Leptospiral Pathogenomics
Abstract
:1. Introduction
1.1. Leptospirosis

1.2. Leptospira
1.3. Maintenance Hosts
1.4. Pathogenomics: An Overview
2. Software
3. Leptospira Genomics and Pathogenomics Studies
3.1. Mobile Genetic Elements
3.1.1. Bacteriophages
3.1.2. Evidence for Plasmid Transfer
3.1.3. Repetitive Elements
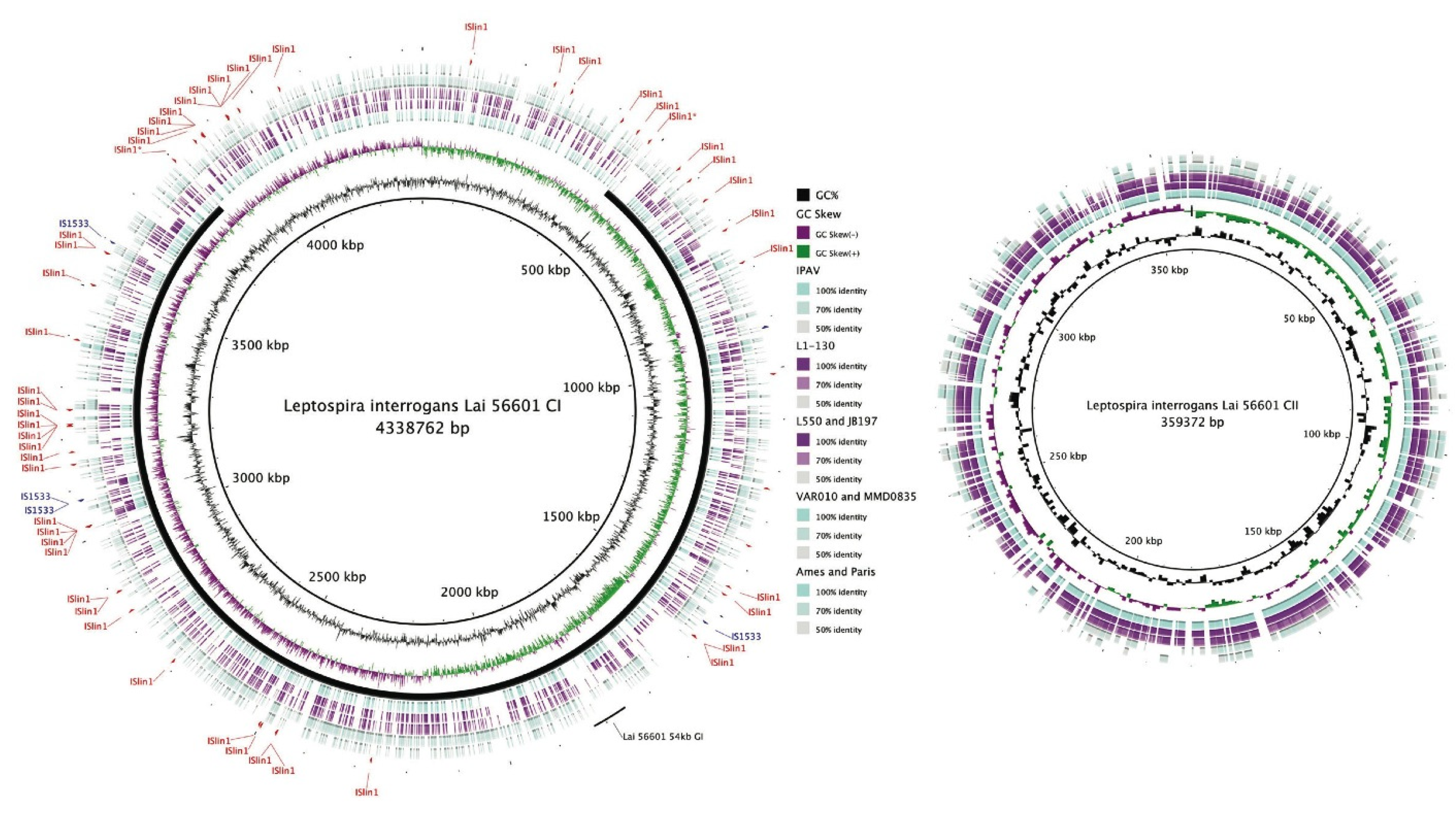
3.1.4. O-Antigen Diversity
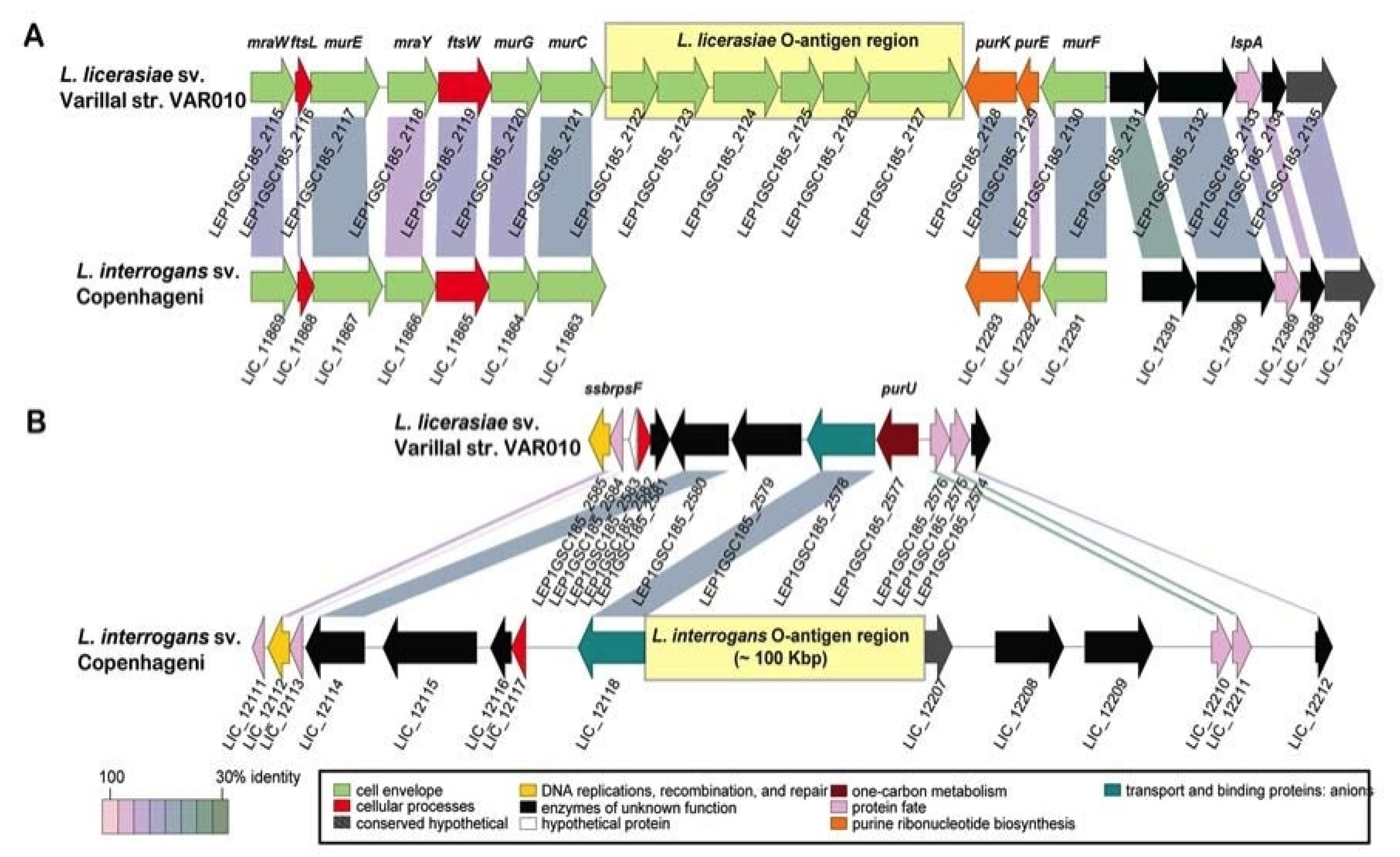
3.2. Virulence Mechanisms and Pathogenesis
3.2.1. Genetic Manipulation
3.2.2. Attenuation Studies in L. interrogans Serovar Lai
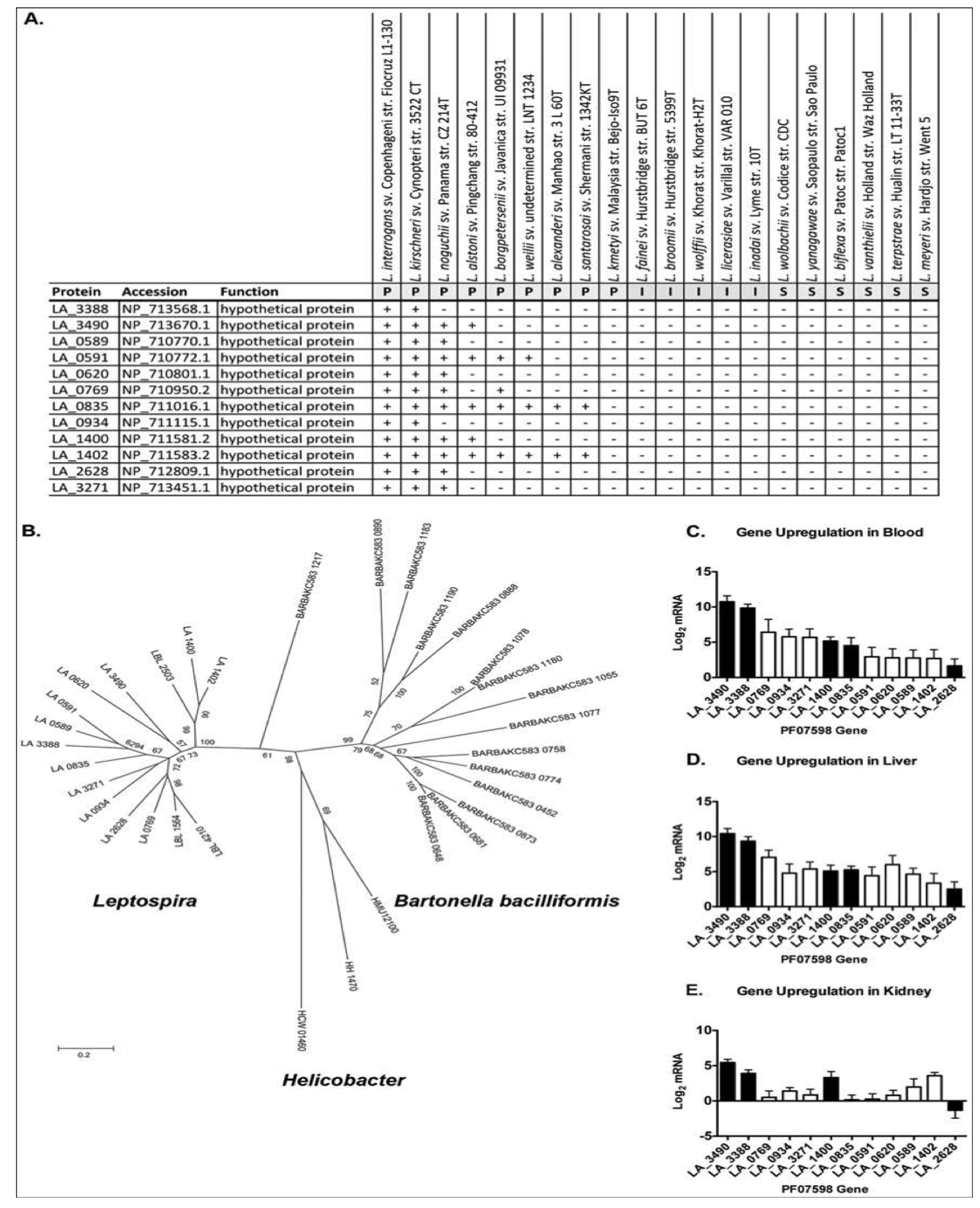
3.2.3. Comparisons of L. borgpetersenii, L. interrogans, L. licerasiae, and L. biflexa
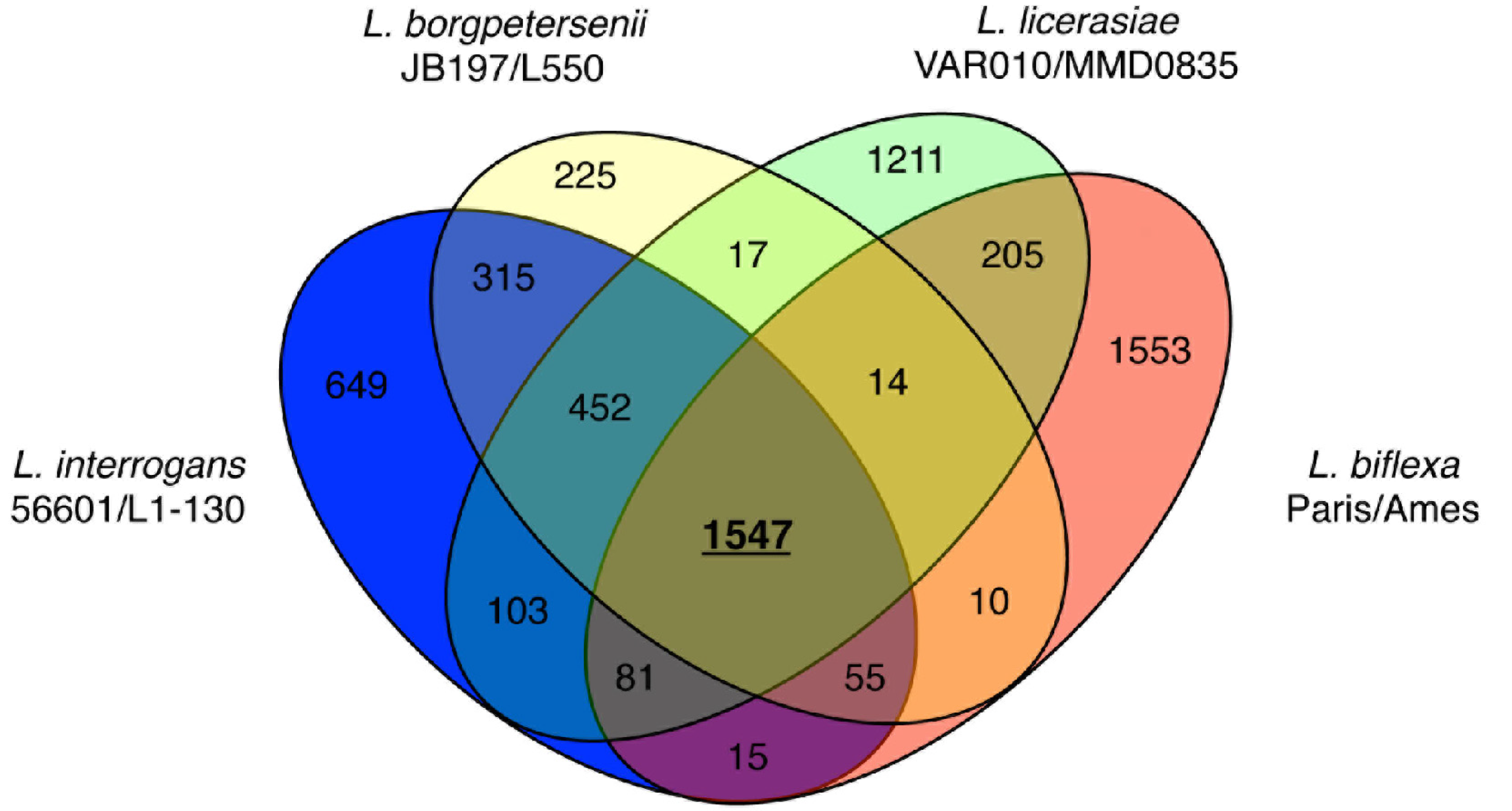
3.3. Vaccine Candidates
3.4. Metabolism
4. Future Directions
- To obtain whole genome information for all known Leptospira species. Currently there are nine named pathogenic Leptospira species, five intermediate Leptospira species, and six saprophytic Leptospira species; more have recently been informally reported and inevitably these numbers will grow over coming years. This information