Recent Advances and Future Perspectives in Microbial Phototrophy in Antarctic Sea Ice
Abstract
:1. Introduction
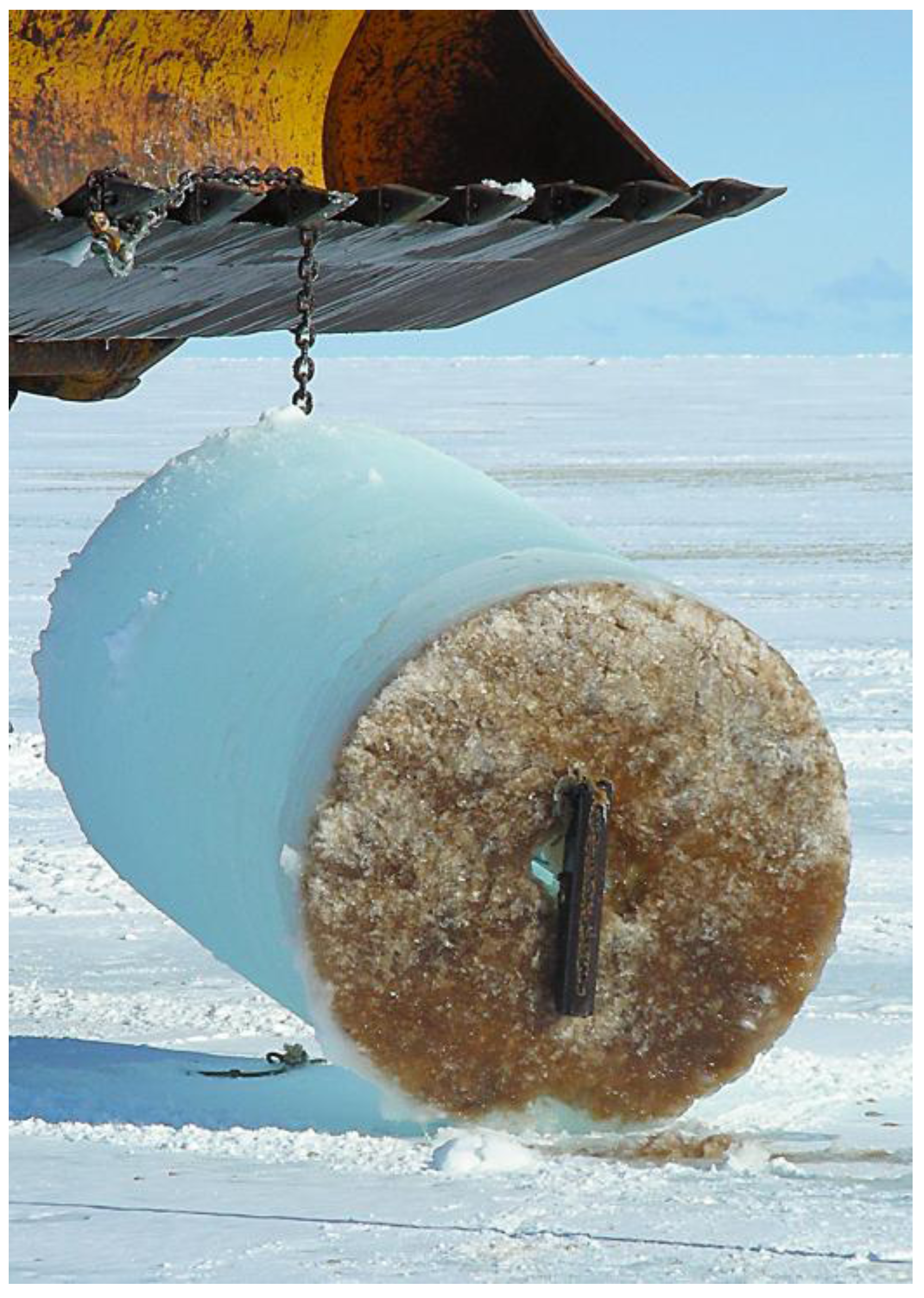

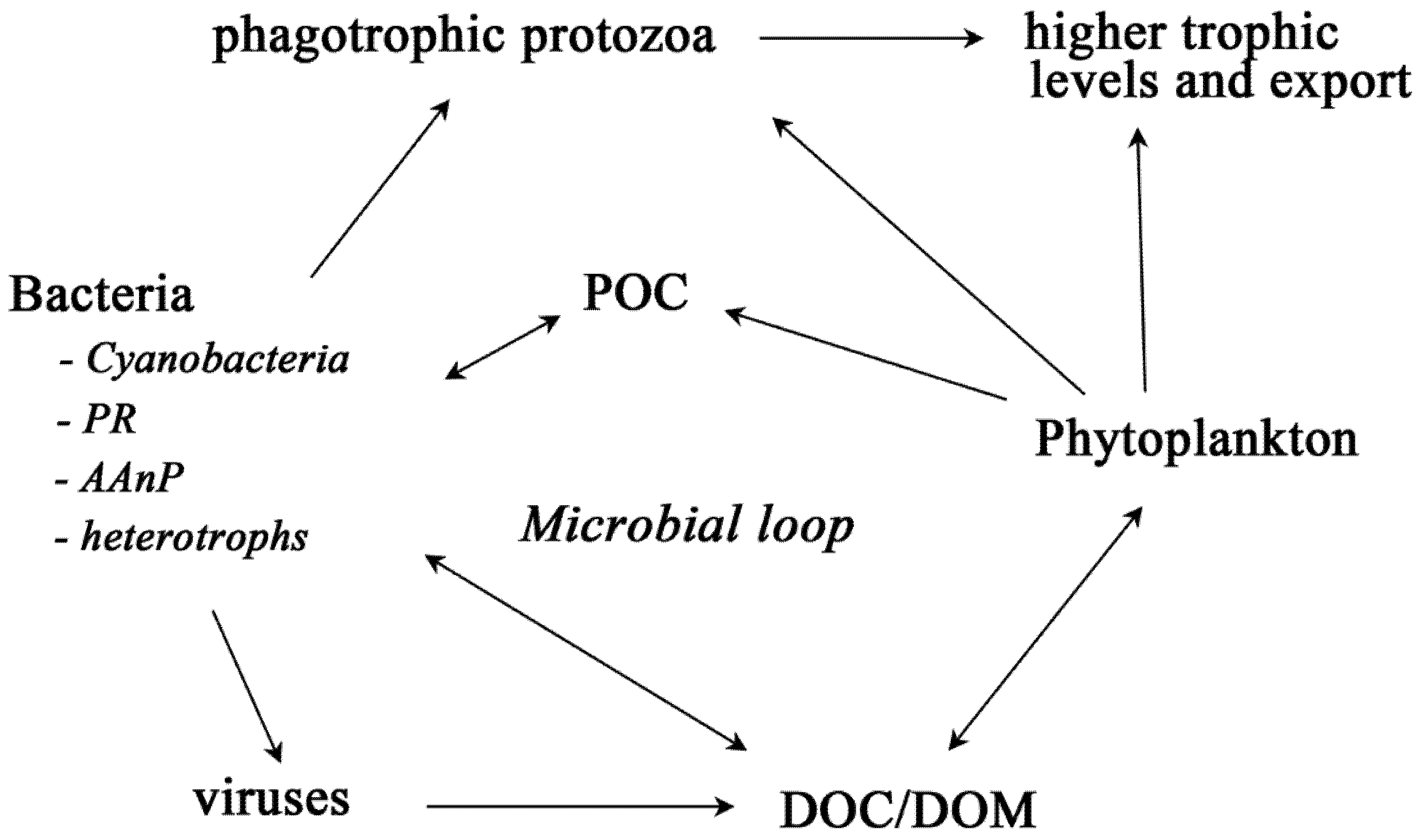
2. Bacteria with Light-Harvesting Capabilities
2.1. Cyanobacteria
2.2. Bacteriochlorophyll
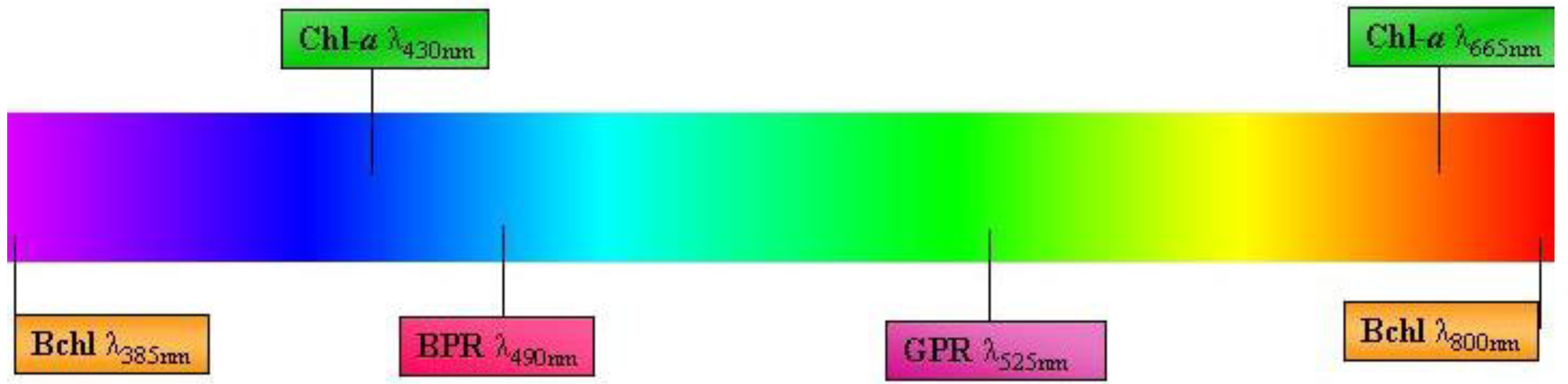

2.3. Proteorhodopsin
3. Future Research and the Significance of Light-Harvesting Pigments for Antarctic Sea Ice
4. Concluding Remarks
Acknowledgments
References
- Xu, J. Microbial ecology in the age of genomics and metagenomics: Concepts, tools, and recent advances. Mol. Ecol. 2006, 15, 1713–1731. [Google Scholar] [CrossRef]
- Junge, K.; Imhoff, F.; Staley, T.; Deming, J.W. Phylogenetic diversity of numerically important Arctic sea-ice bacteria cultured at subzero temperature. Microb. Ecol. 2002, 43, 315–328. [Google Scholar] [CrossRef]
- Nagy, M.L.; Pérez, A.; Garcia-Pichel, F. The prokaryotic diversity of biological soil crusts in the Sonoran Desert (Organ Pipe Cactus National Monument, AZ). FEMS Microb. Ecol. 2005, 54, 233–245. [Google Scholar]
- McCliment, E.A.; Voglesonger, K.M.; O’Day, P.A.; Dunn, E.E.; Holloway, J.R.; Cary, S.C. Colonization of nascent, deep-sea hydrothermal vents by a novel Archaeal and Nanoarchaeal assemblage. Environ. Microbiol. 2006, 8, 114–125. [Google Scholar] [CrossRef]
- Soo, R.M.; Wood, S.A.; Grzymski, J.J.; McDonald, I.R.; Cary, S.C. Microbial biodiversity of thermophilic communities in hot mineral soils of Tramway Ridge, Mount Erebus, Antarctica. Environ. Microbiol. 2009, 11, 715–728. [Google Scholar] [CrossRef]
- Pomeroy, L.R.; Williams, P.J.I.; Azam, F.; Hobbie, J.E. The microbial loop. Oceanography 2007, 20, 28–33. [Google Scholar] [CrossRef]
- Pace, N.R. A molecular view of microbial diversity and the biosphere. Science 1997, 276, 734–740. [Google Scholar] [CrossRef]
- Pedrós-Alió, C. Marine microbial diversity: Can it be determined? Trends Microbiol. 2006, 14, 257–263. [Google Scholar]
- Delille, D. Marine bacterioplankton at the Weddell Sea ice edge, distribution of psychrophilic and psychrotrophic populations. Polar Biol. 1992, 12, 205–210. [Google Scholar]
- Legendre, L.; Ackley, S.F.; Dieckmann, G.S.; Gulliksen, B.; Horner, R.; Hoshiai, T.; Melnikov, I.A.; Reeburgh, W.S.; Spindler, M.; Sullivan, C.W. Ecology of sea ice biota. 2. Global significance. Polar Biol. 1992, 12, 429–444. [Google Scholar]
- Kottmeier, S.T.; Sullivan, C.W. Sea ice microbial communities (SIMCO) 9. Effects of temperature and salinity on rates of metabolism and growth of autotrophs and heterotrophs. Polar Biol. 1988, 8, 293–304. [Google Scholar] [CrossRef]
- McMinn, A.; Ashworth, C.; Ryan, K.G. In situ net primary productivity of an Antarctic fast ice bottom algal community. Aquat. Microb. Ecol. 2000, 21, 177–185. [Google Scholar] [CrossRef]
- Ryan, K.G.; McMinn, A.; Mitchell, K.A.; Trenerry, L. Mycosporine-like amino acids in Antarctic sea ice algae, and their response to UVB radiation. Z. Naturforsch. C 2002, 57, 1–7. [Google Scholar]
- Ryan, K.G.; Cowie, R.O.M.; Liggins, E.; McNaughtan, D.; Martin, A.; Davy, S.K. The short-term effect of irradiance on the photosynthetic properties of Antarctic fast-ice microalgal communities. J. Phycol. 2009, 45, 1290–1298. [Google Scholar] [CrossRef]
