Novel Polyethers from Screening Actinoallomurus spp.
Abstract
:1. Introduction
2. Results and Discussion
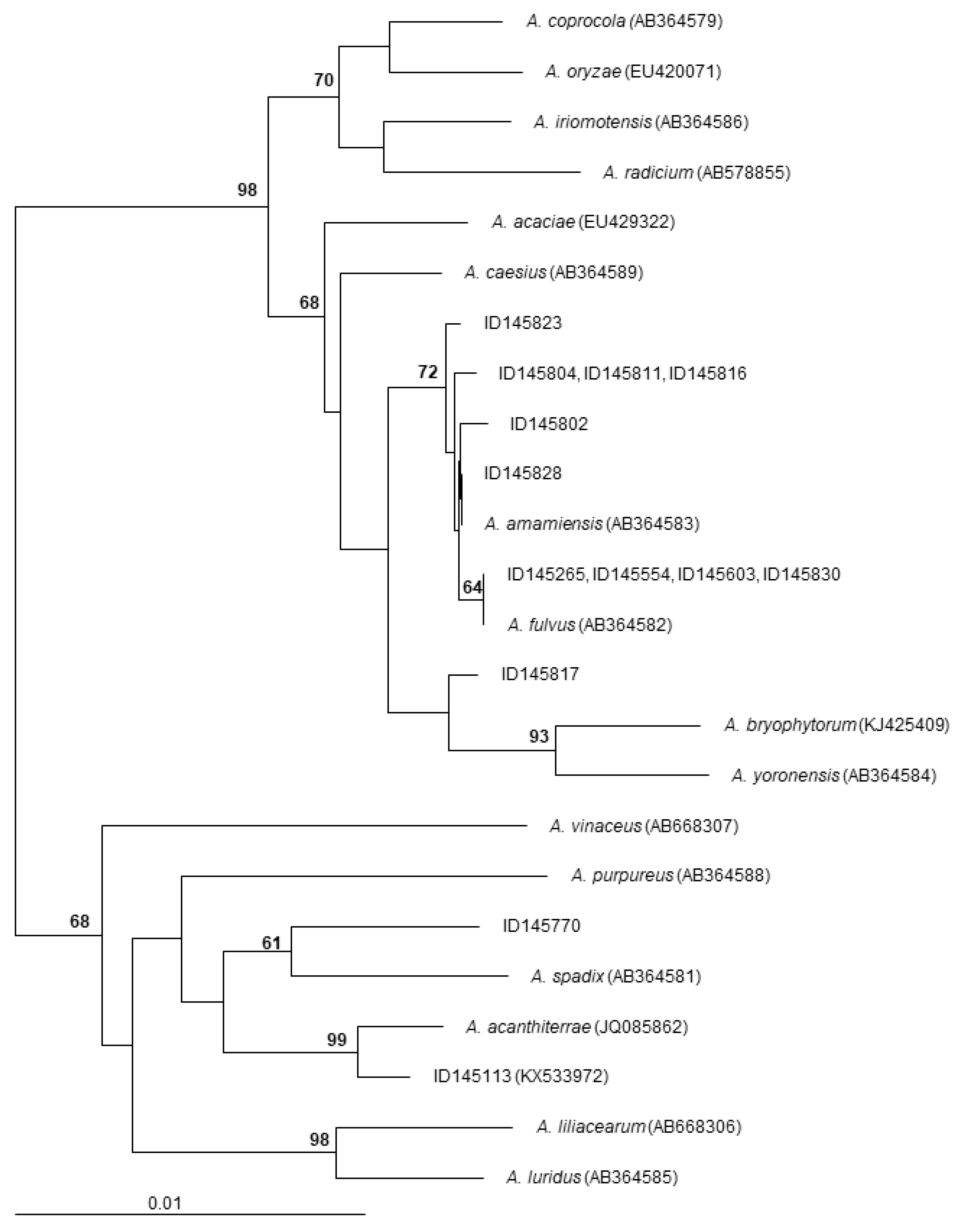
2.1. The Screened Set
2.2. Coumermycins, Spirotetronates, Lantibiotics and Diketopiperazines
2.3. Aromatic Polyketides
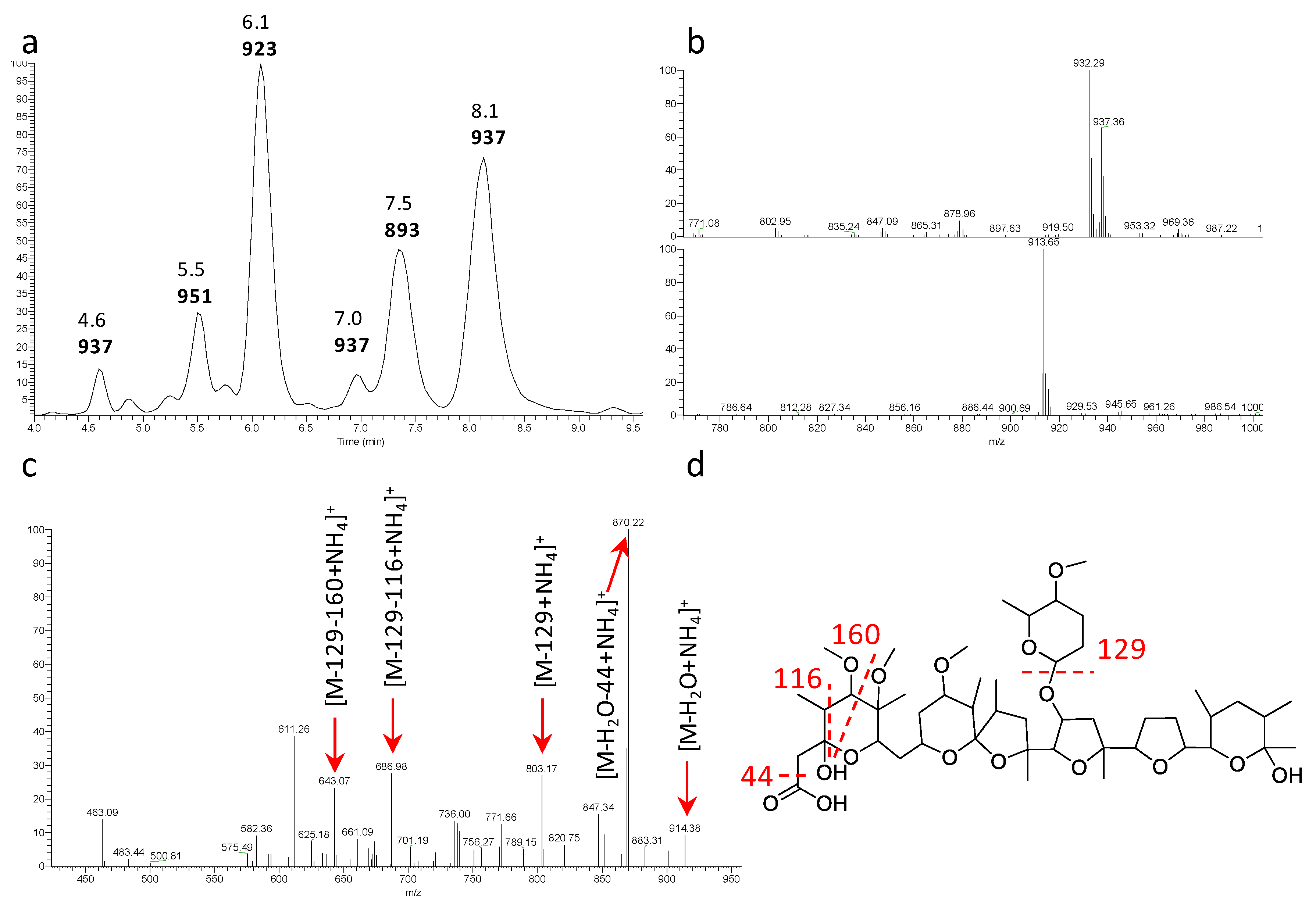
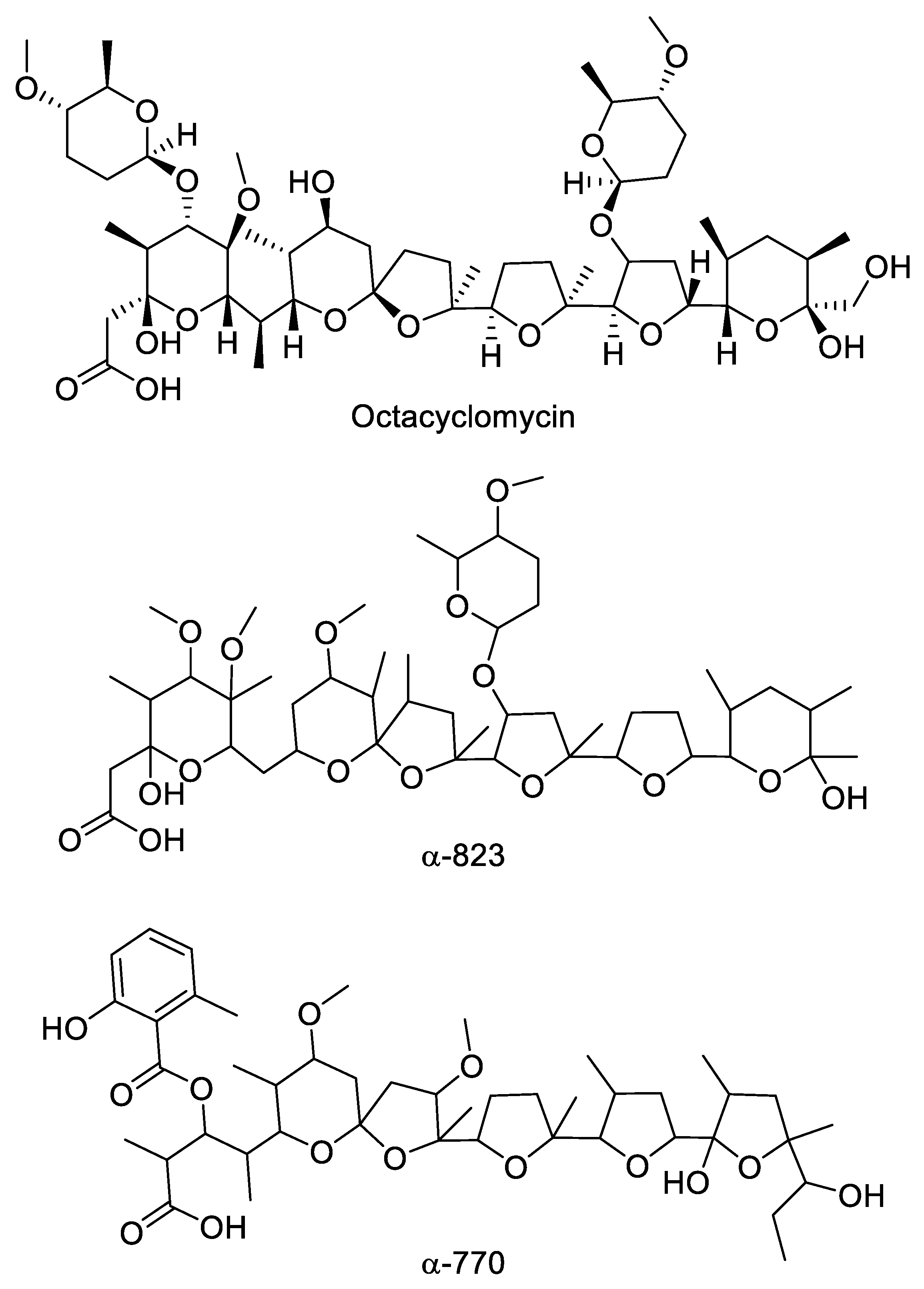
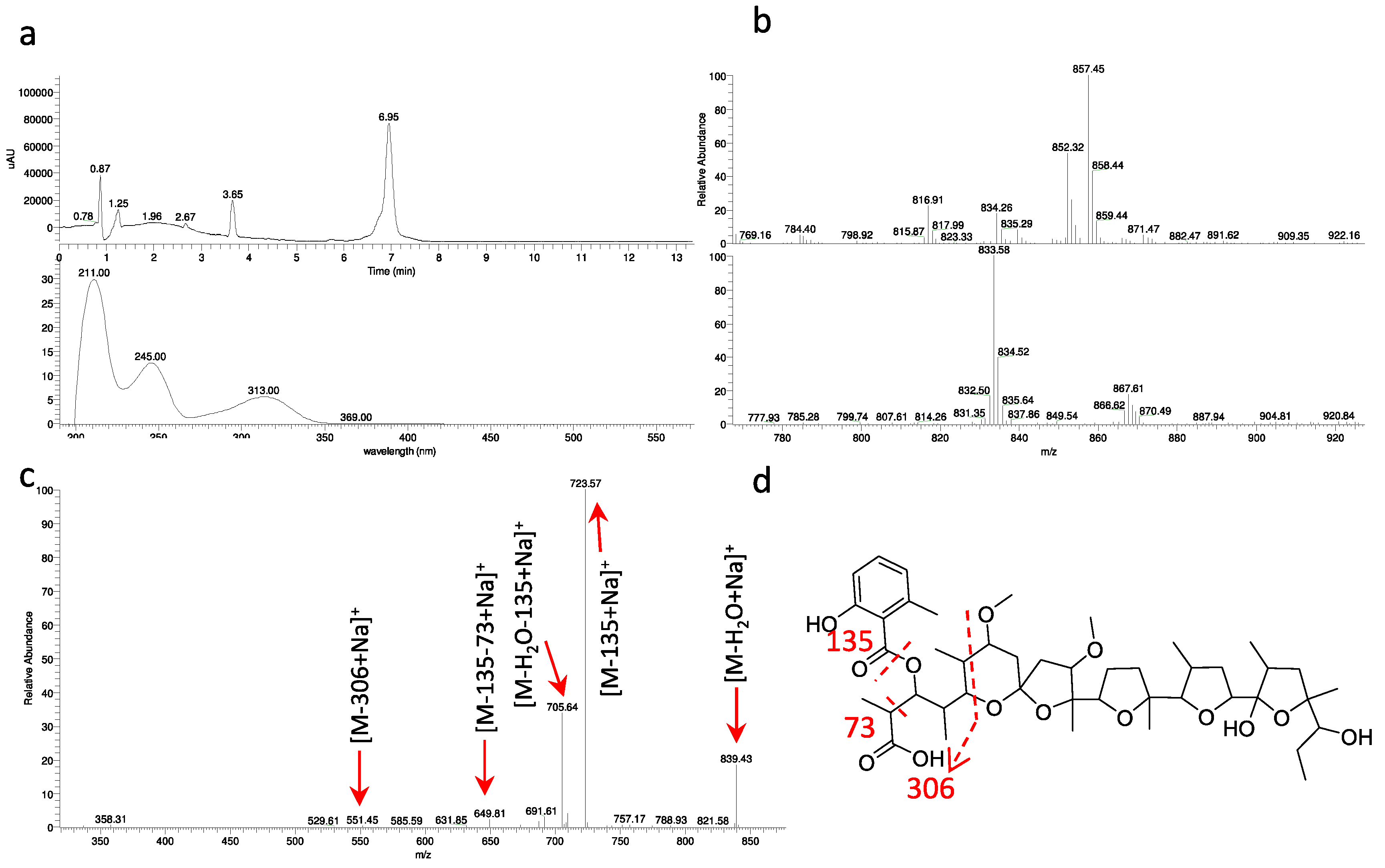
2.4. Polyethers
3. Materials and Methods
3.1. Bacterial Strains and Media
3.2. Preparation of Extracts
3.3. Antibacterial Assays
3.4. Analytical Procedures
3.5. Purification of Polyethers
4. Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Genilloud, O. Actinomycetes: Still a source of novel antibiotics. Nat. Prod. Rep. 2017, 34, 1203–1232. [Google Scholar] [CrossRef] [PubMed]
- Monciardini, P.; Iorio, M.; Maffioli, S.; Sosio, M.; Donadio, S. Discovering new bioactive molecules from microbial sources. Microb. Biotechnol. 2014, 7, 209–220. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wright, G.D. Opportunities for natural products in 21st century antibiotic discovery. Nat. Prod. Rep. 2017, 34, 694–701. [Google Scholar] [CrossRef] [PubMed]
- Jaspars, M.; Challis, G. Microbiology: A talented genus. Nature 2014, 506, 38–39. [Google Scholar] [CrossRef] [PubMed]
- Donadio, S.; Busti, E.; Monciardini, P.; Bamonte, R.; Mazza, P.; Sosio, M.; Cavaletti, L. Sources of polyketides and non-ribosomal peptides. Ernst Schering Res. Found. Workshop 2005, 51, 19–41. [Google Scholar]
- Busti, E.; Cavaletti, L.; Monciardini, P.; Schumann, P.; Rohde, M.; Sosio, M.; Donadio, S. Catenulispora acidiphila gen. nov., sp. nov., a novel, mycelium-forming actinomycete, and proposal of Catenulisporaceae fam. nov. Int. J. Syst. Evol. Microbiol. 2006, 56, 1741–1746. [Google Scholar] [CrossRef] [PubMed]