will provide the basis for identifying a minimal number of molecular markers for multilocus sequence typing that can differentiate infecting leptospires directly from human samples without the need for bacterial isolation. Conserved protein markers that are the targets of antibody recognition or antigen detection will be identified. Accomplishing these goals depends on obtaining whole genome sequence of a globally diverse and representative set of Leptospira strains.
- To delineate taxonomic and phylogenetic relationships among Leptospira species. Current methods to classify new species or serovars and to identify the emergence of new leptospiral causes of human disease are cumbersome and insufficiently informative. Genomic-level information will allow us to determine serovar without the need for serological typing, will provide fresh insights into the utility of serovar as a tool for strain identification, and if shown to be robust, will facilitate the development of molecular-based serovar typing. Complete genome sequence of reference strains used for serological diagnosis is critical for refining and optimizing efficient diagnosis.
- To understand the mechanisms of leptospirosis pathogenesis and determinants of clinical outcome. Correlations of genetic polymorphisms and virulence will be identified between isolates at the species, serovar, and strain level. This will require sequencing of isolates associated with distinct clinical presentations and outcomes. This data will provide the fundamental basis for hypothesis-driven research to determine virulence factors and for vaccine development.
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Bharti, A.R.; Nally, J.E.; Ricaldi, J.N.; Matthias, M.A.; Diaz, M.M.; Lovett, M.A.; Levett, P.N.; Gilman, R.H.; Willig, M.R.; Gotuzzo, E. Leptospirosis: a zoonotic disease of global importance. Lancet Infect. Dis. 2003, 3, 757–771. [Google Scholar] [CrossRef]
- Abela-Ridder, B.; Sikkema, R.; Hartskeerl, R.A. Estimating the burden of human leptospirosis. Int. J. Antimicrob. Agents 2010, 36 (Suppl. 1), S5–S7. [Google Scholar] [CrossRef]
