Structure and Function of the LmbE-like Superfamily
Abstract
:1. Introduction
2. LmbE-like Enzymes
| Enzyme | UniProtKB Entry a | PIG-L Domain b | Metal Ligands c | Proposed GBC c |
|---|---|---|---|---|
| PIG-L | O35790 | 44–168 | His49-Asp52-His157 | Asp51 |
| MshB | O50426 | 8–158 | His13-Asp16-His147 | Asp15 |
| BshB | Q81ST8 | 7–124 | His12-Asp15-His113 | Asp14 |
| Teicoplanin deacetylase | Q6ZZJ1 | 11–175 | His16-Asp19-His164 | Asp18 |
| A40926 deacetylase | Q7WZ70 | 11–172 | His16-Asp19-His161 | Asp18 |
| TT1542 | Q84BR2 | 5–122 | His10-Asp13-His111 | Asp12 |
| Mca | L7N5N8 | 7–152 | His12-Asp15-His142 | Asp14 |
| Bca | Q81WT0 | 8–138 | His13-Asp16-His127 | Asp15 |
| TkDAC | Q6F4N1 | 35–162 | His40-Asp43-His151 | Asp42 |
| BtrD | Q4H4F3 | 9–177 | His14-Asp17-His166 | Asp16 |
| Enzyme | PDB accession code | Bound Ligands a | Bound Metal Ion | Reference |
|---|---|---|---|---|
| MshB | 1Q74 | PE4 | Zn | [4] |
| 1Q7T | BOG, SO4 | - | [12] | |
| 4EWL | Act, GOL, PE4 | Zn | [5] | |
| BshB (BC1534) | 2IXD | Act | Zn | [13] |
| Teicoplanin deacetylase | 3DFF | GOL, PG4 | Zn | [6] |
| 3DFK | DKA | Zn | [6] | |
| 3DFM | SO4 | Zn | [6] | |
| 2X9L | BOG | Zn | [14] | |
| 2XAD | BMA, T55, GCS, NAG | - | [14] | |
| A40926 deacetylase | 3DFI | - | Zn | [6] |
| TT1542 | 1UAN | - | - | [15] |
2.1. N-acetyl-d-glucosaminylphosphatidylinositol Deacetylase (PIG-L)