- Ryan, K.G.; Tay, M.L.; Martin, A.; McMinn, A.; Davy, S.K. Chlorophyll fluorescence imaging analysis of the responses of Antarctic bottom-ice algae to light and salinity during melting. J. Exp. Mar. Biol. Ecol. 2011, 399, 156–161. [Google Scholar] [CrossRef]
- Arrigo, K.R.; Thomas, D.N. Large scale importance of sea ice biology in the Southern Ocean. Antarct. Sci. 2004, 16, 471–486. [Google Scholar] [CrossRef]
- Daly, K.L. Overwintering development, growth, and feeding of larval Euphausia superba in the Antarctic marginal ice zone. Limnol. Oceanogr. 1990, 35, 1564–1576. [Google Scholar] [CrossRef]
- Kottmeier, S.T.; Sullivan, C.W. Bacterial biomass and production in pack ice of Antarctic marginal ice edge zones. Deep-Sea Res. 1990, 37, 1311–1330. [Google Scholar] [CrossRef]
- Giesenhagen, H.C.; Detmer, A.E.; de Wall, J.; Weber, A.; Gradinger, R.R.; Jochem, F.J. How are Antarctic planktonic microbial food webs and algal blooms affected by melting of sea ice? Microcosm simulations. Aquat. Microb. Ecol. 1999, 20, 183–201. [Google Scholar] [CrossRef]
- Lizotte, M.P. The contribution of sea ice algae to Antarctic marine primary production. Am. Zool. 2001, 41, 57–73. [Google Scholar] [CrossRef]
- Delille, D.; Rosiers, C. Seasonal changes of Antarctic marine bacterioplankton and sea ice bacterial assemblages. Polar Biol. 1996, 16, 27–34. [Google Scholar]
- Azam, F.; Worden, A.Z. Microbes, molecules, and marine ecosystems. Science 2004, 303, 1622–1624. [Google Scholar] [CrossRef]
- Murray, A.E.; Grzymski, J.J. Diversity and genomics of Antarctic marine micro-organisms. Phil. Trans. R. Soc. B 2007, 362, 2259–2271. [Google Scholar] [CrossRef]
- Sullivan, C.W.; Palmisano, A.C. Sea ice microbial communities: distribution, abundance, and diversity of ice bacteria in McMurdo Sound, Antarctica, in 1980. Appl. Environ. Microbiol. 1984, 47, 788–795. [Google Scholar]
- Sullivan, C.W. Reciprocal interactions of the organisms and their environmen. In Sea Ice Biota; Horner, R., Ed.; Boca Raton Chemical Rubber Company: Boca Raton, FL, USA, 1985; pp. 160–171. [Google Scholar]
- Martin, A.; Hall, J.A.; O’Toole, R.; Davy, S.K.; Ryan, K.G. High single-cell metabolic activity in Antarctic sea ice bacteria. Aquat. Microb. Ecol. 2008, 52, 25–31. [Google Scholar] [CrossRef]
- Martin, A.; Hall, J.A.; Ryan, K.G. Low salinity and high-level UV-B radiation reduce single-cell activity in Antarctic sea ice bacteria. Appl. Environ. Microbiol. 2009, 75, 7570–7573. [Google Scholar]
- Brown, M.V.; Bowman, J.P. A molecular phylogenetic survey of sea-ice microbial communities. FEMS Microbiol. Ecol. 2001, 35, 267–275. [Google Scholar] [CrossRef]
- Junge, K.; Krembs, C.; Deming, J.W.; Stierle, A.; Eicken, H. A microscopic approach to investigate bacteria under in situ conditions in sea-ice samples. Ann. Glaciol. 2001, 33, 304–310. [Google Scholar] [CrossRef]
- Gentile, G.; Giuliano, L.; D’Auria, G.; Smedile, F.; Azzaro, M.; de Domenico, M.; Yakimov, M.M. Study of bacterial communities in Antarctic coastal waters by a combination of 16S rRNA and 16S rDNA sequencing. Environ. Microbiol. 2006, 8, 2150–2161. [Google Scholar] [CrossRef]
- Karl, D.M. Microbial oceanography: Paradigms, processes and promise. Nat. Rev. Microbiol. 2007, 5, 759–769. [Google Scholar] [CrossRef]
- Kumari, N.; Srivastava, A.K.; Bhargava, P.; Rai, L.C. Molecular approaches towards assessment of cyanobacterial biodiversity. Afr. J. Biotechnol. 2009, 8, 4284–4298. [Google Scholar]
