Biology and Function of miR159 in Plants
Abstract
:1. Introduction
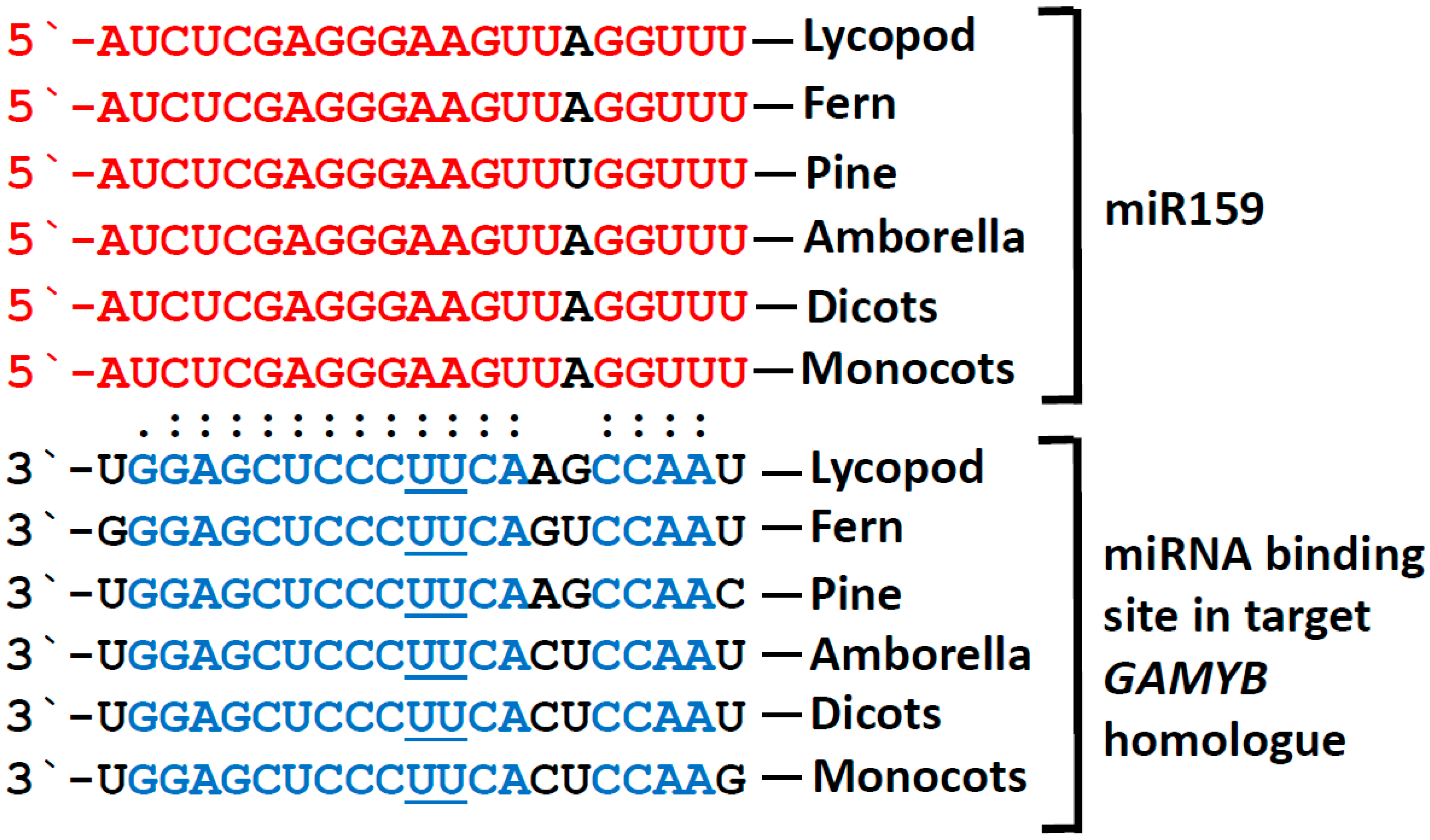
2. MiR159 is Strongly Conserved and Highly Abundant Throughout the Plant Kingdom
3. GAMYB and GAMYB-like Genes are the Only Conserved Targets of miR159
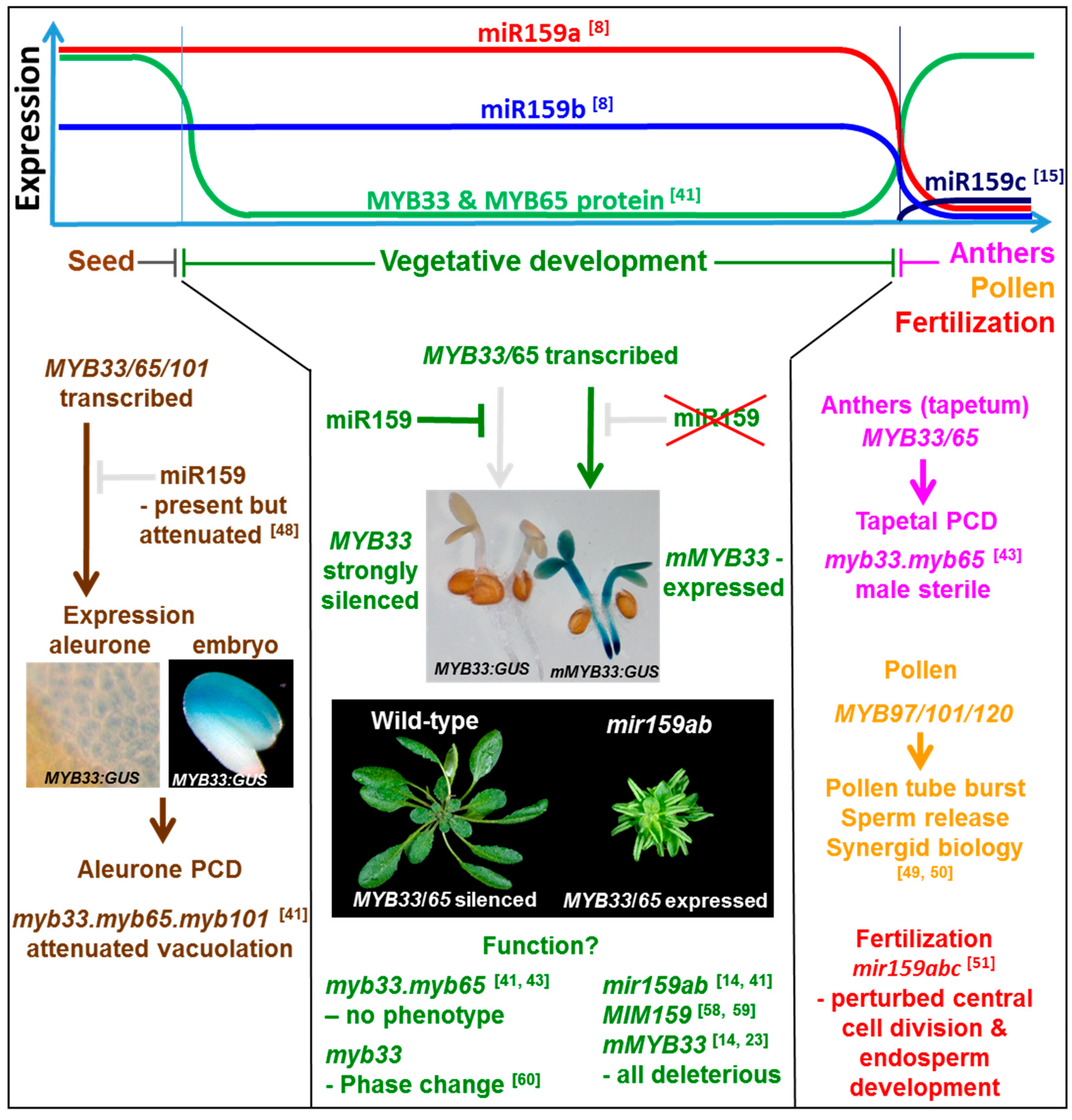
4. The miR159-GAMYB Pathway in Arabidopsis
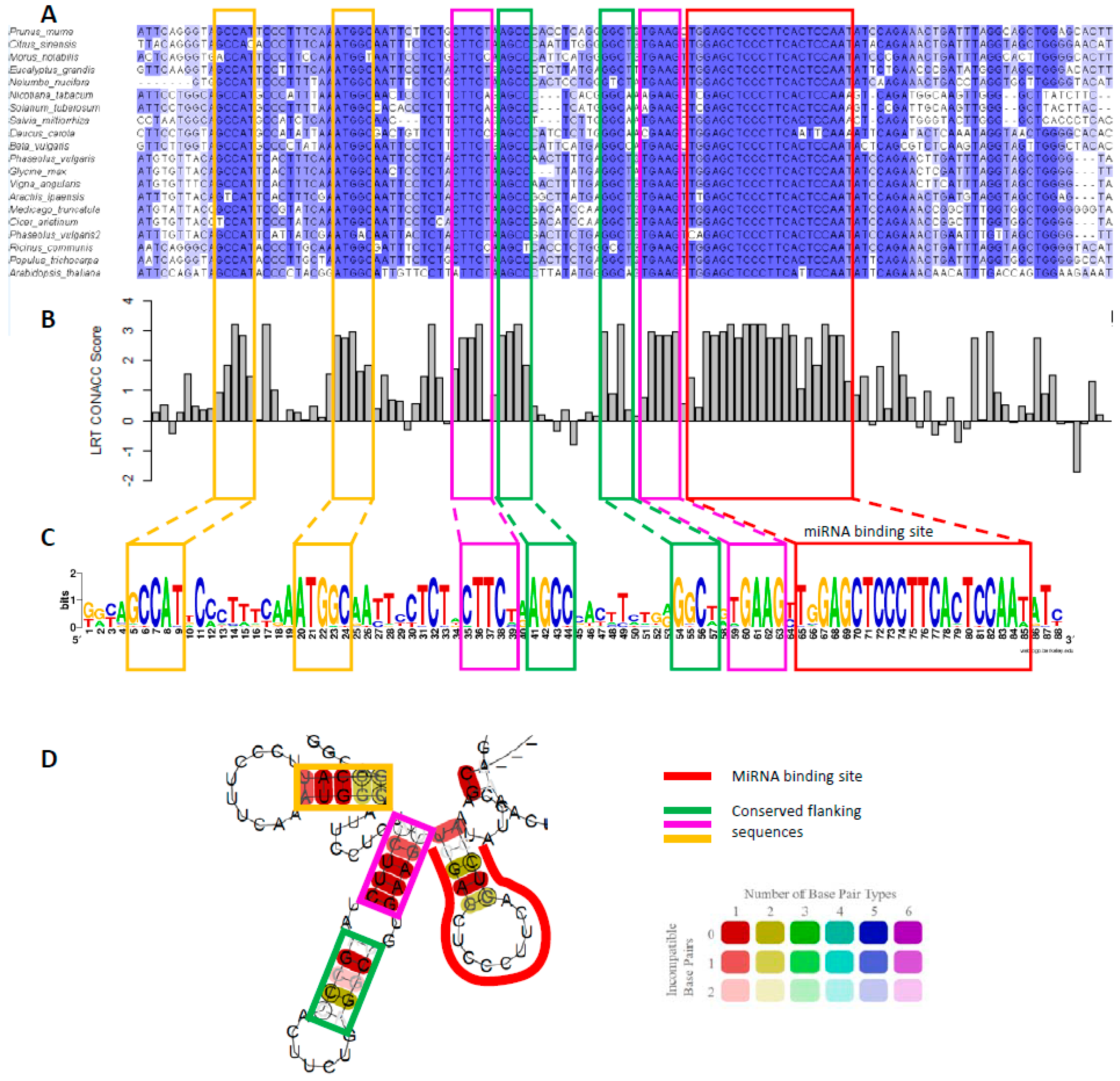
5. Conserved RNA Secondary Structures in MYB33/65 Promote miR159-Mediated Silencing
6. The Function of miR159-MYB Pathway in Plant Development
6.1. A Role in Male Reproductive Development
6.2. A Role in Seed Development
6.3. The Role of miR159-GAMYB Pathway in Vegetative Tissues
6.4. A Role of miR159 in Controlling GA-Mediated Flowering-Time and Growth?
6.5. Fruit and Reproductive Development
7. The Function of the miR159-MYB Pathway in Plant Stress
7.1. Abiotic Stress
7.2. Biotic Stress
8. Conclusions and Some Unresolved Questions
- Why are MYB33 and MYB65 transcribed in vegetative tissues where failure to fully repress them results in a detrimental effect? What selective advantage does this give the plant?
- One hypothesis is that if miR159 is inhibited by a certain trigger, and strong MYB33/65 expression occurs, growth inhibition (or another unknown process) may result in a beneficial outcome (e.g., drought conditions to slow growth). However, currently, no triggers to inhibit miR159 to enable strong MYB expression are known.
- A second hypothesis would be that MYB33/65 are not silenced in all vegetative tissues, but in certain cells they are expressed where they confer a selective advantage. Some evidence suggests GAMYB is involved in the transition to flowering, and VPC in Arabidopsis. But currently there is much conflicting data. For instance in Arabidopsis, overexpressing miR159 represses flowering-time, and inhibition of miR159 represses VPC. Other studies have found no role for miR159 in flowering. More work is needed here to clarify these roles, and how conserved they are across species.
- Why is expression of GAMYB in vegetative tissues deleterious and how does it inhibit growth? What down-stream events are these genes triggering? Although some studies have started to address this, more work is needed for a clearer understanding.
- Is GAMYB function related to the way it is regulated, i.e., strongly transcribed, only to then be strongly silenced by miR159? Does miR159 have a role in stress response? Again, many studies have identified changes to miR159 levels in response to a host of different biotic/abiotic stresses, but currently there is no clearly defined role for this miR159 concerning stress tolerance/response.
- How does the conserved RNA secondary structure associated with the miR159-binding sites of GAMYB genes promote their silencing by miR159? Can this structure facilitate a complex regulatory mechanism, enabling strong silencing in some tissues, but poor silencing in others, depending on a dynamic secondary structure configuration? i.e., acting like a riboswitch concerning silencing.
- What is the role of miR159-mediate regulation on non-GAMYB targets? For example, DUO1 has a conserved miR159-binding site, but the role of miR159 in controlling the expression of this gene remains unclear.
- What is the role of miR159 in female fertility? Why are Arabidopsis mir159ab seeds small and misshapen (likewise rice STTM159 grains are small)? Why does the central cell still divide in some mir159abc ovules? How can a seed still form (from mir159abc pollen) with a viable embryo when the endosperm divisions stop apparently so early?