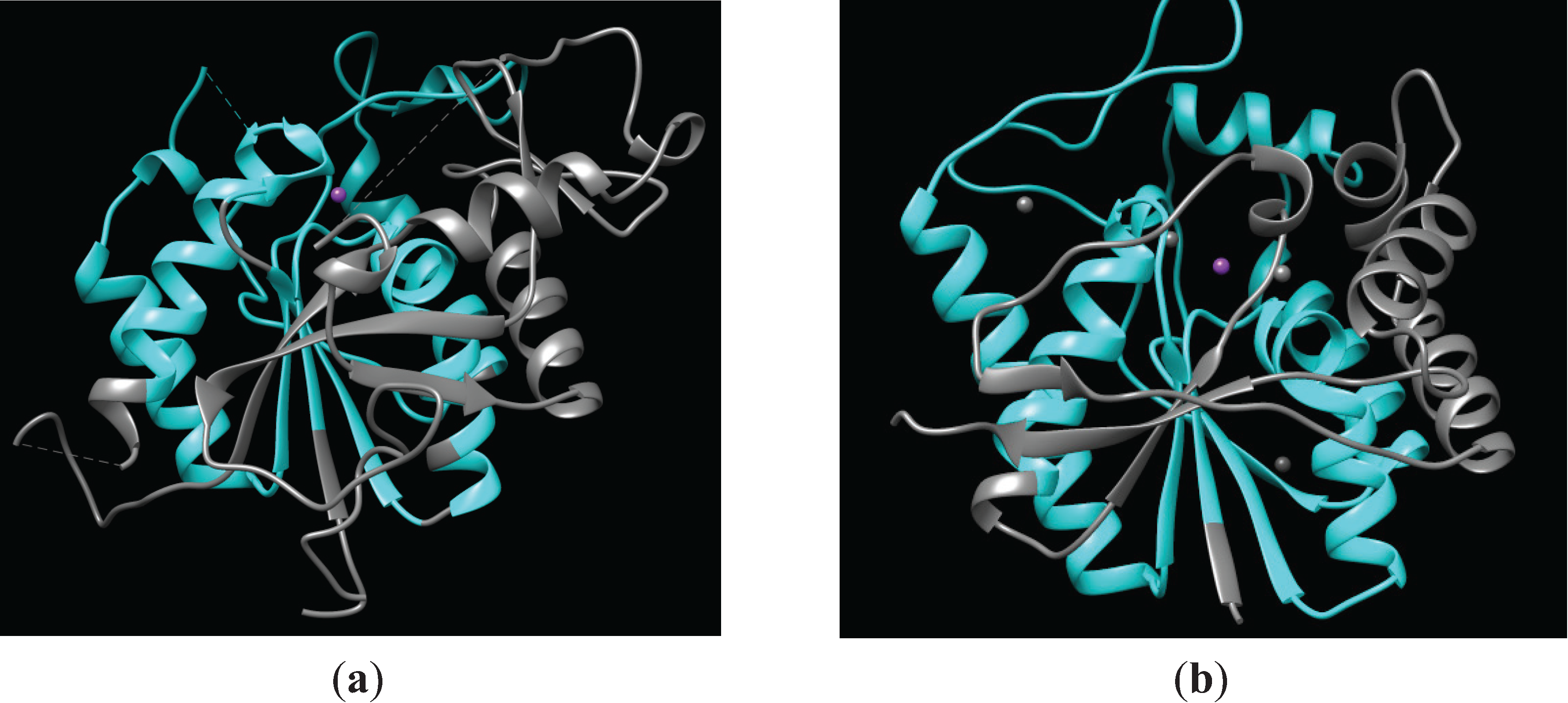
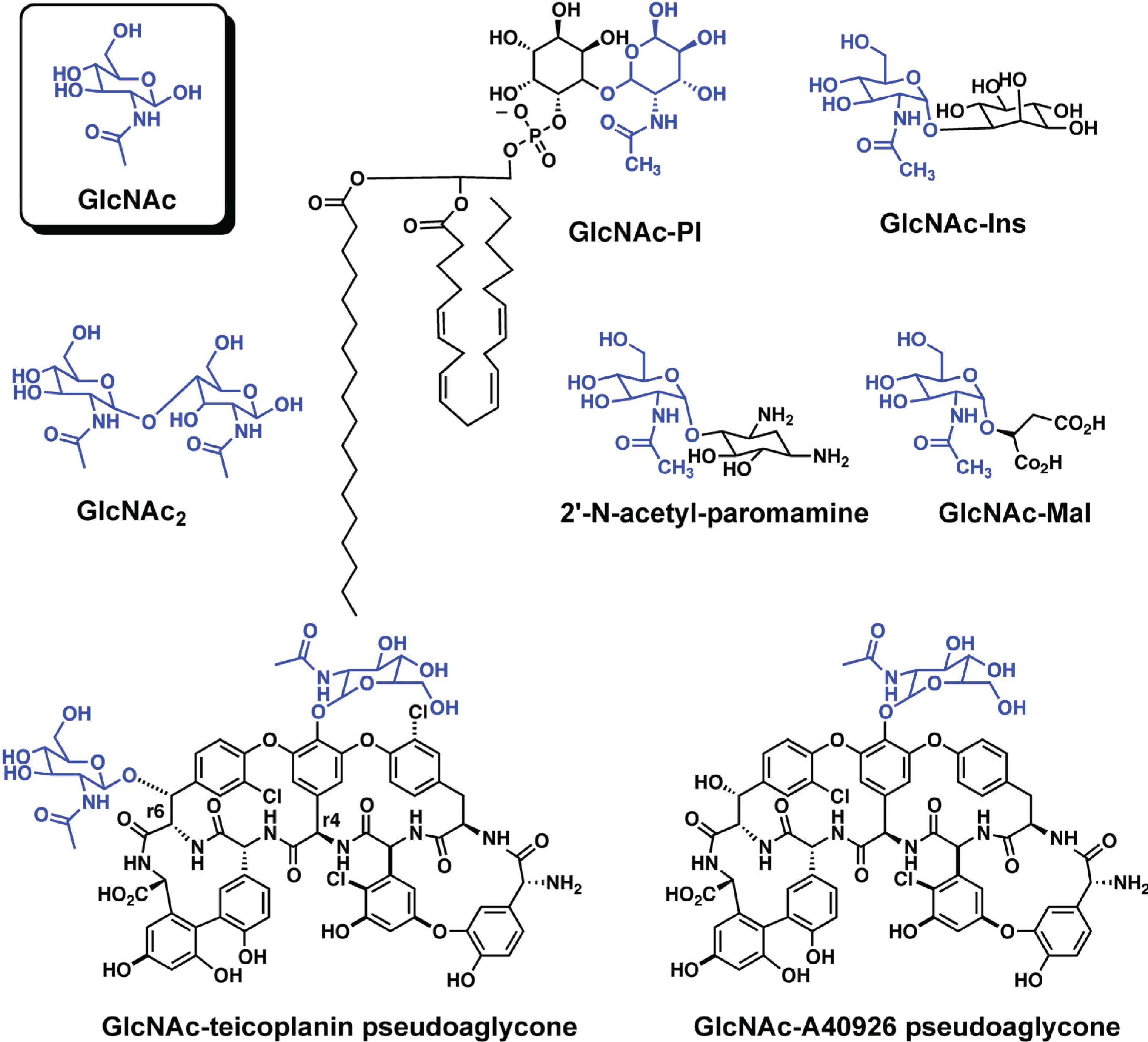
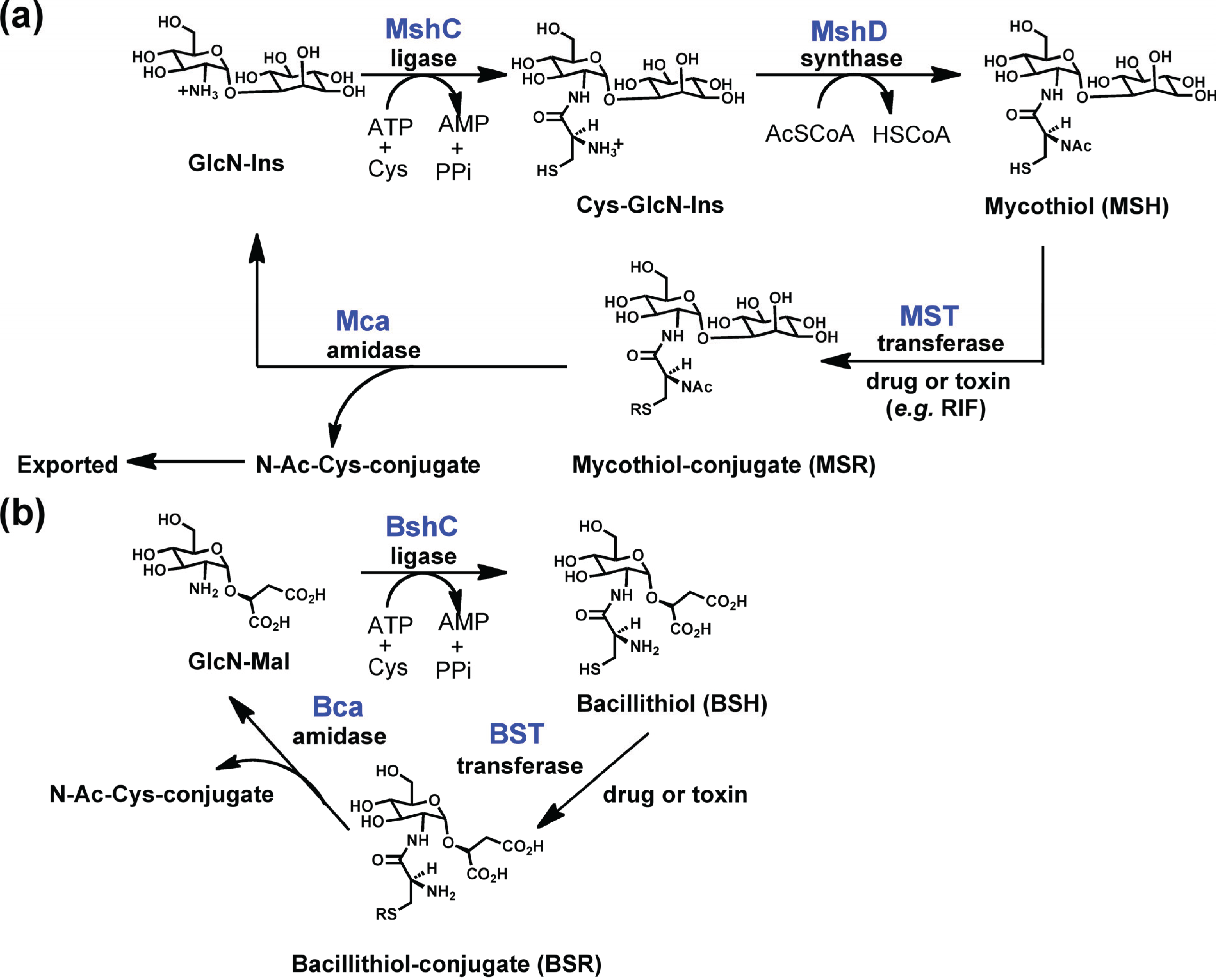
2.1.1. Structure of PIG-L
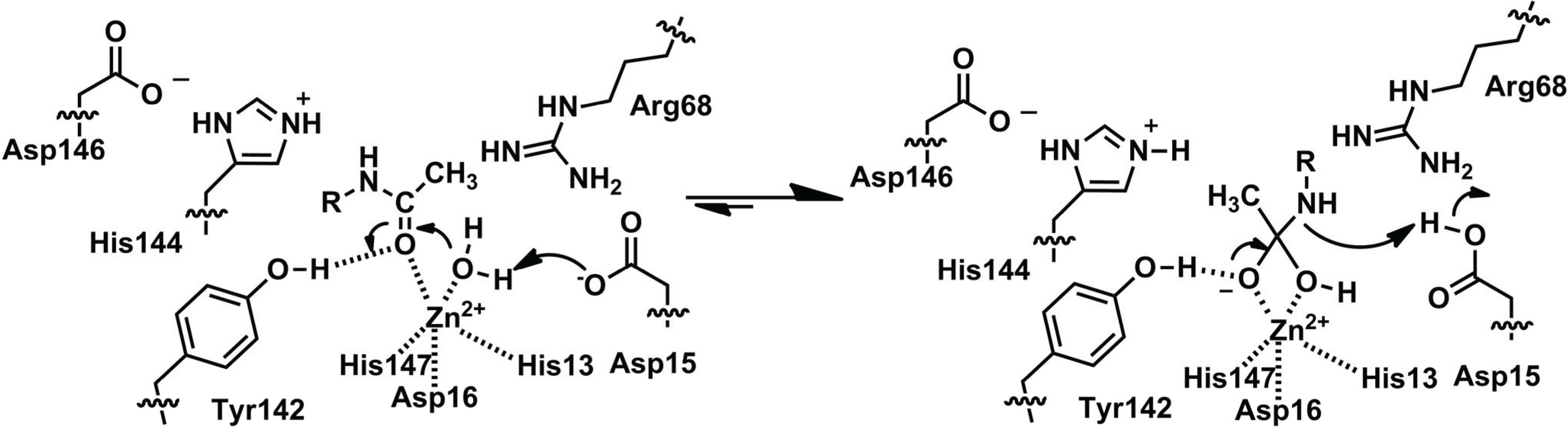
2.1.2. Function of PIG-L
2.2. N-acetyl-1-d-myo-inosityl-2-amino-2-deoxy-α-d-glucopyranoside Deacetylase (MshB)
2.2.1. Structure of MshB
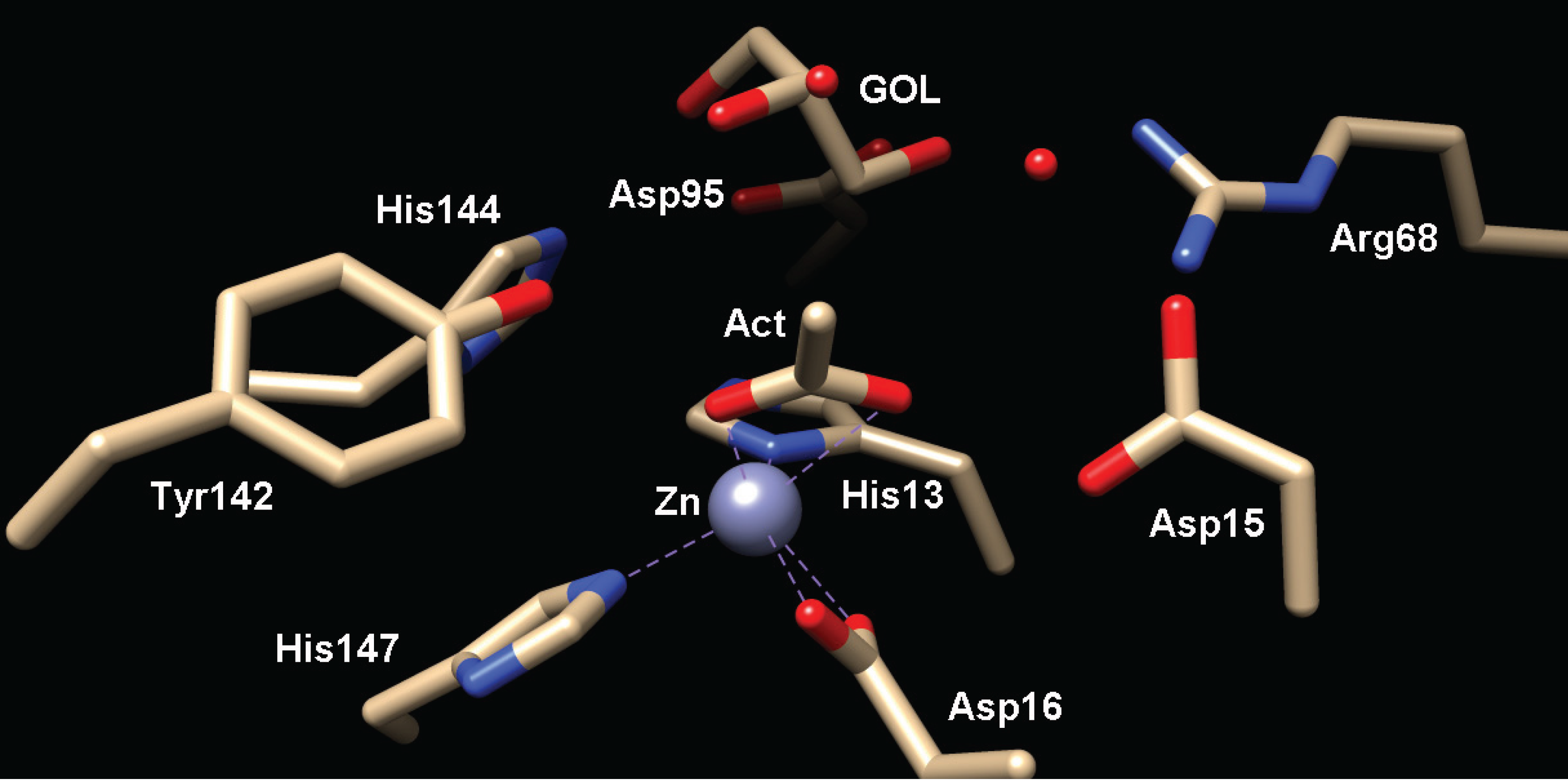
2.2.2. Function of MshB
2.3. N-acetylglucosamine-maleate Deacetylase (BshB)
2.3.1. Structure of BshB
2.3.2. Function of BshB
2.4. Teicoplanin Deacetylase and A40926 Deacetylase
2.4.1. Structure of Teicoplanin Deacetylase and A40926 Deacetylase
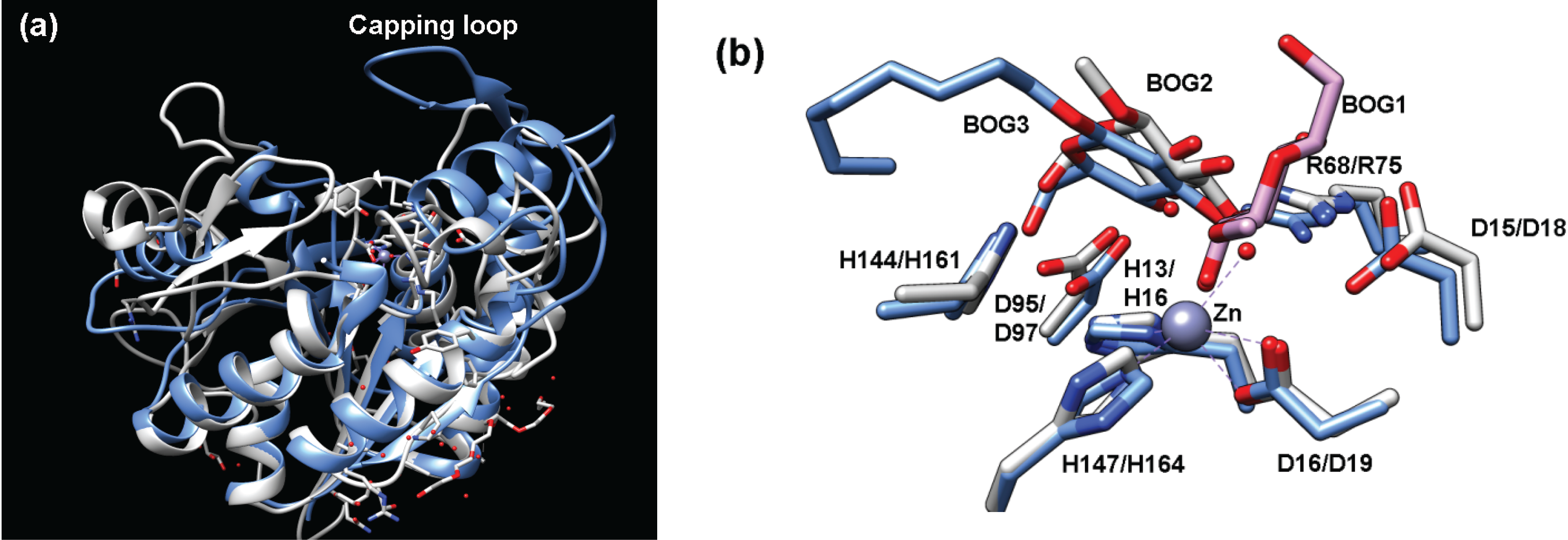
2.4.2. Function of Teicoplanin Deacetylase and A40926 Deacetylase