- Madigan, M.T. Anoxygenic phototrophic bacteria from extreme environments. Photosynth. Res. 2003, 76, 157–171. [Google Scholar] [CrossRef]
- Lanyi, J.K. Bacteriorhodopsin. Annu. Rev. Physiol. 2004, 66, 665–688. [Google Scholar] [CrossRef]
- Bryant, D.A.; Frigaard, N.-U. Prokaryotic photosynthesis and phototrophy illuminated. Trends Microbiol. 2006, 14, 488–496. [Google Scholar] [CrossRef]
- Fuhrman, J.A.; Schwalbach, M.S.; Stingl, U. Proteorhodopsins: An array of physiological roles? Nat. Rev. Microbiol. 2008, 6, 488–494. [Google Scholar]
- Yurkov, V.; Csotonyi, J.T. New light on aerobic anoxygenic phototrophs. In The Purple Phototrophic Bacteria; Hunter, C.N., Daldal, F., Thurnauer, M.C., Beatty, J.T., Eds.; Springer Science: Dordrecht, The Netherlands, 2009; pp. 31–55. [Google Scholar]
- Kolber, Z.S.; Gerald, F.; Lang, A.S.; Beatty, J.T.; Blankenship, R.E.; VanDover, C.L.; Vetriani, C.; Koblizek, M.; Rathgeber, C.; Falkowski, P.G. Contribution of aerobic photoheterotrophic bacteria to the carbon cycle in the ocean. Science 2001, 292, 2492–2495. [Google Scholar]
- Rathgeber, C.; Beattyc, J.T.; Yurkov, V. Aerobic phototrophic bacteria: New evidence for the diversity, ecological importance and applied potential of this previously overlooked group. Photosynth. Res. 2004, 81, 113–128. [Google Scholar] [CrossRef]
- Jiang, H.; Dong, H.; Yu, B.; Lv, G.; Deng, S.; Wu, Y.; Dai, M.; Jiao, N. Abundance and diversity of aerobic anoxygenic phototrophic bacteria in saline lakes on the Tibetan plateau. FEMS Microbiol. Ecol. 2009, 67, 268–278. [Google Scholar] [CrossRef]
- Hawes, I.; Schwarz, A.-M.J. Absorption and utilization of irradiance by cyanobacterial mats in two ice-covered antarctic lakes with contrasting light climates. J. Phycol. 2001, 37, 5–15. [Google Scholar] [CrossRef]
- Taton, A.; Grubisic, S.; Brambilla, E.; de Wit, R.; Wilmotte, A. Cyanobacterial diversity in natural and artificial microbial mats of Lake Fryxell (McMurdo Dry Valleys, Antarctica): A morphological and molecular approach. Appl. Environ. Microbiol. 2003, 69, 5157–5169. [Google Scholar] [CrossRef]
- Jungblut, A.D.; Hawes, I.; Mountfort, D.; Hitzfeld, B.; Dietrich, D.R.; Burns, B.P.; Neilan, B.A. Diversity within cyanobacterial mat communities in variable salinity meltwater ponds of McMurdo Ice Shelf, Antarctica. Environ. Microbiol. 2005, 7, 519–529. [Google Scholar] [CrossRef]
- Vincent, W.F. Cold tolerance in cyanobacteria and life in the cryosphere. In Cellular Origin and Life in Extreme Habitats and Astrobiology; Seckbach, J., Ed.; Springer Science: Dordrecht, The Netherlands, 2007; pp. 289–301. [Google Scholar]
- Jungblut, A.D.; Lovejoy, C.; Vincent, W.F. Global distribution of cyanobacterial ecotypes in the cold biosphere. ISME J. 2010, 4, 191–202. [Google Scholar] [CrossRef]
- Tang, E.P.Y.; Vincent, W.F. Strategies of thermal adaptation by high-latitude cyanobacteria. New Phytol. 1999, 142, 315–323. [Google Scholar]
- Zakhia, F.; Jungblut, A.-D.; Taton, A.; Vincent, W.F.; Wilmotte, A. Cyanobacteria in cold ecosystems. In Psychrophiles: From Biodiversity to Biotechnology; Margesin, R., Schinner, F., Marx, J.-C., Gerday, C., Eds.; Springer Science: Heidelberg, Germany, 2008; pp. 121–135. [Google Scholar]
- Partensky, F.; Hess, W.R.; Vaulot, D. Prochlorococcus, a marine photosynthetic prokaryote of global significance. Microbiol. Mol. Biol. Rev. 1999, 63, 106–127. [Google Scholar]