- Cavaletti, L.; Monciardini, P.; Schumann, P.; Rohde, M.; Bamonte, R.; Busti, E.; Sosio, M.; Donadio, S. Actinospica robiniae gen. nov., sp. nov. and Actinospica acidiphila sp. nov.: Proposal for Actinospicaceae fam. nov. and Catenulisporinae subord. nov. in the order Actinomycetales. Int. J. Syst. Evol. Microbiol. 2006, 56, 1747–1753. [Google Scholar] [CrossRef] [PubMed]
- Monciardini, P.; Cavaletti, L.; Ranghetti, A.; Schumann, P.; Rohde, M.; Bamonte, R.; Sosio, M.; Mezzelani, A.; Donadio, S. Novel members of the family Micromonosporaceae, Rugosimonospora acidiphila gen. nov., sp. nov. and Rugosimonospora africana sp. nov. Int. J. Syst. Evol. Microbiol. 2009, 59, 2752–2758. [Google Scholar] [CrossRef] [PubMed]
- Tamura, T.; Ishida, Y.; Nozawa, Y.; Otoguro, M.; Suzuki, K. Transfer of Actinomadura spadix Nonomura and Ohara 1971 to Actinoallomurus spadix gen. nov., comb. nov., and description of Actinoallomurus amamiensis sp. nov., Actinoallomurus caesius sp. nov., Actinoallomurus coprocola sp. nov., Actinoallomurus fulvus sp. nov., Actinoallomurus iriomotensis sp. nov., Actinoallomurus luridus sp. nov., Actinoallomurus purpureus sp. nov. and Actinoallomurus yoronensis sp. nov. Int. J. Syst. Evol. Microbiol. 2009, 59, 1867–1874. [Google Scholar] [PubMed]
- Thamchaipenet, A.; Indananda, C.; Bunyoo, C.; Duangmal, K.; Matsumoto, A.; Takahashi, Y. Actinoallomurus acaciae sp. nov., an endophytic actinomycete isolated from Acacia auriculiformis A. Cunn. ex Benth. Int. J. Syst. Evol. Microbiol. 2010, 60, 554–559. [Google Scholar] [CrossRef] [PubMed]