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Axtell, M.J.; Bartel, D.P. Antiquity of microRNAs and their targets in land plants. Plant Cell 2005, 17, 1658–1673. [Google Scholar] [CrossRef] [PubMed]
- Chávez Montes, R.A.; de Fátima Rosas-Cárdenas, F.; De Paoli, E.; Accerbi, M.; Rymarquis, L.A.; Mahalingam, G.; Marsch-Martínez, N.; Meyers, B.C.; Green, P.J.; de Folter, S. Sample sequencing of vascular plants demonstrates widespread conservation and divergence of microRNAs. Nat. Commun. 2014, 5, 3722. [Google Scholar] [CrossRef] [PubMed]
- You, C.; Cui, J.; Wang, H.; Qi, X.; Kuo, L.Y.; Ma, H.; Gao, L.; Mo, B.; Chen, X. Conservation and divergence of small RNA pathways and microRNAs in land plants. Genome Biol. 2017, 18, 158. [Google Scholar] [CrossRef] [PubMed]
- Axtell, M.J.; Meyers, B.C. Revisiting criteria for plant microRNA annotation in the era of big data. Plant Cell 2018, 30, 272–284. [Google Scholar] [CrossRef] [PubMed]
- Alaba, S.; Piszczalka, P.; Pietrykowska, H.; Pacak, A.M.; Sierocka, I.; Nuc, P.W.; Singh, K.; Plewka, P.; Sulkowska, A.; Jarmolowski, A.; et al. The liverwort Pellia endiviifolia shares microtranscriptomic traits that are common to green algae and land plants. Plant J. 2014, 80, 331–344. [Google Scholar]
- Fahlgren, N.; Howell, M.D.; Kasschau, K.D.; Chapman, E.J.; Sullivan, C.M.; Cumbie, J.S.; Givan, S.A.; Law, T.F.; Grant, S.R.; Dang, J.L.; et al. High-throughput sequencing of Arabidopsis microRNAs: Evidence for frequent birth and death of MIRNA genes. PLoS ONE 2007, 2, e219. [Google Scholar] [CrossRef] [PubMed]
- Jeong, D.H.; Park, S.; Zhai, J.; Gurazada, S.G.R.; De Paoli, E.; Meyers, B.C.; Green, P.J. Massive analysis of rice small RNAs: Mechanistic implications of regulated microRNAs and variants for differential target RNA cleavage. Plant Cell 2011, 23, 4185–4207. [Google Scholar] [CrossRef]
- Rajagopalan, R.; Vaucheret, H.; Trejo, J.; Bartel, D.P. A diverse and evolutionarily fluid set of microRNAs in Arabidopsis thaliana. Genes Dev. 2006, 20, 3407–3425. [Google Scholar] [CrossRef] [PubMed]
- Szittya, G.; Moxon, S.; Santos, D.M.; Jing, R.; Fevereiro, M.P.; Moulton, V.; Dalmay, T. High-throughput sequencing of Medicago truncatula short RNAs identifies eight new miRNA families. BMC Genom. 2008, 9, 593. [Google Scholar] [CrossRef]
- Mao, W.; Li, Z.; Xia, X.; Li, Y.; Yu, J. A combined approach of high-throughput sequencing and degradome analysis reveals tissue specific expression of microRNAs and their targets in cucumber. PLoS ONE 2012, 7, e33040. [Google Scholar] [CrossRef]
- Kozomara, A.; Birgaoanu, M.; Griffiths-Jones, S. miRBase: From microRNA sequences to function. Nucleic Acids Res. 2019, 47, D155–D162. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Chia, J.M.; Kumari, S.; Stein, J.C.; Liu, Z.; Narechania, A.; Maher, C.A.; Guill, K.; McMullen, M.D.; Ware, D. A genome-wide characterization of microRNA genes in maize. PLoS Genet. 2009, 5, e1000716. [Google Scholar] [CrossRef] [PubMed]
- Schwab, R.; Palatnik, J.F.; Riester, M.; Schommer, C.; Schmid, M.; Weigel, D. Specific effects of microRNAs on the plant transcriptome. Dev. Cell 2005, 8, 517–527. [Google Scholar] [CrossRef] [PubMed]
- Allen, R.S.; Li, J.; Stahle, M.I.; Dubroué, A.; Gubler, F.; Millar, A.A. Genetic analysis reveals functional redundancy and the major target genes of the Arabidopsis miR159 family. Proc. Natl. Acad. Sci. USA 2007, 104, 16371–16376. [Google Scholar] [CrossRef] [PubMed]