- Chisholm, S.W.; Olson, R.J.; Zettler, E.R.; Goericke, R.; Waterbury, J.B.; Welschmeyer, N.A. A novel free-living prochlorophyte abundant in the oceanic euphotic zone. Nature 1988, 334, 340–343. [Google Scholar] [CrossRef]
- Murphy, L.S.; Haugen, E.M. The distribution and abundance of phototrophic ultraplankton in the North-Atlantic. Limnol. Oceanogr. 1985, 30, 47–58. [Google Scholar] [CrossRef]
- Marchant, H.J.; Davidson, A.T.; Wright, S.W. The distribution and abundance of chroococcoid cyanobacteria in the Southern Ocean. Proc. NIPR Symp. Polar Biol. 1987, 1, 1–9. [Google Scholar]
- Vincent, W.F. Cyanobacterial dominance in the polar regions. In The Ecology of Cyanobacteria; Whitton, B.A., Potts, M., Eds.; Kluwer Academic Publishers: Dordrecht, the Netherlands, 2000; pp. 321–340. [Google Scholar]
- Walker, T.D.; Marchant, H.J. The seasonal occurrence of chroococcoid cyanobacteria at an Antarctic coastal site. Polar Biol. 1989, 9, 193–196. [Google Scholar] [CrossRef]
- Andreoli, C.; Moro, I.; La Rocca, N.; Dalla Valle, L.; Masiero, L.; Rascio, N.; Dalla Vecchia, F. Ecological, physiological and biomolecular surveys on microalgae from Ross Sea (Antarctica). Ital. J. Zool. 2000, 67, 147–156. [Google Scholar]
- Koh, E.Y.; Cowie, R.O.M.; Simpson, A.M.; O’Toole, R.; Ryan, K.G. The origin of cyanobacteria in Antarctic sea ice: Marine or freshwater? Environ. Microbiol. Rep. 2012, 4, 479–483. [Google Scholar] [CrossRef]
- Castiglioni, B.; Rizzi, E.; Frosini, A.; Sivonen, K.; Rajaniemi, P.; Rantala, A.; Mugnai, M.A.; Ventura, S.; Wilmotte, A.; Boutte, C. Development of a universal microarray based on the ligation detection reaction and 16S rRNA gene polymorphism to target diversity of cyanobacteria. Appl. Environ. Microbiol. 2004, 70, 7161–7172. [Google Scholar] [CrossRef]
- Hughes, K.A.; McCartney, H.A.; Lachlan-Cope, T.A.; Pearce, D.A. A preliminary study of airborne microbial biodiversity over peninsular Antarctica. Cell. Mol. Biol. 2004, 50, 537–542. [Google Scholar]
- Pearce, D.A.; Bridge, P.D.; Hughes, K.A.; Sattler, B.; Psenner, R.; Russell, N.J. Microorganisms in the atmosphere over Antarctica. FEMS Microbiol. Ecol. 2009, 69, 143–157. [Google Scholar] [CrossRef]
- Harding, T.; Jungblut, A.D.; Lovejoy, C.; Vincent, W.F. Microbes in high Arctic snow and implications for the cold biosphere. Appl. Environ. Microbiol. 2011, 77, 3234–3243. [Google Scholar] [CrossRef]
- Bottos, E.M.; Vincent, W.F.; Greer, C.W.; Whyte, L.G. Prokaryotic diversity of arctic ice shelf microbial mats. Environ. Microbiol. 2008, 10, 950–966. [Google Scholar] [CrossRef]
- Marshall, G.J.; Turner, J. Katabatic wind propagation over the western Ross Sea observed using ERS-1 scatterometer data. Antarct. Sci. 1997, 9, 221–226. [Google Scholar]
- Raymond, J.; Blankenship, R.E. The evolutionary development of the protein complement of photosystem 2. Biochim. Biophys. Acta. 2004, 1655, 133–139. [Google Scholar] [CrossRef]
- Azam, F.; Smith, D.C.; Hollibaugh, J.T. The role of the microbial loop in Antarctic pelagic ecosystems. Polar Res. 1991, 10, 239–243. [Google Scholar] [CrossRef]
- Achenbach, L.A.; Carey, J.; Madigan, M.T. Photosynthetic and phylogenetic primers for detection of anoxygenic phototrophs in natural environments. Appl. Environ. Microbiol. 2001, 67, 2922–2926. [Google Scholar] [CrossRef]