- Ko, A.I.; Galvao Reis, M.; Ribeiro Dourado, C.M.; Johnson, W.D., Jr.; Riley, L.W. Urban epidemic of severe leptospirosis in Brazil. Salvador Leptospirosis Study Group. Lancet 1999, 354, 820–825. [Google Scholar] [CrossRef]
- Gouveia, E.L.; Metcalfe, J.; de Carvalho, A.L.; Aires, T.S.; Villasboas-Bisneto, J.C.; Queirroz, A.; Santos, A.C.; Salgado, K.; Reis, M.G.; Ko, A.I. Leptospirosis-associated severe pulmonary hemorrhagic syndrome, Salvador, Brazil. Emerg. Infect. Dis. 2008, 14, 505–508. [Google Scholar] [CrossRef]
- Zaki, S.R.; Shieh, W.J.; The Epidemic Working Group at Ministry of Health in Nicaragua. Leptospirosis associated with outbreak of acute febrile illness and pulmonary haemorrhage, Nicaragua, 1995. Lancet 1996, 347, 535–536. [Google Scholar] [CrossRef]
- Trevejo, R.T.; Rigau-Perez, J.G.; Ashford, D.A.; McClure, E.M.; Jarquin-Gonzalez, C.; Amador, J.J.; de los Reyes, J.O.; Gonzalez, A.; Zaki, S.R.; Shieh, W.J.; et al. Epidemic leptospirosis associated with pulmonary hemorrhage-Nicaragua, 1995. J. Infect. Dis. 1998, 178, 1457–1463. [Google Scholar] [CrossRef]
- Golubev, M.V.; Litvin, V. Mechanism of the maintenance of the infective capacity of soil in a natural focus of leptospirosis. Zh. Mikrobiol. Epidemiol. Immunobiol. 1983, 43–46. [Google Scholar]