- Indananda, C.; Thamchaipenet, A.; Matsumoto, A.; Inahashi, Y.; Duangmal, K.; Takahashi, Y. Actinoallomurus oryzae sp. nov., an endophytic actinomycete isolated from roots of a Thai jasmine rice plant. Int. J. Syst. Evol. Microbiol. 2011, 61, 737–741. [Google Scholar] [CrossRef] [PubMed]
- Pozzi, R.; Simone, M.; Mazzetti, C.; Maffioli, S.; Monciardini, P.; Cavaletti, L.; Bamonte, R.; Sosio, M.; Donadio, S. The genus Actinoallomurus and some of its metabolites. J. Antibiot. 2011, 64, 133–139. [Google Scholar] [CrossRef] [PubMed]
- Koyama, R.; Matsumoto, A.; Inahashi, Y.; Omura, S.; Takahashi, Y. Isolation of actinomycetes from the root of the plant, Ophiopogon japonicus, and proposal of two new species, Actinoallomurus liliacearum sp. nov. and Actinoallomurus vinaceus sp. nov. J. Antibiot. 2012, 65, 335–340. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, A.; Fukuda, A.; Inahashi, Y.; Omura, S.; Takahashi, Y. Actinoallomurus radicium sp. nov., isolated from the roots of two plant species. Int. J. Syst. Evol. Microbiol. 2012, 62, 295–298. [Google Scholar] [CrossRef] [PubMed]
- Inahashi, Y.; Iwatsuki, M.; Ishiyama, A.; Matsumoto, A.; Hirose, T.; Oshita, J.; Sunazuka, T.; Panbangred, W.; Takahashi, Y.; Kaiser, M.; et al. Actinoallolides A-E, new anti-trypanosomal macrolides, produced by an endophytic actinomycete, Actinoallomurus fulvus MK10-036. Org. Lett. 2015, 17, 864–867. [Google Scholar] [CrossRef] [PubMed]
- Mazzetti, C.; Ornaghi, M.; Gaspari, E.; Parapini, S.; Maffioli, S.; Sosio, M.; Donadio, S. Halogenated spirotetronates from Actinoallomurus. J. Nat. Prod. 2012, 75, 1044–1050. [Google Scholar] [CrossRef] [PubMed]
- Cruz, J.C.S.; Maffioli, S.; Bernasconi, A.; Brunati, C.; Gaspari, E.; Sosio, M.; Wellington, E.; Donadio, S. Allocyclinones, hyperchlorinated angucyclinones and common metabolites from Actinoallomurus. J. Antibiot. 2017, 70, 73–78. [Google Scholar] [CrossRef] [PubMed]