- Allen, R.S.; Li, J.; Alonso-Peral, M.M.; White, R.G.; Gubler, F.; Millar, A.A. MicroR159 regulation of most conserved targets in Arabidopsis has negligible phenotypic effects. Silence 2010, 1, 18. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.H.; Cho, J.S.; Lee, J.H.; Bae, S.Y.; Choi, Y.I.; Park, E.J.; Lee, H.; Ko, J.H. Poplar MYB transcription factor PtrMYB012 and its Arabidopsis AtGAMYB orthologs are differentially repressed by the Arabidopsis miR159 family. Tree Physiol. 2018, 38, 801–812. [Google Scholar] [CrossRef] [PubMed]
- Palatnik, J.F.; Wollmann, H.; Schommer, C.; Schwab, R.; Boisbouvier, J.; Rodriguez, R.; Warthmann, N.; Allen, E.; Dezulian, T.; Huson, D.; et al. Sequence and expression differences underlie functional specialization of Arabidopsis microRNAs miR159 and miR319. Dev. Cell 2007, 13, 115–125. [Google Scholar] [CrossRef]
- Bologna, N.G.; Mateos, J.L.; Bresso, E.G.; Palatnik, J.F. A loop-to-base processing mechanism underlies the biogenesis of plant microRNAs miR319 and miR159. EMBO J. 2009, 28, 3646–3656. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.; Li, C.; Ding, G.; Jin, Y. Evolution of MIR159/319 microRNA genes and their post-transcriptional regulatory link to siRNA pathways. BMC Evol. Biol. 2011, 11, 122. [Google Scholar] [CrossRef]
- Aya, K.; Hiwatashi, Y.; Kojima, M.; Sakakibara, H.; Ueguchi-Tanaka, M.; Hasebe, M.; Matsuoka, M. The Gibberellin perception system evolved to regulate a pre-existing GAMYB-mediated system during land plant evolution. Nat. Commun. 2011, 2, 544. [Google Scholar] [CrossRef]
- Tsuzuki, M.; Nishihama, R.; Ishizaki, K.; Kurihara, Y.; Matsui, M.; Bowman, J.L.; Kohchi, T.; Hamada, T.; Watanabe, Y. Profiling and characterization of small RNAs in the Liverwort, Marchantia polymorpha, belonging to the first diverged land plants. Plant Cell Physiol. 2016, 57, 359–372. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.S.; Bowman, J.L. MicroRNAs in Marchantia polymorpha. New Phytol. 2018, 220, 409–416. [Google Scholar] [CrossRef] [PubMed]
- Palatnik, J.F.; Allen, E.; Wu, X.; Schommer, C.; Schwab, R.; Carrington, J.C.; Weigel, D. Control of leaf morphogenesis by microRNAs. Nature 2003, 425, 257–263. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, J.; Reichel, M.; Li, Y.; Millar, A.A. The functional scope of plant microRNA-mediated silencing. Trends Plant Sci. 2014, 19, 750–756. [Google Scholar] [CrossRef] [PubMed]
- Addo-Quaye, C.; Eshoo, T.W.; Bartel, D.P.; Axtell, M.J. Endogenous siRNA and miRNA targets identified by sequencing of the Arabidopsis degradome. Curr. Biol. 2008, 18, 758–762. [Google Scholar] [CrossRef] [PubMed]
- Song, Q.X.; Liu, Y.F.; Hu, X.Y.; Zhang, W.K.; Ma, B.; Chen, S.Y.; Zhang, J.S. Identification of miRNAs and their target genes in developing soybean seeds by deep sequencing. BMC Plant Biol. 2011, 11, 5. [Google Scholar] [CrossRef] [PubMed]
- Liu, N.; Tu, L.; Tang, W.; Gao, W.; Lindsey, K.; Zhang, X. Small RNA and degradome profiling reveals a role for miRNAs and their targets in the developing fibers of Gossypium barbadense. New Phytol. 2015, 206, 352–367. [Google Scholar]
- Zhang, J.; Zeng, R.; Chen, J.; Liu, X.; Liao, Q. Identification of conserved microRNAs and their targets from Solanum lycopersicum Mill. Gene 2008, 423, 1–7. [Google Scholar] [CrossRef]