- Tonge, J.L.; Smith, D.J. Leptospirosis acquired from soil. Med. J. Aust. 1961, 48, 711–712. [Google Scholar]
- Zaitsev, S.V.; Chernukha Iu, G.; Evdokimova, O.A.; Belov, A.S. Survival rate of Leptospira pomona in the soil at a natural leptospirosis focus. Zh. Mikrobiol. Epidemiol. Immunobiol. 1989, 64–68. [Google Scholar]
- Caldas, E.M.; Sampaio, M.B. Leptospirosis in the city of Salvador, Bahia, Brazil: a case-control seroepidemiologic study. Int. J. Zoonoses 1979, 6, 85–96. [Google Scholar]
- Monahan, A.M.; Miller, I.S.; Nally, J.E. Leptospirosis: risks during recreational activities. J. Appl. Microbiol. 2009, 107, 707–716. [Google Scholar] [CrossRef]
- Gaynor, K.; Katz, A.R.; Park, S.Y.; Nakata, M.; Clark, T.A.; Effler, P.V. Leptospirosis on Oahu: an outbreak associated with flooding of a university campus. Am. J. Trop. Med. Hyg. 2007, 76, 882–885. [Google Scholar]
- Pellizzer, P.; Todescato, A.; Benedetti, P.; Colussi, P.; Conz, P.; Cinco, M. Leptospirosis following a flood in the Veneto area, North-east Italy. Ann. Ig. 2006, 18, 453–456. [Google Scholar]