- Stackebrandt, E.; Rainey, F.A.; Ward-Rainey, N. Anoxygenic phototrophy across the phylogenetic spectrum: Current understanding and future perspectives. Arch. Microbiol. 1996, 166, 211–223. [Google Scholar] [CrossRef]
- Shiba, T.; Simidu, U.; Taga, N. Distribution of aerobic bacteria which contain bacteriochlorophyll a. Appl. Environ. Microbiol. 1979, 38, 43–45. [Google Scholar]
- Kolber, Z.S.; van Dover, C.L.; Niederman, R.A.; Falkowski, P.G. Bacterial photosynthesis in surface waters of the open ocean. Nature 2000, 407, 177–179. [Google Scholar] [CrossRef]
- Yurkov, W.; Beatty, J.T. Aerobic anoxygenic phototrophic bacteria. Microbiol. Mol. Biol. Rev. 1998, 62, 695–724. [Google Scholar]
- Goericke, R. Bacteriochlorophyll a in the ocean: is anoxygenic bacterial photosynthesis important? Limnol. Oceanogr. 2002, 47, 290–295. [Google Scholar] [CrossRef]
- Gregor, J.; Klug, G. Regulation of bacterial photosynthesis genes by oxygen and light. FEMS Microbiol. Lett. 1999, 179, 1–9. [Google Scholar] [CrossRef]
- Nishimura, K.; Shimada, H.; Ohta, H.; Masuda, T.; Shioi, Y.Z.; Takamiya, K. Expression of the puf operon in an aerobic photosynthetic bacterium, Roseobacter denitrificans. Plant Cell Physiol. 1996, 37, 153–159. [Google Scholar] [CrossRef]
- Masuda, S.; Nagashima, K.V.P.; Shimada, K.; Matsuura, K. Transcriptional control of expression of genes for photosynthetic reaction center and light-harvesting proteins in the purple bacterium Rhodovulum sulfidophilum. J. Bacteriol. 2000, 182, 2778–2786. [Google Scholar] [CrossRef]
- Yutin, N.; Beja, O. Putative novel photosynthetic reaction centre organizations in marine aerobic anoxygenic photosynthetic bacteria: Insights from metagenomics and environmental genomics. Environ. Microbiol. 2005, 7, 2027–2033. [Google Scholar] [CrossRef]
- Béjà, O.; Suzuki, M.T.; Heidelberg, J.F.; Nelson, W.C.; Preston, C.M.; Hamada, T.; Eisen, J.A.; Fraser, C.M.; DeLong, E.F. Unsuspected diversity among marine aerobic anoxygenic phototrophs. Nature 2002, 415, 630–633. [Google Scholar] [CrossRef]
- Lambeck, K.; Esat, T.M.; Potter, E.K. Links between climate and sea levels for the past three million years. Nature 2002, 419, 199–206. [Google Scholar] [CrossRef]
- Oz, A.; Sabehi, G.; Koblizek, M.; Massana, R.; Beja, O. Roseobacter-like bacteria in Red and Mediterranean Sea aerobic anoxygenic photosynthetic populations. Appl. Environ. Microbiol. 2005, 71, 344–353. [Google Scholar] [CrossRef]
- Yutin, N.; Suzuki, M.T.; Beja, O. Novel primers reveal wider diversity among marine aerobic anoxygenic phototrophs. Appl. Environ. Microbiol. 2005, 71, 8958–8962. [Google Scholar] [CrossRef]
- Cottrell, M.T.; Kirchman, D.L. Photoheterotrophic microbes in the Arctic Ocean in summer and winter. Appl. Environ. Microbiol. 2009, 75, 4958–4966. [Google Scholar] [CrossRef]
- Jiao, N.Z.; Zhang, F.; Hong, N. Significant roles of bacteriochlorophylla supplemental to chlorophylla in the ocean. ISME J. 2010, 4, 595–597. [Google Scholar] [CrossRef]
- Schwalbach, M.S.; Fuhrman, J.A. Wide-ranging abundances of aerobic anoxygenic phototrophic bacteria in the world ocean revealed by epifluorescence microscopy and quantitative PCR. Limnol. Oceanogr. 2005, 50, 620–628. [Google Scholar] [CrossRef]
- Cottrell, M.T.; Michelou, V.K.; Nemcek, N.; DiTullio, G.; Kirchman, D.L. Carbon cycling by microbes influenced by light in the Northeast Atlantic Ocean. Aquat. Microb. Ecol. 2008, 50, 239–250. [Google Scholar] [CrossRef]