2.5. TT1542
Structure of TT1542
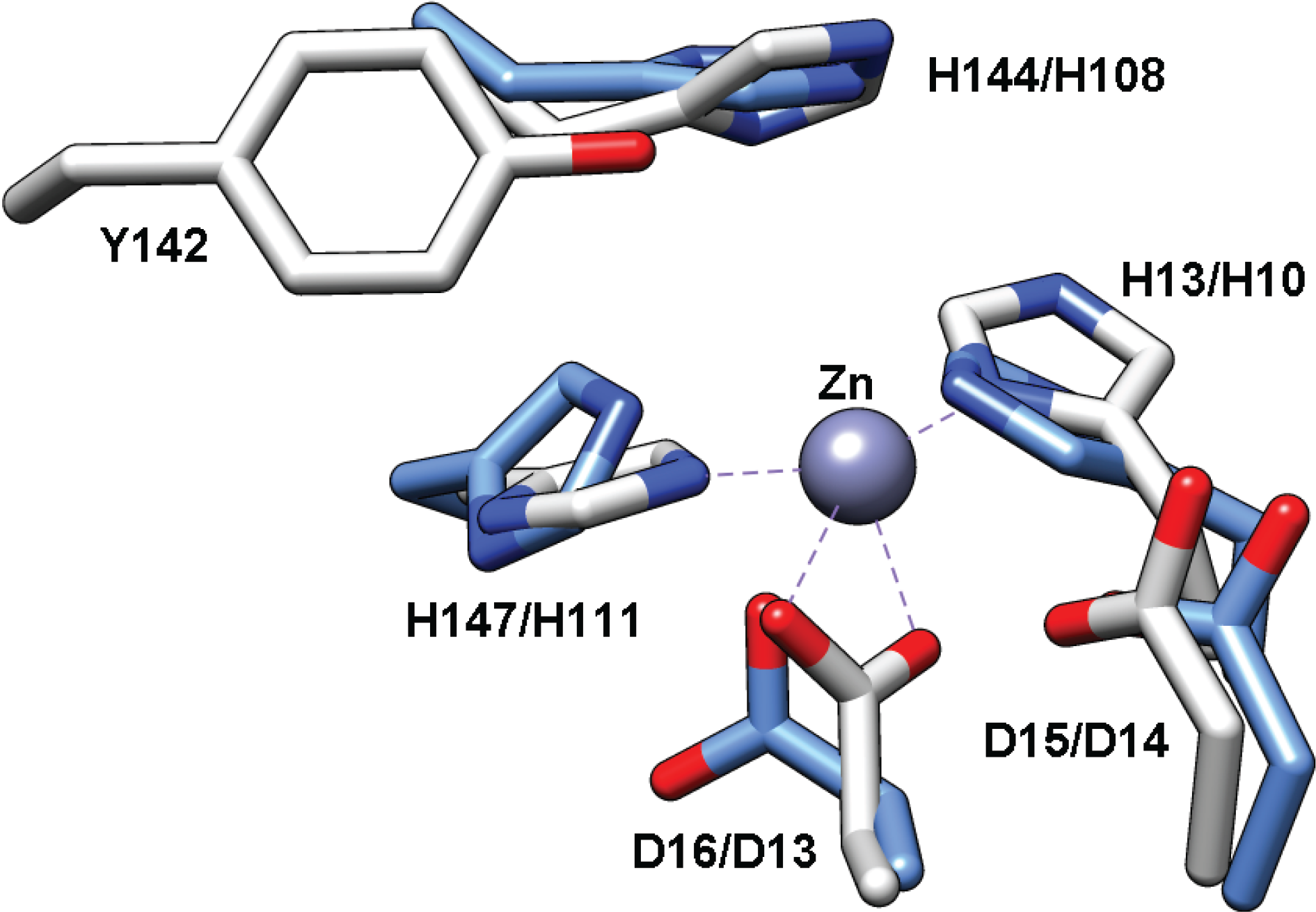
2.6. Mycothiol-Conjugate Amidase (Mca) and Bacillithiol-conjugate Amidase (Bca)
2.6.1. Structure of Mca and Bca
2.6.2. Function of Mca and Bca
2.7. Other Enzymes
3. Implications and Conclusions
Abbreviations
| Act | acetate |
| Bca | bacillithiol-conjugate amidase |
| BMA | β-d-Mannose |
| BOG | β-octylglucoside |
| BSH | bacillithiol |
| BshB | N-acetylglucosamine-maleate deacetylase |
| DKA | decanoic acid |
| GAC | general acid catalyst |
| GABC | general acid-base catalyst |
| GalNAc | acetylgalactosaminyltransferase |
| GBC | general base catalyst |
| GCS | d-glucosamine |
| GlcNAc | N-acetylglucosamine |