- Brief report: Leptospirosis after flooding of a university campus--Hawaii, 2004. MMWR Morb. Mortal. Wkly. Rep. 2006, 55, 125–127.
- Vanasco, N.B.; Fusco, S.; Zanuttini, J.C.; Manattini, S.; Dalla Fontana, M.L.; Prez, J.; Cerrano, D.; Sequeira, M.D. Outbreak of human leptospirosis after a flood in Reconquista, Santa Fe, 1998. Rev. Argent Microbiol. 2002, 34, 124–131. [Google Scholar]
- Easton, A. Leptospirosis in Philippine floods. BMJ 1999, 319, 212. [Google Scholar] [CrossRef]
- Adler, B.; Faine, S. Host immunological mechanisms in the resistance of mice to leptospiral infections. Infect. Immun. 1977, 17, 67–72. [Google Scholar]
- Chassin, C.; Picardeau, M.; Goujon, J.M.; Bourhy, P.; Quellard, N.; Darche, S.; Badell, E.; d'Andon, M.F.; Winter, N.; Lacroix-Lamande, S.; et al. TLR4- and TLR2-mediated B cell responses control the clearance of the bacterial pathogen, Leptospira interrogans. J. Immunol. 2009, 183, 2669–2677. [Google Scholar] [CrossRef]
- Outbreak of leptospirosis among white-water rafters--Costa Rica, 1996. MMWR Morb. Mortal. Wkly. Rep. 1997, 46, 577–579.