- An, F.M.; Chan, M.T. Transcriptome-wide characterization of miRNA-directed and non-miRNA-directed endonucleolytic cleavage using degradome analysis under low ambient temperature in Phalaenopsis aphrodite subsp. formosana. Plant Cell Physiol. 2012, 53, 1737–1750. [Google Scholar] [CrossRef]
- Luo, X.; Gao, Z.; Shi, T.; Cheng, Z.; Zhang, Z.; Ni, Z. Identification of miRNAs and their target genes in peach (Prunus persica L.) using high-throughput sequencing and degradome analysis. PLoS ONE 2013, 8, e79090. [Google Scholar] [CrossRef]
- Sun, F.; Guo, G.; Du, J.; Guo, W.; Peng, H.; Ni, Z.; Sun, Q.; Yao, Y. Whole-genome discovery of miRNAs and their targets in wheat (Triticum aestivum L.). BMC Plant Biol. 2014, 14, 142. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.F.; Zheng, Y.; Addo-Quaye, C.; Zhang, L.; Saini, A.; Jagadeeswaran, G.; Axtell, M.J.; Zhang, W.; Sunkar, R. Transcriptome-wide identification of microRNA targets in rice. Plant J. 2010, 62, 742–759. [Google Scholar] [CrossRef]
- Curaba, J.; Spriggs, A.; Taylor, J.; Li, Z.; Helliwell, C. miRNA regulation in the early development of barley seed. BMC Plant Biol. 2012, 12, 120. [Google Scholar] [CrossRef] [PubMed]
- Buxdorf, K.; Hendelman, A.; Stav, R.; Lapidot, M.; Ori, N.; Arazi, T. Identification and characterization of a novel miR159 target not related to MYB in tomato. Planta 2010, 232, 1009–1022. [Google Scholar] [CrossRef] [PubMed]
- Da Silva, E.M.; Silva, G.F.F.E.; Bidoia, D.B.; da Silva Azevedo, M.; de Jesus, F.A.; Pino, L.E.; Peres, L.E.P.; Carrera, E.; López-Díaz, I.; Nogueira, F.T.S. microRNA159-targeted SlGAMYB transcription factors are required for fruit set in tomato. Plant J. 2017, 92, 95–109. [Google Scholar] [CrossRef] [PubMed]
- Wan, L.C.; Zhang, H.; Lu, S.; Zhang, L.; Qiu, Z.; Zhao, Y.; Zeng, Q.Y.; Lin, J. Transcriptome-wide identification and characterization of miRNAs from Pinus densata. BMC Genom. 2012, 13, 132. [Google Scholar] [CrossRef] [PubMed]
- Amborella Genome Project. The Amborella genome and the evolution of flowering plants. Science 2013, 342, 1241089. [Google Scholar] [CrossRef]
- Li, W.F.; Zhang, S.G.; Han, S.Y.; Wu, T.; Zhang, J.H.; Qi, L.W. Regulation of LaMYB33 by miR159 during maintenance of embryogenic potential and somatic embryo maturation in Larix kaempferi (Lamb.) Carr. Plant Cell Tissue Organ Cult. 2013, 113, 131–136. [Google Scholar] [CrossRef]
- Pappas, M.D.C.R.; Pappas, G.J.; Grattapaglia, D. Genome-wide discovery and validation of Eucalyptus small RNAs reveals variable patterns of conservation and diversity across species of Myrtaceae. BMC Genom. 2015, 16, 1113. [Google Scholar] [CrossRef]
- Dai, X.; Zhuang, Z.; Zhao, P.X. psRNATarget: A plant small RNA target analysis server (2017 release). Nucleic Acids Res. 2018, 46, W49–W54. [Google Scholar] [CrossRef]
- Alonso-Peral, M.M.; Li, J.; Li, Y.; Allen, R.S.; Schnippenkoetter, W.; Ohms, S.; White, R.G.; Millar, A.A. The microRNA159-regulated GAMYB-like genes inhibit growth and promote programmed cell death in Arabidopsis. Plant Physiol. 2010, 154, 757–771. [Google Scholar] [CrossRef] [PubMed]
- German, M.A.; Pillay, M.; Jeong, D.H.; Hetawal, A.; Luo, S.; Janardhanan, P.; Kannan, V.; Rymarquis, L.A.; Nobuta, K.; German, R.; et al. Global identification of microRNA–target RNA pairs by parallel analysis of RNA ends. Nat. Biotech. 2008, 26, 941. [Google Scholar] [CrossRef] [PubMed]
- Millar, A.A.; Gubler, F. The Arabidopsis GAMYB-like genes, MYB33 and MYB65, are microRNA-regulated genes that redundantly facilitate anther development. Plant Cell 2005, 17, 705–721. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Z.; Reichel, M.; Deveson, I.; Wong, G.; Li, J.; Millar, A.A. Target RNA secondary structure is a major determinant of miR159 efficacy. Plant Physiol. 2017, 174, 1764–1778. [Google Scholar] [CrossRef] [PubMed]