| GlcNAc-Ins | N-acetyl-1-d-myo-inosityl-2-amino-2-deoxy-α-d-glucopyranoside GlcNAc-PI, N-acetyl-glucosaminylphosphatidylinositol |
| GlcNAc-Mal | N-acetylglucosamine-maleate |
| GlcN-Ins | 1-d-myo-inosityl-2-amino-2-deoxy-α-d-glucopyranoside |
| GlcN-Mal | glucosamine-maleate |
| GlcN-PI | glucosaminylphosphatidylinositol |
| GOL | glycerol |
| GPGTF | guanylyl-2',5-phosphoguanosine transferase |
| GPI | glycosylphosphatidylinositol |
| LpxC | UDP-3-O-(R-3-hydroxymyristoyl)-N-acetylglucosamine deacetylase |
| Mca | mycothiol-conjugate amidase |
| MRSA | methicillin-resistant Staphylococcus aureus |
| MSH | mycothiol |
| MshB | N-acetyl-1-d-myo-2-amino-2-deoxy-α-d-glucopyranoside deacetylase |
| NAG | N-acetyl-d-glucosamine |
| PE4 | polyethylene glycol 4000 |
| PG4 | tetraethylene glycol |
| PIG-L | N-acetyl-d-glucosaminylphosphatidylinositol deacetylase |
| TB | tuberculosis |
| TkDAC | Thermococcus kodakaraenis diacetylchitobiose deacetylase |
| TT55 | 8-methylnonanoic acid; VRE, vancomycin-resistant Enterococcus |
| VRSA | vancomycin-resistant Staphylococcus aureus |
| WT | wild-type |
Author Contributions
Conflicts of Interest
References
- Hernick, M.; Fierke, C. Mechanisms of metal-dependent hydrolases in metabolism, reference module in chemistry, molecular sciences and chemical engineering. Available online: http://mrw.elsevier.com/chem/ (accessed on 5 January 2014).