- Boland, M.; Sayers, G.; Coleman, T.; Bergin, C.; Sheehan, N.; Creamer, E.; O'Connell, M.; Jones, L.; Zochowski, W. A cluster of leptospirosis cases in canoeists following a competition on the River Liffey. Epidemiol. Infect. 2004, 132, 195–200. [Google Scholar] [CrossRef]
- Morgan, J.; Bornstein, S.L.; Karpati, A.M.; Bruce, M.; Bolin, C.A.; Austin, C.C.; Woods, C.W.; Lingappa, J.; Langkop, C.; Davis, B.; et al. Outbreak of leptospirosis among triathlon participants and community residents in Springfield, Illinois, 1998. Clin. Infect. Dis. 2002, 34, 1593–1599. [Google Scholar] [CrossRef]
- Sejvar, J.; Bancroft, E.; Winthrop, K.; Bettinger, J.; Bajani, M.; Bragg, S.; Shutt, K.; Kaiser, R.; Marano, N.; Popovic, T.; et al. Leptospirosis in "Eco-Challenge" athletes, Malaysian Borneo, 2000. Emerg. Infect. Dis. 2003, 9, 702–707. [Google Scholar] [CrossRef]
- Ganoza, C.A.; Matthias, M.A.; Collins-Richards, D.; Brouwer, K.C.; Cunningham, C.B.; Segura, E.R.; Gilman, R.H.; Gotuzzo, E.; Vinetz, J.M. Determining risk for severe leptospirosis by molecular analysis of environmental surface waters for pathogenic Leptospira. PLoS Med. 2006, 3, e308. [Google Scholar] [CrossRef]
- Brenner, D.J.; Kaufmann, A.F.; Sulzer, K.R.; Steigerwalt, A.G.; Rogers, F.C.; Weyant, R.S. Further determination of DNA relatedness between serogroups and serovars in the family Leptospiraceae with a proposal for Leptospira alexanderi sp. nov., and four new Leptospira genomospecies. Int. J. Syst. Bacteriol. 1999, 49, Pt. 2. 839–858. [Google Scholar] [CrossRef]
- Slack, A.T.; Khairani-Bejo, S.; Symonds, M.L.; Dohnt, M.F.; Galloway, R.L.; Steigerwalt, A.G.; Bahaman, A.R.; Craig, S.; Harrower, B.J.; Smythe, L.D. Leptospira kmetyi sp. nov., isolated from an environmental source in Malaysia. Int. J. Syst. Evol. Microbiol. 2009, 59, 705–708. [Google Scholar] [CrossRef]
- Schmid, G.P.; Steere, A.C.; Kornblatt, A.N.; Kaufmann, A.F.; Moss, C.W.; Johnson, R.C.; Hovind-Hougen, K.; Brenner, D.J. Newly recognized Leptospira species ("Leptospira inadai" serovar lyme) isolated from human skin. J. Clin. Microbiol. 1986, 24, 484–486. [Google Scholar]
- Petersen, A.M.; Boye, K.; Blom, J.; Schlichting, P.; Krogfelt, K.A. First isolation of Leptospira fainei serovar Hurstbridge from two human patients with Weil’s syndrome. J. Med. Microbiol. 2001, 50, 96–100. [Google Scholar]
- Matthias, M.A.; Ricaldi, J.N.; Cespedes, M.; Diaz, M.M.; Galloway, R.L.; Saito, M.; Steigerwalt, A.G.; Patra, K.P.; Ore, C.V.; Gotuzzo, E.; et al. Human leptospirosis caused by a new, antigenically unique Leptospira associated with a Rattus species reservoir in the Peruvian Amazon. PLoS Negl. Trop. Dis. 2008, 2, e213. [Google Scholar] [CrossRef]
- Levett, P.N.; Morey, R.E.; Galloway, R.L.; Steigerwalt, A.G. Leptospira broomii sp. nov., isolated from humans with leptospirosis. Int. J. Syst. Evol. Microbiol. 2006, 56, 671–673. [Google Scholar] [CrossRef]