- Seitz, H. Redefining microRNA targets. Curr. Biol. 2009, 19, 870–873. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Alonso-Peral, M.; Wong, G.; Wang, M.B.; Millar, A.A. Ubiquitous miR159 repression of MYB33/65 in Arabidopsis rosettes is robust and is not perturbed by a wide range of stresses. BMC Plant Biol. 2016, 16, 179. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Reichel, M.; Millar, A.A. Determinants beyond both complementarity and cleavage govern microR159 efficacy in Arabidopsis. PLoS Genet. 2014, 10, e1004232. [Google Scholar] [CrossRef] [PubMed]
- Alonso-Peral, M.M.; Sun, C.; Millar, A.A. MicroRNA159 can act as a switch or tuning microRNA independently of its abundance in Arabidopsis. PLoS ONE 2012, 7, e34751. [Google Scholar] [CrossRef] [PubMed]
- Leydon, A.R.; Beale, K.M.; Woroniecka, K.; Castner, E.; Chen, J.; Horgan, C.; Palanivelu, R.; Johnson, M.A. Three MYB transcription factors control pollen tube differentiation required for sperm release. Curr. Biol. 2013, 23, 1209–1214. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.; Tan, Z.M.; Zhu, L.; Niu, Q.K.; Zhou, J.J.; Li, M.; Chen, L.Q.; Zhang, X.Q.; Ye, D. MYB97, MYB101 and MYB120 function as male factors that control pollen tube-synergid interaction in Arabidopsis thaliana fertilization. PLoS Genet. 2013, 9, e1003933. [Google Scholar] [CrossRef]
- Zhao, Y.; Wang, S.; Wu, W.; Li, L.; Jiang, T.; Zheng, B. Clearance of maternal barriers by paternal miR159 to initiate endosperm nuclear division in Arabidopsis. Nat. Commun. 2018, 9, 5011. [Google Scholar] [CrossRef] [PubMed]
- Achard, P.; Herr, A.; Baulcombe, D.C.; Harberd, N.P. Modulation of floral development by a gibberellin-regulated microRNA. Development 2004, 131, 3357–3365. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alves-Junior, L.; Niemeier, S.; Hauenschild, A.; Rehmsmeier, M.; Merkle, T. Comprehensive prediction of novel microRNA targets in Arabidopsis thaliana. Nucleic Acids Res. 2009, 37, 4010–4021. [Google Scholar] [CrossRef] [PubMed]
- Pollard, K.S.; Hubisz, M.J.; Rosenbloom, K.R.; Siepel, A. Detection of nonneutral substitution rates on mammalian phylogenies. Genome Res. 2010, 20, 110–121. [Google Scholar] [CrossRef]
- Hubisz, M.J.; Pollard, K.S.; Siepel, A. PHAST and RPHAST: Phylogenetic analysis with space/time models. Brief Bioinform. 2011, 12, 41–51. [Google Scholar] [CrossRef] [PubMed]
- Crooks, G.; Hon, G.; Chandonia, J.; Brenner, S. WebLogo: A sequence logo generator. Genome Res. 2004, 14, 1188–1190. [Google Scholar] [CrossRef] [PubMed]
- Bernhart, S.H.; Hofacker, I.L.; Will, S.; Gruber, A.R.; Stadler, P.F. RNAalifold: Improved consensus structure prediction for RNA alignments. BMC Bioinform. 2008, 9, 1–13. [Google Scholar] [CrossRef]
- Todesco, M.; Rubio-Somoza, I.; Paz-Ares, J.; Weigel, D. A collection of target mimics for comprehensive analysis of microRNA function in Arabidopsis thaliana. PLoS Genet. 2010, 6, e1001031. [Google Scholar] [CrossRef]
- Rubio-Somoza, I.; Weigel, D. Coordination of flower maturation by a regulatory circuit of three microRNAs. PLoS Genet. 2013, 9, e1003374. [Google Scholar] [CrossRef]
- Guo, C.; Xu, Y.; Shi, M.; Lai, Y.; Wu, X.; Wang, H.; Zhu, Z.; Poethig, R.S.; Wu, G. Repression of miR156 by miR159 regulates the timing of the juvenile-to-adult transition in Arabidopsis. Plant Cell 2017, 29, 1293–1304. [Google Scholar] [CrossRef]