- Punta, M.; Coggill, P.C.; Eberhardt, R.Y.; Mistry, J.; Tate, J.; Boursnell, C.; Pang, N.; Forslund, K.; Ceric, G.; Clements, J.; et al. The Pfam protein families database. Nucleic Acids Res. 2012, 40, D290–D301. [Google Scholar] [CrossRef]
- Nakamura, N.; Inoue, N.; Watanabe, R.; Takahashi, M.; Takeda, J.; Stevens, V.L.; Kinoshita, T. Expression cloning of PIG-L, a candidate N-acetylglucosaminyl-phosphatidylinositol deacetylase. J. Biol. Chem. 1997, 272, 15834–15840. [Google Scholar]
- Maynes, J.T.; Garen, C.; Cherney, M.M.; Newton, G.; Arad, D.; Av-Gay, Y.; Fahey, R.C.; James, M.N.G. The crystal structure of 1-d-myo-inosityl-2-acetamido-2-deoxy-alpha-d-glucopyranoside deacetylase (MshB) from Mycobacterium tuberculosis reveals a zinc hydrolase with a lactate dehydrogenase fold. J. Biol. Chem. 2003, 278, 47166–47170. [Google Scholar]
- Broadley, S.G.; Gumbart, J.C.; Weber, B.W.; Marakalala, M.J.; Steenkamp, D.J.; Sewell, B.T. A new crystal form of MshB from Mycobacterium tuberculosis with glycerol and acetate in the active site suggests the catalytic mechanism. Acta Crystallogr. D Biol. Crystallogr. 2012, 68, 1450–1459. [Google Scholar] [CrossRef]
- Zou, Y.; Brunzelle, J.S.; Nair, S.K. Crystal structures of lipoglycopeptide antibiotic deacetylases: Implications for the biosynthesis of A40926 and teicoplanin. Chem. Biol. 2008, 15, 533–545. [Google Scholar] [CrossRef]
- Newton, G.L.; Buchmeier, N.; Fahey, R.C. Biosynthesis and functions of mycothiol, the unique protective thiol of actinobacteria. Microbiol. Mol. Biol. Rev. 2008, 72, 471–494. [Google Scholar] [CrossRef]
- Hernick, M. Mycothiol, a target for potentiation of rifampin and other antibiotics against M. tuberculosis. Expert Rev. Anti Infect. Ther. 2013, 11, 49–67. [Google Scholar] [CrossRef]
- Gaballa, A.; Newton, G.L.; Antelmann, H.; Parsonage, D.; Upton, H.; Rawat, M.; Claiborne, A.; Fahey, R.C.; Helmann, J.D. Biosynthesis and functions of bacillithiol, a major low-molecular-weight thiol in Bacilli. Proc. Natl. Acad. Sci. USA 2010, 107, 6482–6486. [Google Scholar] [CrossRef]
- Urbaniak, M.D.; Ferguson, M.A.J. The GlcNAc-PI de-N-acetylase: Structure, function, and activity. Enzymes 2009, 26, 49–64. [Google Scholar] [CrossRef]
- Magrane, M.; Consortium, U. UniProt Knowledgebase: A hub of integrated protein data. Database 2011. [Google Scholar] [CrossRef]
- McCarthy, A.A.; Peterson, N.A.; Knijff, R.; Baker, E.N. Crystal structure of MshB from Mycobacterium tuberculosis, a deacetylase involved in mycothiol biosynthesis. J. Mol. Biol. 2004, 335, 1131–1141. [Google Scholar] [CrossRef]
- Fadouloglou, V.E.; Deli, A.; Glykos, N.M.; Psylinakis, E.; Bouriotis, V.; Kokkinidis, M. Crystal structure of the BcZBP, a zinc-binding protein from Bacillus cereus—Functional insights from structural data. FEBS J. 2007, 274, 3044–3054. [Google Scholar] [CrossRef]
- Chan, H.-C.; Huang, Y.-T.; Lyu, S.-Y.; Huang, C.-J.; Li, Y.-S.; Liu, Y.-C.; Chou, C.-C.; Tsai, M.-D.; Li, T.-L. Regioselective deacetylation based on teicoplanin-complexed Orf2* crystal structures. Mol. BioSyst. 2011, 7, 1224–1231. [Google Scholar] [CrossRef] [Green Version]
- Handa, N.; Terada, T.; Kamewari, Y.; Hamana, H.; Tame, J.R.H.; Park, S.-Y.; Kinoshita, K.; Ota, M.; Nakamura, H.; Kuramitsu, S.; et al. Crystal structure of the conserved protein TT1542 from Thermus thermophilus HB8. Protein Sci. 2003, 12, 1621–1632. [Google Scholar]
- Urbaniak, M.D.; Crossman, A.; Chang, T.; Smith, T.K.; van Aalten, D.M.F.; Ferguson, M.A.J. The N-Acetyl-d-glucosaminylphosphatidylinositol de-N-acetylase of glycosylphosphatidylinositol biosynthesis is a zinc metalloenzyme. J. Biol. Chem. 2005, 280, 22831–22838. [Google Scholar]
- Ashraf, M.; Yadav, B.; Perinthottathil, S.; Kumar, K.S.; Vats, D.; Muthuswami, R.; Komath, S.S. N-Acetyl-d-glucosaminylphosphatidylinositol de-N-acetylase from Entamoeba histolytica: Metal alters catalytic rates but not substrate affinity. J. Biol. Chem. 2011, 286, 2543–2549. [Google Scholar]
- Huang, X.; Kocabas, E.; Hernick, M. The activity and cofactor preferences of N-acetyl-1-d-myo-inositol-2-amino-2-deoxy-α--d-glucopyranoside deacetylase (MshB) change depending on environmental conditions. J. Biol. Chem. 2011, 286, 20275–20282. [Google Scholar] [CrossRef]