- Karr, E.A.; Sattley, W.M.; Jung, D.O.; Madigan, M.T.; Achenbach, L.A. Remarkable diversity of phototrophic purple bacteria in a permanently frozen Antarctic lake. Appl. Environ. Microbiol. 2003, 69, 4910–4914. [Google Scholar] [CrossRef]
- Perreault, N.N.; Greer, C.W.; Andersen, D.T.; Tille, S.; Lacrampe-Couloume, G.; Lollar, B.S.; Whyte, L.G. Heterotrophic and autotrophic microbial populations in cold perennial springs of the high Arctic. Appl. Environ. Microbiol. 2008, 74, 6898–6907. [Google Scholar] [CrossRef]
- Koh, E.Y.; Phua, W.; Ryan, K.G. Aerobic anoxygenic phototrophic bacteria in Antarctic sea ice and seawater. Environ. Microbiol. Rep. 2011, 3, 710–716. [Google Scholar] [CrossRef]
- Giebel, H.A.; Brinkhoff, T.; Zwisler, W.; Selje, N.; Simon, M. Distribution of Roseobacter RCA and SAR11 lineages and distinct bacterial communities from the subtropics to the Southern Ocean. Environ. Microbiol. 2009, 11, 2164–2178. [Google Scholar] [CrossRef]
- Giebel, H.A.; Kalhoefer, D.; Lemke, A.; Thole, S.; Gahl-Janssen, R.; Simon, M.; Brinkhoff, T. Distribution of Roseobacter RCA and SAR11 lineages in the North Sea and characteristics of an abundant RCA isolate. ISME J. 2011, 5, 8–19. [Google Scholar] [CrossRef]
- Berghoff, B.A.; Glaeser, J.; Nuss, A.M.; Zobawa, M.; Lottspeich, F.; Klug, G. Anoxygenic photosynthesis and photooxidative stress: A particular challenge for Roseobacter. Environ. Microbiol. 2011, 13, 775–791. [Google Scholar] [CrossRef]
- Cowie, R.O.M.; Maas, E.W.; Ryan, K.G. Archaeal diversity revealed in Antarctic sea ice. Antarct. Sci. 2011, 23, 531–536. [Google Scholar] [CrossRef]
- Béja, O.; Aravind, L.; Koonin, E.V.; Suzuki, M.T.; Hadd, A.; Nguyen, L.P.; Jovanovich, S.B.; Gates, C.M.; Feldman, R.A.; Spudich, J.L. Bacterial rhodopsin: Evidence for a new type of phototrophy in the sea. Science 2000, 289, 1902–1906. [Google Scholar]
- Beja, O.; Spudich, E.N.; Spudich, J.L.; Leclerc, M.; Delong, E.F. Proteorhodopsin phototrophy in the ocean. Nature 2001, 411, 786–789. [Google Scholar] [CrossRef]
- Atamna-Ismaeel, N.; Sabehi, G.; Sharon, I.; Witzel, K.P.; Labrenz, M.; Jürgens, K.; Barkay, T.; Stomp, M.; Huisman, J.; Beja, O. Widespread distribution of proteorhodopsins in freshwater and brackish ecosystems. ISME J. 2008, 2, 656–662. [Google Scholar] [CrossRef]
- De La Torre, J.R.; Christianson, L.M.; Béjà, O.; Suzuki, M.T.; Karl, D.M.; Heidelberg, J.; DeLong, E.F. Proteorhodopsin genes are distributed among divergent marine bacterial taxa. Proc. Natl. Acad. Sci. 2003, 100, 12830–12835. [Google Scholar]
- Giovannoni, S.J.; Bibbs, L.; Cho, J.C.; Stapels, M.D.; Desiderio, R.; Vergin, K.L.; Rappé, M.S.; Laney, S.; Wilhelm, L.J.; Tripp, H.J. Proteorhodopsin in the ubiquitous marine bacterium SAR11. Nature 2005, 438, 82–85. [Google Scholar] [CrossRef]
- Campbell, B.J.; Waidner, L.A.; Cottrell, M.T.; Kirchman, D.L. Abundant proteorhodopsin genes in the North Atlantic Ocean. Environ. Microbiol. 2007, 10, 99–109. [Google Scholar]
- Rusch, D.B.; Halpern, A.L.; Sutton, G.; Heidelberg, K.B.; Williamson, S.; Yooseph, S.; Wu, D.; Eisen, J.A.; Hoffman, J.M.; Remington, K. The Sorcerer II global ocean sampling expedition: Northwest Atlantic through Eastern Tropical Pacific. PLoS Biol. 2007, 5, e77. [Google Scholar] [CrossRef]