- Iorio, M.; Cruz, J.; Simone, M.; Bernasconi, A.; Brunati, C.; Sosio, M.; Donadio, S.; Maffioli, S.I. Antibacterial paramagnetic quinones from Actinoallomurus. J. Nat. Prod. 2017, 80, 819–827. [Google Scholar] [CrossRef] [PubMed]
- Cruz, J.C.S.; Iorio, M.; Monciardini, P.; Simone, M.; Brunati, C.; Gaspari, E.; Maffioli, S.I.; Wellington, E.; Sosio, M.; Donadio, S. Brominated variant of the lantibiotic NAI-107 with enhanced antibacterial potency. J. Nat. Prod. 2015, 78, 2642–2647. [Google Scholar] [CrossRef] [PubMed]
- Kammerer, B.; Kahlich, R.; Laufer, S.; Li, S.M.; Heide, L.; Gleiter, C.H. Mass spectrometric pathway monitoring of secondary metabolites: Systematic analysis of culture extracts of Streptomyces species. Anal. Biochem. 2004, 335, 17–29. [Google Scholar] [CrossRef] [PubMed]
- Vieweg, L.; Reichau, S.; Schobert, R.; Leadlay, P.F.; Süssmuth, R.D. Recent advances in the field of bioactive tetronates. Nat. Prod. Rep. 2014, 31, 1554–1584. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lam, K.S.; Hesler, G.A.; Gustavson, D.R.; Berry, R.L.; Tomita, K.; MacBeth, J.L.; Ross, J.; Miller, D.; Forenza, S. Pyrrolosporin A, a new antitumor antibiotic from Micromonospora sp. C39217-R109-7. I. Taxonomy of producing organism, fermentation and biological activity. J. Antibiot. 1996, 49, 860–864. [Google Scholar] [CrossRef] [PubMed]
- Schroeder, D.R.; Colson, K.L.; Klohr, S.E.; Lee, M.S.; Matson, J.A.; Brinen, L.S.; Clardy, J. Pyrrolosporin A, a new antitumor antibiotic from Micromonospora sp. C39217-R109-7. II. Isolation, physic-chemical properties, spectroscopic study and X-ray analysis. J. Antibiot. 1996, 9, 865–872. [Google Scholar] [CrossRef]