- Zhao, Y.; Wen, H.; Teotia, S.; Du, Y.; Zhang, J.; Li, J.; Sun, H.; Tang, G.; Peng, T.; Zhao, Q. Suppression of microRNA159 impacts multiple agronomic traits in rice (Oryza sativa L.). BMC Plant Biol. 2017, 17, 215. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Zhang, J.; Yan, J.; Gou, F.; Mao, Y.; Tang, G.; Botella, J.R.; Zhu, J.K. Short tandem target mimic rice lines uncover functions of miRNAs in regulating important agronomic traits. Proc. Natl. Acad. Sci. USA 2017, 114, 5277–5282. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tsuji, H.; Aya, K.; Ueguchi-Tanaka, M.; Shimada, Y.; Nakazono, M.; Watanabe, R.; Nishizawa, N.K.; Gomi, K.; Shimada, A.; Kitano, H.; et al. GAMYB controls different sets of genes and is differentially regulated by microRNA in aleurone cells and anthers. Plant J. 2006, 47, 427–444. [Google Scholar] [CrossRef] [PubMed]
- Kaneko, M.; Inukai, Y.; Ueguchi-Tanaka, M.; Itoh, H.; Izawa, T.; Kobayashi, Y.; Hattori, T.; Miyao, A.; Hirochika, H.; Ashikari, M.; et al. Loss-of-function mutations of the rice GAMYB gene impair α-amylase expression in aleurone and flower development. Plant Cell 2004, 16, 33–44. [Google Scholar] [CrossRef] [PubMed]
- Murray, F.; Kalla, R.; Jacobsen, J.; Gubler, F. A role for HvGAMYB in anther development. Plant J. 2003, 33, 481–491. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Sun, F.; Cao, H.; Peng, H.; Ni, Z.; Sun, Q.; Yao, Y. TamiR159 directed wheat TaGAMYB cleavage and its involvement in anther development and heat response. PLoS ONE 2012, 7, e48445. [Google Scholar] [CrossRef]
- Li, X.; Bian, H.; Song, D.; Ma, S.; Han, N.; Wang, J.; Zhu, M. Flowering time control in ornamental gloxinia (Sinningia speciosa) by manipulation of miR159 expression. Ann. Bot. 2013, 111, 791–799. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, X.; Liu, B.; Wang, W.; Liu, X.; Chen, C.; Liu, X.; Yang, S.; Ren, H. A GAMYB homologue CsGAMYB1 regulates sex expression of cucumber via an ethylene-independent pathway. J. Exp. Bot. 2014, 65, 3201–3213. [Google Scholar] [CrossRef]
- Vallarino, J.G.; Osorio, S.; Bombarely, A.; Casañal, A.; Cruz-Rus, E.; Sánchez-Sevilla, J.F.; Amaya, I.; Giavalisco, P.; Fernie, A.R.; Botella, M.A.; et al. Central role of FaGAMYB in the transition of the strawberry receptacle from development to ripening. New Phytol. 2015, 208, 482–496. [Google Scholar] [CrossRef]
- Plackett, A.R.; Thomas, S.G.; Wilson, Z.A.; Hedden, P. Gibberellin control of stamen development: A fertile field. Trends Plant Sci. 2011, 16, 568–578. [Google Scholar] [CrossRef]
- Aya, K.; Ueguchi-Tanaka, M.; Kondo, M.; Hamada, K.; Yano, K.; Nishimura, M.; Matsuoka, M. Gibberellin modulates anther development in rice via the transcriptional regulation of GAMYB. Plant Cell 2009, 21, 1453–1472. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; De Storme, N.; Geelen, D. Gibberellin induces diploid pollen formation by interfering with meiotic cytokinesis. Plant Physiol. 2017, 173, 338–353. [Google Scholar] [CrossRef] [PubMed]
- Gubler, F.; Kalla, R.; Roberts, J.K.; Jacobsen, J.V. Gibberellin-regulated expression of a myb gene in barley aleurone cells: Evidence for Myb transactivation of a high-pI alpha-amylase gene promoter. Plant Cell 1995, 7, 1879–1891. [Google Scholar] [CrossRef] [PubMed]