- Zhao, M.; Chen, F.; Jiao, N. Genetic diversity and abundance of Flavobacterial proteorhodopsin in China Seas. Appl. Environ. Microbiol. 2009, 75, 529–533. [Google Scholar] [CrossRef] [Green Version]
- Sabehi, G.; Loy, A.; Jung, K.H.; Partha, R.; Spudich, J.L.; Isaacson, T.; Hirschberg, J.; Wagner, M.; Béjà, O. New insights into metabolic properties of marine bacteria encoding proteorhodopsins. PLoS Biol. 2005, 3, e273. [Google Scholar]
- Venter, J.C.; Remington, K.; Heidelberg, J.F.; Halpern, A.L.; Rusch, D.; Eisen, J.A.; Wu, D.; Paulsen, I.; Nelson, K.E.; Nelson, W. Environmental genome shotgun sequencing of the Sargasso Sea. Science 2004, 304, 66–74. [Google Scholar]
- Poretsky, R.S.; Bano, N.; Buchan, A.; LeCleir, G.; Kleikemper, J.; Pickering, M.; Pate, W.M.; Moran, M.A.; Hollibaugh, J.T. Analysis of microbial gene transcripts in environmental samples. Appl. Environ. Microbiol. 2005, 71, 4121–4126. [Google Scholar]
- Gómez-Consarnau, L.; González, J.M.; Coll-Lladó, M.; Gourdon, P.; Pascher, T.; Neutze, R.; Pedrós-Alió, C.; Pinhassi, J. Light stimulates growth of proteorhodopsin-containing marine Flavobacteria. Nature 2007, 445, 210–213. [Google Scholar] [CrossRef]
- Michelou, V.K.; Cottrell, M.T.; Kirchman, D.L. Light-stimulated bacterial production and amino acid assimilation by cyanobacteria and other microbes in the North Atlantic Ocean. Appl. Environ. Microbiol. 2007, 73, 5539–5546. [Google Scholar] [CrossRef]
- Steindler, L.; Schwalbach, M.S.; Smith, D.P.; Chan, F.; Giovannoni, S.J. Energy starved Candidatus Pelagibacter ubique substitutes light-mediated ATP production for endogenous carbon respiration. PLoS One 2011, 6, e19725. [Google Scholar]
- Eiler, A. Evidence for the ubiquity of mixotrophic bacteria in the upper ocean: Implications and consequence. Appl. Environ. Microbiol. 2006, 72, 7431–7437. [Google Scholar] [CrossRef]
- Koh, E.Y.; Atamna-Ismaeel, N.; Martin, A.; Cowie, R.O.M.; Beja, O.; Davy, S.K.; Maas, E.W.; Ryan, K.G. Proteorhodopsin-bearing bacteria in Antarctic sea ice. Appl. Environ. Microbiol. 2010, 76, 5918–5925. [Google Scholar] [CrossRef]
- Buckley, R.G.; Trodahl, H.J. Scattering and absorption of visible light by sea ice. Nature 1987, 326, 867–869. [Google Scholar] [CrossRef]
- Man, D.; Wang, W.; Sabehi, G.; Aravind, L.; Post, A.F.; Massana, R.; Spudich, E.N.; Spudich, J.L.; Béjà, O. Diversification and spectral tuning in marine proteorhodopsins. EMBO J. 2003, 22, 1725–1731. [Google Scholar] [CrossRef]
- Sabehi, G.; Kirkup, B.C.; Rozenberg, M.; Stambler, N.; Polz, M.F.; Beja, O. Adaptation and spectral tuning in divergent marine proteorhodopsins from the eastern Mediterranean and the Sargasso Seas. ISME J. 2007, 1, 48–55. [Google Scholar] [CrossRef]
- DeLong, E.F.; Beja, O. The light-driven proton pump proteorhodopsin enhances bacterial survival during tough times. PLoS Biol. 2010, 8, e1000359. [Google Scholar] [CrossRef]
- Terrado, R.; Lovejoy, C.; Massana, R.; Vincent, W.F. Microbial food web responses to light and nutrients beneath the coastal Arctic Ocean sea ice during the winter-spring transition. J. Mar. Sys. 2008, 74, 964–977. [Google Scholar] [CrossRef]
- Collins, R.E.; Rocap, G.; Deming, J.W. Persistence of bacterial and archaeal communities in sea ice through an Arctic winter. Environ. Microbiol. 2010, 12, 1828–1841. [Google Scholar] [CrossRef]