- Arnison, P.G.; Bibb, M.J.; Bierbaum, G.; Bowers, A.A.; Bugni, T.S.; Bulaj, G.; Camarero, J.A.; Campopiano, D.J.; Challis, G.L.; Clardy, J.; et al. Ribosomally synthesized and post-translationally modified peptide natural products: Overview and recommendations for a universal nomenclature. Nat. Prod. Rep. 2013, 30, 108–160. [Google Scholar] [CrossRef] [PubMed]
- Lee, M.D. Antibiotics from Microbispora. U.S. Patent 6,551,591, 22 April 2003. [Google Scholar]
- Lazzarini, A.; Gastaldo, L.; Candiani, G.; Ciciliato, I.; Losi, D.; Marinelli, F.; Selva, E.; Parenti, F. Antibiotic 107891, Its Factors A1 and A2, Pharmaceutically Acceptable Salts and Compositions, and Use Thereof. U.S. Patent 7,351,687, 1 April 2008. [Google Scholar]
- Belin, P.; Moutiez, M.; Lautru, S.; Seguin, J.; Pernodet, J.L.; Gondry, M. The nonribosomal synthesis of diketopiperazines in tRNA-dependent cyclodipeptide synthase pathways. Nat. Prod. Rep. 2012, 29, 961–979. [Google Scholar] [CrossRef] [PubMed]
- Oki, T.; Kinoshi, M.; Tomatsu, K.; Tomita, K.; Saitoh, K.; Tsunakawa, M.; Nishio, M.; Miyaki, T.; Kawaguchi, H. Pradimicin, a novel class of potent antifungal antibiotics. J. Antibiot. 1999, 11, 1701–1704. [Google Scholar] [CrossRef]
- Tadashi, N.; Yukio, T.; Yuko, K.; Mamoru, I.; Taneto, T.; Shinji, M.; Masaji, S.; Michio, K. Novel antibiotic substance SF-2361, production and used thereof. Japan Patent 61260888, 19 November 1986. [Google Scholar]
- Kevin, D.A., II; Meujo, D.A.F.; Hamann, M.T. Polyether ionophores: Broad-spectrum and promising biologically active molecules for the control of drug-resistant bacteria and parasites. Expert Opin. Drug Discov. 2009, 4, 109–146. [Google Scholar] [CrossRef] [PubMed]