- Gubler, F.; Raventos, D.; Keys, M.; Watts, R.; Mundy, J.; Jacobsen, J.V. Target genes and regulatory domains of the GAMYB transcriptional activator in cereal aleurone. Plant J. 1999, 17, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Guo, W.J.; Ho, T.H.D. An abscisic acid-induced protein; HVA22, inhibits gibberellin-mediated programmed cell death in cereal aleurone cells. Plant Physiol. 2008, 147, 1710–1722. [Google Scholar] [CrossRef] [PubMed]
- Reichel, M.; Li, Y.; Li, J.; Millar, A.A. Inhibiting plant microRNA activity: Molecular SPONGEs, target MIMICs and STTMs all display variable efficacies against target microRNAs. Plant Biotech. J. 2015, 13, 915–926. [Google Scholar] [CrossRef] [PubMed]
- Gong, X.; Bewley, D.J. A GAMYB-like gene in tomato and its expression during seed germination. Planta 2008, 228, 563–572. [Google Scholar] [CrossRef] [PubMed]
- Xue, T.; Liu, Z.; Dai, X.; Xiang, F. Primary root growth in Arabidopsis thaliana is inhibited by the miR159 mediated repression of MYB33, MYB65 and MYB101. Plant Sci. 2017, 262, 182–189. [Google Scholar] [CrossRef]
- Spanudakis, E.; Jackson, S. The role of microRNAs in the control of flowering time. J. Exp. Bot. 2014, 65, 365–380. [Google Scholar] [CrossRef]
- Conti, L. Hormonal control of the floral transition: Can one catch them all? Dev. Biol. 2017, 430, 288–301. [Google Scholar] [CrossRef]
- Gocal, G.F.; Sheldon, C.C.; Gubler, F.; Moritz, T.; Bagnall, D.J.; MacMillan, C.P.; Li, S.F.; Parish, R.W.; Dennis, E.S.; Weigel, D.; et al. GAMYB-like genes, flowering, and gibberellin signaling in Arabidopsis. Plant Physiol. 2001, 127, 1682–1693. [Google Scholar] [CrossRef] [PubMed]
- Blazquez, M.A.; Green, R.; Nilsson, O.; Sussman, M.R.; Weigel, D. Gibberellins promote flowering of arabidopsis by activating the LEAFY promoter. Plant Cell 1998, 10, 791–800. [Google Scholar] [CrossRef]
- Csukasi, F.; Donaire, L.; Casañal, A.; Martínez-Priego, L.; Botella, M.A.; Medina-Escobar, N.; Llave, C.; Valpuesta, V. Two strawberry miR159 family members display developmental-specific expression patterns in the fruit receptacle and cooperatively regulate Fa-GAMYB. New Phytol. 2012, 195, 47–57. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Jogaiah, S.; Zhang, W.; Abdelrahman, M.; Fang, J.G. Spatio-temporal expression of miRNA159 family members and their GAMYB target gene during the modulation of gibberellin-induced grapevine parthenocarpy. J. Exp. Bot. 2018, 69, 3639–3650. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B. MicroRNA: A new target for improving plant tolerance to abiotic stress. J. Exp. Bot. 2015, 66, 1749–1761. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.H.; Tian, X.; Li, Y.J.; Wu, C.A.; Zheng, C.C. Microarray-based analysis of stress-regulated microRNAs in Arabidopsis thaliana. RNA 2008, 14, 836–843. [Google Scholar] [CrossRef] [PubMed]
- Reyes, J.L.; Chua, N.H. ABA induction of miR159 controls transcript levels of two MYB factors during Arabidopsis seed germination. Plant J. 2007, 49, 592–606. [Google Scholar] [CrossRef]
- Pieczynski, M.; Marczewski, W.; Hennig, J.; Dolata, J.; Bielewicz, D.; Piontek, P.; Wyrzykowska, A.; Krusiewicz, D.; Strzelczyk-Zyta, D.; Konopka-Postupolska, D.; et al. Down-regulation of CBP80 gene expression as a strategy to engineer a drought-tolerant potato. Plant Biotechnol. J. 2013, 11, 459–469. [Google Scholar] [CrossRef]
- Zhang, T.; Zhao, Y.L.; Zhao, J.H.; Wang, S.; Jin, Y.; Chen, Z.Q.; Fang, Y.Y.; Hua, C.L.; Ding, S.W.; Guo, H.S. Cotton plants export microRNAs to inhibit virulence gene expression in a fungal pathogen. Nat. Plants 2016, 2, 16153. [Google Scholar] [CrossRef]