- Rogers, I.L.; Gammon, D.W.; Naidoo, K.J. Conformational preferences of plumbagin with phenyl-1-thioglucoside conjugates in solution and bound to MshB determined by aromatic association. Carbohydr. Res. 2013, 371, 52–60. [Google Scholar] [CrossRef]
- Huang, X.; Hernick, M. Automated docking studies provide insights into molecular determinants of ligand recognition by N-acetyl-1-d-myo-inosityl-2-amino-2-deoxy-α-d-glucopyranoside deacetylase (MshB). Biopolymers. 2014, 101, 406–417. [Google Scholar] [CrossRef]
- Huang, X.; Hernick, M. Examination of mechanism of N-Acetyl-1-d-myo-inosityl-2-amino-2-deoxy-α-d-glucopyranoside deacetylase (MshB) reveals unexpected role for dynamic tyrosine. J. Biol. Chem. 2012, 287, 10424–10434. [Google Scholar] [CrossRef]
- Newton, G.L.; Ko, M.; Ta, P.; Av-Gay, Y.; Fahey, R.C. Purification and characterization of Mycobacterium tuberculosis 1d-myo-inosityl-2-acetamido-2-deoxy-a-d-glucopyranoside deacetylase, MshB, a mycothiol biosynthetic enzyme. Protein Expr. Purif. 2006, 47, 542–550. [Google Scholar] [CrossRef]
- Fang, Z.; Roberts, A.A.; Weidman, K.; Sharma, S.V.; Claiborne, A.; Hamilton, C.J.; dos Santos, P.C. Cross-functionalities of Bacillus deacetylases involved in bacillithiol biosynthesis and bacillithiol-S-conjugate detoxification pathways. Biochem. J. 2013, 454, 239–247. [Google Scholar]
- Fadouloglou, V.E.; Stavrakoudis, A.; Bouriotis, V.; Kokkinidis, M.; Glykos, N.M. Molecular dynamics simulations of BcZBP, a deacetylase from Bacillus cereus: Active site loops determine substrate accessibility and specificity. J. Chem. Theory Comput. 2009, 5, 3299–3311. [Google Scholar] [CrossRef]
- Deli, A.; Koutsioulis, D.; Fadouloglou, V.E.; Spiliotopoulou, P.; Balomenou, S.; Arnaouteli, S.; Tzanodaskalaki, M.; Mavromatis, K.; Kokkinidis, M.; Bouriotis, V. LmbE proteins from Bacillus cereus are de-N-acetylases with broad substrate specificity and are highly similar to proteins in Bacillus anthracis. FEBS J. 2010, 277, 2740–2753. [Google Scholar] [CrossRef]
- Truman, A.W.; Robinson, L.; Spencer, J.B. Identification of a deacetylase involved in the maturation of teicoplanin. ChemBioChem 2006, 7, 1670–1675. [Google Scholar] [CrossRef]
- Ho, J.-Y.; Huang, Y.-T.; Wu, C.-J.; Li, Y.-S.; Tsai, M.-D.; Li, T.-L. Glycopeptide biosynthesis: Äâ Dbv21/Orf2* from dbv/tcp gene clusters Are N-Ac-Glm teicoplanin pseudoaglycone deacetylases and Orf15 from cep gene cluster is a Glc-1-P thymidyltransferase. J. Am. Chem. Soc. 2006, 128, 13694–13695. [Google Scholar]
- Newton, G.L.; Leung, S.S.; Wakabayashi, J.I.; Rawat, M.; Fahey, R.C. The DinB superfamily includes novel mycothiol, bacillithiol, and glutathione S-transferases. Biochemistry 2011, 50, 10751–10760. [Google Scholar] [CrossRef]
- Newton, G.L.; Av-Gay, Y.; Fahey, R.C. A novel mycothiol-dependent detoxification pathway in mycobacteria involving mycothiol S-conjugate amidase. Biochemistry 2000, 39, 10739–10746. [Google Scholar] [CrossRef]
- Steffek, M.; Newton, G.L.; Av-Gay, Y.; Fahey, R.C. Characterization of Mycobacterium tuberculosis mycothiol S-conjugate amidase. Biochemistry 2003, 42, 12067–12076. [Google Scholar] [CrossRef]
- Truman, A.W.; Huang, F.; Llewellyn, N.M.; Spencer, J.B. Characterization of the enzyme BtrD from Bacillus circulans and revision of its functional assignment in the biosynthesis of butirosin. Angew. Chem. Int. Ed. 2007, 46, 1462–1464. [Google Scholar] [CrossRef]
- Tanaka, T.; Fukui, T.; Fujiwara, S.; Atomi, H.; Imanaka, T. Concerted action of diacetylchitobiose deacetylase and Exo-β-d-glucosaminidase in a novel chitinolytic pathway in the hyperthermophilic archaeon Thermococcus kodakaraensis KOD1. J. Biol. Chem. 2004, 279, 30021–30027. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Viars, S.; Valentine, J.; Hernick, M. Structure and Function of the LmbE-like Superfamily. Biomolecules 2014, 4, 527-545. https://doi.org/10.3390/biom4020527
Viars S, Valentine J, Hernick M. Structure and Function of the LmbE-like Superfamily. Biomolecules. 2014; 4(2):527-545. https://doi.org/10.3390/biom4020527
Chicago/Turabian StyleViars, Shane, Jason Valentine, and Marcy Hernick. 2014. "Structure and Function of the LmbE-like Superfamily" Biomolecules 4, no. 2: 527-545. https://doi.org/10.3390/biom4020527