- Funayama, S.; Nozoe, S.; Tronquet, C.; Anraku, Y.; Komiyama, K.; Omura, S. Isolation and structure of a new polyether antibiotic, octacyclomycin. J. Antibiot. 1992, 45, 1686–1691. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, G.; Kobayashi, K.; Sakurai, T.; Isono, K. Cationomycin, a new polyether ionophore antibiotic produced by Actinomadura nov. sp. J. Antibiot. 1981, 34, 1513–1514. [Google Scholar] [CrossRef] [PubMed]
- Minami, A.; Oguri, H.; Watanabe, K.; Oikawa, H. Biosynthetic machinery of ionophore polyether lasalocid: Enzymatic construction of polyether skeleton. Curr. Opin. Chem. Biol. 2013, 17, 555–561. [Google Scholar] [CrossRef] [PubMed]
- Ubukata, M.; Uzawa, J.; Isono, K. Biosynthesis of cationomycin: Direct and indirect incorporation of 13C-acetate and application of homoscalar correlated 2-D carbon-13 NMR and double quantum coherence. J. Am. Chem. Soc. 1984, 106, 2213–2214. [Google Scholar] [CrossRef]
- Rutkowski, J.; Brzezinski, B. Structures and properties of naturally occurring polyether antibiotics. BioMed Res. Int. 2013, 2013, 162513. [Google Scholar] [CrossRef] [PubMed]
- Zhou, S.; Wang, F.; Wong, E.T.; Fonkem, E.; Hsieh, T.C.; Wu, JM.; Wu, E. Salinomycin: A novel anti-cancer agent with known anti-coccidial activities. Curr. Med. Chem. 2013, 20, 4095–4101. [Google Scholar] [CrossRef] [PubMed]
- D’Alessandro, S.; Corbett, Y.; Ilboudo, D.P.; Misiano, P.; Dahiya, N.; Abay, S.M.; Habluetzel, A.; Grande, R.; Gismondo, M.R.; Dechering, K.J.; et al. Salinomycin and other ionophores as a new class of antimalarial drugs with transmission-blocking activity. Antimicrob. Agents Chemother. 2015, 59, 5135–5144. [Google Scholar] [CrossRef] [PubMed]