- Medina, C.; da Rocha, M.; Magliano, M.; Ratpopoulo, A.; Revel, B.; Marteu, N.; Magnone, V.; Lebrigand, K.; Cabrera, J.; Barcala, M.; et al. Characterization of microRNAs from Arabidopsis galls highlights a role for miR159 in the plant response to the root-knot nematode Meloidogyne incognita. New Phytol. 2017, 216, 882–896. [Google Scholar] [CrossRef]
| At Number | Score | Name | 5’-RACE | Degradome | miR159 OE | miR159ab | |
|---|---|---|---|---|---|---|---|
| 1 | AT4G37770 | 1.5 | ACS8 | [53] | [13] | ||
| 2 | AT2G32460 | 2 | MYB101 | [15,17,53] | [13] | ||
| 3 | AT3G60460 | 2 | DUO1 | [15,17,53] | |||
| 4 | AT2G26950 | 2 | MYB104 | ||||
| 5 | AT4G26930 | 2 | MYB97 | ||||
| 6 | AT5G06100 | 2.5 | MYB33 | [15,23] | [25,42] | [52] | [14,41] |
| 7 | AT3G11440 | 2.5 | MYB65 | [23] | [25,42] | [14,41] | |
| 8 | AT2G34010 | 2.5 | MRG1 | [53] | [42] | ||
| 9 | AT2G21600 | 2.5 | RER1B | ||||
| 10 | AT5G55020 | 2.5 | MYB120 | [15] | [13] | ||
| 11 | AT4G27330 | 2.5 | SPL | ||||
| 12 | AT5G27395 | 2.5 | Tim44-related | ||||
| 13 | AT3G61740 | 3 | SDG14, ATX3 | ||||
| 14 | AT1G29010 | 3 | MRG-LIKE | ||||
| 15 | AT4G31240 | 3 | NRX2 | ||||
| 16 | AT2G26960 | 3 | MYB81 | [15] | |||
| 17 | AT2G22810 | 3 | ACS4 | ||||
| 18 | AT3G08850 | 3 | RAPTOR1B | ||||
| 19 | AT5G55930 | 3.5 | OPT1 | [13] | [13] | [41] | |
| 20 | AT2G44450 | 3.5 | beta gluc 15 |
| Species | Approach | Phenotype | Ref. |
|---|---|---|---|
| Arabidopsis | T-DNA mir159ab mutant | Pleiotropic defects, stunted growth, curled leaves, reduced apical dominance. | [14] |
| Arabidopsis | T-DNA mir159c mutant | none | [15] |
| Arabidopsis | T-DNA mir159abc mutant | Perturbed fertilization | [51] |
| Arabidopsis | MIM159 mimic–loss-of-function. | Pleiotropic defects, stunted growth, curled leaves, defective sepals, petals and anthers. | [58,59] |
| Arabidopsis (Col-0) | miR159a overexpression | Male sterility | [13] |
| Arabidopsis (Ler) | miR159a overexpression | Male sterility, delayed flowering-time | [52] |
| Arabidopsis | T-DNA myb33.myb65 mutant | Male sterile | [43] |
| Arabidopsis | T-DNA myb33 mutant | Altered phase change | [60] |
| Rice | STTM159 mimic–loss-of-function. | Stunted growth, curled leaves, smaller seeds | [61,62] |
| Rice | miR159 overexpression | Delayed heading, shorten internode I, malformed flowers, male sterility. | [63] |
| Rice | gamyb-1 insertion mutant | Male sterility | [64] |
| Barley | miR159 overexpression | Male sterility | [65] |
| Wheat | miR159 overexpression | Delayed heading, male sterility, increased tillering. | [66] |
| Gloxinia | MIM159 mimic loss-of-function, miR159 over-expression (OE). | Accelerated flowering (MIM159) or delayed flowering (miR159 OE) | [67] |
| Tomato | miR159 overexpression | Fruit set, parthenocarpy, ovule development, seedless fruits | [35] |
| Cucumber | RNAi against GAMYB | Altered ratio of male to female flowers | [68] |
| Strawberry | RNAi against GAMYB | Inhibition of receptacle ripening | [69] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Millar, A.A.; Lohe, A.; Wong, G. Biology and Function of miR159 in Plants. Plants 2019, 8, 255. https://doi.org/10.3390/plants8080255
Millar AA, Lohe A, Wong G. Biology and Function of miR159 in Plants. Plants. 2019; 8(8):255. https://doi.org/10.3390/plants8080255
Chicago/Turabian StyleMillar, Anthony A., Allan Lohe, and Gigi Wong. 2019. "Biology and Function of miR159 in Plants" Plants 8, no. 8: 255. https://doi.org/10.3390/plants8080255





