Y.-C.; Kim, S.-H.; Foo, H.-L.; Tan, C.-H.; van Noort, D.; Park, S.; Yu, H. A gel-free 3D microfluidic cell culture system. Biomaterials 2008, 29, 3237–3244. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Zhao, Z.; Abdul Rahim, N.A.; van Noort, D.; Yu, H. Towards human on a chip: Culturing multiple cell types on a chip with compartmentalized microenvironments. Lab Chip 2009, 9, 3185–3192. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, D.T.T.; van Noort, D.; Jeong, I.K.; Park, S. Endocrine systems on chip for a diabetes treatment model. Biofabrication 2017, 9, 015021. [Google Scholar] [CrossRef] [PubMed]
- Isaiah, J.F. The organ microenvironment and cancer metastasis. Differentiation 2002, 70, 498–505. [Google Scholar] [CrossRef]
- Brandt, B.; Heyder, C.; Gloria-Maercker, E.; Hatzmann, W.; Rötger, A.; Kemming, D.; Zänker, K.S.; Entschladen, F.; Dittmar, T. 3D-extravasation model—Selection of highly motile and metastatic cancer cells. Sem. Cancer Biol. 2005, 15, 387–395. [Google Scholar] [CrossRef] [PubMed]
- Heyder, C.; Gloria, M.; Gloria-Maercker, E.; Hatzmann, W.; Niggermann, D.; Zänker, K.S.; Dittmar, T. Realtime visualization of tumor cell/endothelial cell interactions during transmigration across the endothelial barrier. J. Cancer Res. Clin. Oncol. 2002, 128, 533–538. [Google Scholar] [CrossRef] [PubMed]
- Sheridan, C.; Kishimoto, H.; Fuchs, R.; Mehrotra, S.; Bhat-Nakshatri, P.; Turner, C.H.; Goulet, R., Jr.; Badve, S.; Nakshatri, H. CD44+/CD24− breast cancer cells exhibit enhanced invasive properties: An early step necessary for metastasis. Breast Cancer Res. 2006, 8, R59. [Google Scholar] [CrossRef] [PubMed]
- Brown, N.S.; Bicknell, R. Cell Migration and the Boyden Chamber. In Metastasis Research Protocols; Brooks, S.A., Schumacher, U., Eds.; Humana Press: Totowa, NJ, USA, 2001; pp. 47–54. ISBN 0-89603-610-3. [Google Scholar]
- Overall, C.M.; Kleifeld, O. Tumour microenvironment—Opinion: Validating matrix metalloproteinases as drug targets and anti-targets for cancer therapy. Nat. Rev. Cancer 2006, 6, 227–239. [Google Scholar] [CrossRef] [PubMed]
- Sliva, D. Signaling Pathways Responsible for Cancer Cell Invasion as Targets for Cancer Therapy. Curr. Cancer Drug Targets 2004, 4, 327–336. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Toh, Y.-C.; Raja, A.; Yu, H.; Van Noort, D. A 3D Microfluidic Model to Recapitulate Cancer Cell Migration and Invasion. Bioengineering 2018, 5, 29. https://doi.org/10.3390/bioengineering5020029
Toh Y-C, Raja A, Yu H, Van Noort D. A 3D Microfluidic Model to Recapitulate Cancer Cell Migration and Invasion. Bioengineering. 2018; 5(2):29. https://doi.org/10.3390/bioengineering5020029
Chicago/Turabian StyleToh, Yi-Chin, Anju Raja, Hanry Yu, and Danny Van Noort. 2018. "A 3D Microfluidic Model to Recapitulate Cancer Cell Migration and Invasion" Bioengineering 5, no. 2: 29. https://doi.org/10.3390/bioengineering5020029