- Charlop-Powers, Z.; Owen, J.G.; Reddy, B.V.B.; Ternei, M.A.; Brady, S.F. Chemical-biogeographic survey of secondary metabolism in soil. Proc. Natl. Acad. Sci. USA 2014, 111, 3757–3762. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Charlop-Powers, Z.; Pregitzer, C.C.; Lemetre, C.; Ternei, M.A.; Maniko, J.; Hover, B.M.; Calle, P.Y.; McGuire, K.L.; Garbarino, J.; Forgione, H.M.; et al. Urban park soil microbiomes are a rich reservoir of natural product biosynthetic diversity. Proc. Natl. Acad. Sci. USA 2016, 113, 14811–14816. [Google Scholar] [CrossRef] [PubMed]
- Monciardini, P.; Sosio, M.; Cavaletti, L.; Chiocchini, C.; Donadio, S. New PCR primers for the selective amplifcation of 16S rDNA from different groups of actinomycetes. FEMS Microbiol. Ecol. 2002, 42, 419–429. [Google Scholar] [PubMed]
- Donadio, S.; Monciardini, P.; Sosio, M. Chapter 1. Approaches to discovering novel antibacterial and antifungal agents. Methods Enzymol. 2009, 458, 3–28. [Google Scholar] [PubMed]


| Continent | Analyzed Strains | Active Strains | Active (%) |
|---|---|---|---|
| Europe | 124 | 59 | 48% |
| Africa a | 24 | 17 | 71% |
| Asia a | 12 | 3 | 25% |
| Americas b | 40 | 25 | 62% |
| Strain | Origin | Accession Number a | m/z [M−Na]+ | Compound |
|---|---|---|---|---|
| ID145265 | soil, Nicaragua | MH3933000 | 937 | α-823 |
| ID145554 | soil, Mauritius | MH3933001 | 937 | α-823 |
| ID145603 | soil, Brazil | MH3933002 | 937 | α-823 |
| ID145770 | soil, Niger | MH3933011 | 857 | α-770 |
| ID145802 | soil, Nicaragua | MH3933003 | 937 | α-823 |
| ID145804 | soil, Cameroon | MH3933004 | 937 | α-823 |
| ID145811 | soil, Cameroon | MH3933005 | 937 | α-823 |
| ID145816 | soil, Cameroon | MH3933006 | 937 | α-823 |
| ID145817 | soil, Cameroon | MH3933010 | 1039 | octacyclomycin |
| ID145823 | soil, Venezuela | MH3933007 | 937 | α-823 |
| ID145828 | soil, Nicaragua | MH3933008 | 937 | α-823 |