- Goldberg, S.M.D.; Johnson, J.; Busam, D.; Feldblyum, T.; Ferriera, S.; Friedman, R.; Halpern, A.; Khouri, H.; Kravitz, S.A.; Lauro, F.M. A Sanger/pyrosequencing hybrid approach for the generation of high-quality draft assemblies of marine microbial genomes. Proc. Natl. Acad. Sci. 2006, 103, 11240–11245. [Google Scholar]
- Rothberg, J.M.; Leamon, J.H. The development and impact of 454 sequencing. Nat. Biotechnol. 2008, 26, 1117–1124. [Google Scholar] [CrossRef]
- Stepanauskas, R.; Sieracki, M.E. Matching phylogeny and metabolism in the uncultured marine bacteria, one cell at a time. Proc. Natl. Acad. Sci. 2007, 104, 9052–9057. [Google Scholar] [CrossRef]
- Gentry, T.J.; Wickham, G.S.; Schadt, C.W.; He, Z.; Zhou, J. Microarray applications in microbial ecology research. Microb. Ecol. 2006, 52, 159–175. [Google Scholar] [CrossRef]
- Palmer, C.; Bik, E.M.; Eisen, M.B.; Eckburg, P.B.; Sana, T.R.; Wolber, P.K.; Relman, D.A.; Brown, P.O. Rapid quantitative profiling of complex microbial populations. Nucleic Acids Res. 2006, 34, e5. [Google Scholar] [CrossRef]
- DeSantis, T.; Brodie, E.; Moberg, J.; Zubieta, I.; Piceno, Y.; Andersen, G. High-density universal 16S rRNA microarray analysis reveals broader diversity than typical clone library when sampling the environment. Microb. Ecol. 2007, 53, 371–383. [Google Scholar]
- Pernthaler, A.; Amann, R. Simultaneous fluorescence in situ hybridization of mRNA and rRNA in environmental bacteria. Appl. Environ. Microbiol. 2004, 70, 5426–5433. [Google Scholar] [CrossRef]
- Smith, C.J.; Osborn, A.M. Advantages and limitations of quantitative PCR (Q-PCR)-based approaches in microbial ecology. FEMS Microbiol. Ecol. 2009, 67, 6–20. [Google Scholar] [CrossRef]
- Frias-Lopez, J.; Shi, Y.; Tyson, G.W.; Coleman, M.L.; Schuster, S.C.; Chisholm, S.W.; DeLong, E.F. Microbial community gene expression in ocean surface waters. Proc. Natl. Acad. Sci. 2008, 105, 3805–3810. [Google Scholar]
- Zhang, W.; Li, F.; Nie, L. Integrating multiple “omics” analysis for microbial biology: Application and methodologies. Microbiology 2010, 156, 287–301. [Google Scholar] [CrossRef]
- Herring, C.D.; Raffaelle, M.; Allen, T.E.; Kanin, E.I.; Landick, R.; Ansari, A.Z.; Palsson, B.Ø. Immobilization of Escherichia coli RNA polymerase and location of binding sites by use of chromatin immunoprecipitation and microarrays. J. Bacteriol. 2005, 187, 6166–6174. [Google Scholar] [CrossRef]
- Uyar, E.; Kurokawa, K.; Yoshimura, M.; Ishikawa, S.; Ogasawara, N.; Oshima, T. Differential binding profiles of StpA in wild-type and hns mutant cells: A comparative analysis of cooperative partners by chromatin immunoprecipitation-microarray analysis. J. Bacteriol. 2009, 191, 2388–2391. [Google Scholar] [CrossRef]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Koh, E.Y.; Martin, A.R.; McMinn, A.; Ryan, K.G. Recent Advances and Future Perspectives in Microbial Phototrophy in Antarctic Sea Ice. Biology 2012, 1, 542-556. https://doi.org/10.3390/biology1030542
Koh EY, Martin AR, McMinn A, Ryan KG. Recent Advances and Future Perspectives in Microbial Phototrophy in Antarctic Sea Ice. Biology. 2012; 1(3):542-556. https://doi.org/10.3390/biology1030542
Chicago/Turabian StyleKoh, Eileen Y., Andrew R. Martin, Andrew McMinn, and Ken G. Ryan. 2012. "Recent Advances and Future Perspectives in Microbial Phototrophy in Antarctic Sea Ice" Biology 1, no. 3: 542-556. https://doi.org/10.3390/biology1030542