| ID145830 | soil, Nicaragua | MH3933009 | 937 | α-823 |
| Microorganism a | Code | MIC (µg/mL) | ||
|---|---|---|---|---|
| α-770 | α-823 | Salinomycin | ||
| Staphylococcus aureus (MSSA) | ATCC 6538P | 0.125 | 0.5 | 0.25 |
| S. aureus (MRSA) | L1400 | 0.125 | 2 | 0.5 |
| S. aureus (GISA) | L3797 | 0.125 | 1 | 0.125 |
| Streptococcus pyogenes | L49 | ≤0.03 | ≤0.03 | ≤0.03 |
| S. pneumoniae | L44 | ≤0.03 | 0.125 | 0.125 |
| S. haemolyticus | L1730 | 0.125 | 1 | 0.5 |
| S. epidermidis | ATCC 12228 | 0.125 | 1 | 0.5 |
| Enterococcus faecalis | L559 | ≤0.03 | 0.25 | ≤0.03 |
| E. faecium | L568 | 0.25 | 1 | 1 |
| Bacillus subtilis | ATCC 6633 | ≤0.03 | 0.125 | 0.125 |
| Micrococcus luteus | ATCC 10240 | 0.125 | 0.5 | 0.5 |
| Mycobacterium smegmatis | mc2 155 | 16 | 2 | 64 |
| Moraxella catarrhalis | L3292 | 2 | 8 | 16 |
| Clostridium difficile | L4013 | 0.5 | 0.125 | 0.25 |
| Candida albicans | L145 | >64 | >64 | >64 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Iorio, M.; Tocchetti, A.; Cruz, J.C.S.; Del Gatto, G.; Brunati, C.; Maffioli, S.I.; Sosio, M.; Donadio, S. Novel Polyethers from Screening Actinoallomurus spp. Antibiotics 2018, 7, 47. https://doi.org/10.3390/antibiotics7020047
Iorio M, Tocchetti A, Cruz JCS, Del Gatto G, Brunati C, Maffioli SI, Sosio M, Donadio S. Novel Polyethers from Screening Actinoallomurus spp. Antibiotics. 2018; 7(2):47. https://doi.org/10.3390/antibiotics7020047
Chicago/Turabian StyleIorio, Marianna, Arianna Tocchetti, Joao Carlos Santos Cruz, Giancarlo Del Gatto, Cristina Brunati, Sonia Ilaria Maffioli, Margherita Sosio, and Stefano Donadio. 2018. "Novel Polyethers from Screening Actinoallomurus spp." Antibiotics 7, no. 2: 47. https://doi.org/10.3390/antibiotics7020047