- Slack, A.T.; Kalambaheti, T.; Symonds, M.L.; Dohnt, M.F.; Galloway, R.L.; Steigerwalt, A.G.; Chaicumpa, W.; Bunyaraksyotin, G.; Craig, S.; Harrower, B.J.; et al. Leptospira wolffii sp. nov., isolated from a human with suspected leptospirosis in Thailand. Int. J. Syst. Evol. Microbiol. 2008, 58, 2305–2308. [Google Scholar] [CrossRef]
- Saito, M.; Villanueva, S.Y.; Kawamura, Y.; Iida, K.I.; Tomida, J.; Kanemaru, T.; Kohno, E.; Miyahara, S.; Umeda, A.; Amako, K.; et al. Leptospira idonii sp. nov., isolated from an environmental water in Fukuoka, Japan. Int. J. Syst. Evol. Microbiol. 2012, 63, 2457–2462. [Google Scholar]
- Gravekamp, C.; Korver, H.; Montgomery, J.; Everard, C.O.; Carrington, D.; Ellis, W.A.; Terpstra, W.J. Leptospires isolated from toads and frogs on the Island of Barbados. Zentralbl. Bakteriol. 1991, 275, 403–411. [Google Scholar] [CrossRef]
- Athanazio, D.A.; Silva, E.F.; Santos, C.S.; Rocha, G.M.; Vannier-Santos, M.A.; McBride, A.J.; Ko, A.I.; Reis, M.G. Rattus norvegicus as a model for persistent renal colonization by pathogenic Leptospira interrogans. Acta. Tropica. 2008, 105, 176–180. [Google Scholar] [CrossRef]
- Hathaway, S.C.; Blackmore, D.K. Ecological aspects of the epidemiology of infection with leptospires of the Ballum serogroup in the black rat (Rattus rattus) and the brown rat (Rattus norvegicus) in New Zealand. J. Hyg. (Lond) 1981, 87, 427–436. [Google Scholar] [CrossRef]
- Lloyd-Smith, J.O.; Greig, D.J.; Hietala, S.; Ghneim, G.S.; Palmer, L.; St Leger, J.; Grenfell, B.T.; Gulland, F.M. Cyclical changes in seroprevalence of leptospirosis in California sea lions: endemic and epidemic disease in one host species? BMC Infect. Dis. 2007, 7, 125. [Google Scholar] [CrossRef]
- Ganoza, C.A.; Matthias, M.A.; Saito, M.; Cespedes, M.; Gotuzzo, E.; Vinetz, J.M. Asymptomatic renal colonization of humans in the peruvian Amazon by Leptospira. PLoS Negl. Trop. Dis. 2010, 4, e612. [Google Scholar] [CrossRef]
- Day, T.D.; O'Connor, C.E.; Waas, J.R.; Pearson, A.J.; Matthews, L.R. Transmission of Leptospira interrogans serovar Balcanica infection among socially housed brushtail possums in New Zealand. J. Wildl. Dis. 1998, 34, 576–581. [Google Scholar] [CrossRef]
- Ellis, W.A.; Songer, J.G.; Montgomery, J.; Cassells, J.A. Prevalence of Leptospira interrogans serovar hardjo in the genital and urinary tracts of non-pregnant cattle. Vet. Rec. 1986, 118, 11–13. [Google Scholar]
- Ellis, W.A.; Logan, E.F.; O'Brien, J.J.; Neill, S.D.; Ferguson, H.W.; Hanna, J. Antibodies to Leptospira in the sera of aborted bovine fetuses. Vet. Rec. 1978, 103, 237–239. [Google Scholar]
- Baker, M.F.; Baker, H.J. Pathogenic Leptospira in Malaysian surface waters. I. A method of survey for Leptospira in natural waters and soils. Am. J. Trop. Med. Hyg. 1970, 19, 485–492. [Google Scholar]
- Karaseva, E.V.; Chernukha Yu, G.; Sakhartseva, T.F. Results of the investigation of soil for contamination with pathogenic leptospires. Folia. Parasitol. (Praha) 1977, 24, 301–304. [Google Scholar]
- Ido, Y.; Hoki, R.; Ito, H.; Wani, H. The Rat as a Carrier of Spirochaeta Icterohaemorrhagiae, the Causative Agent of Weil's Disease (Spirochaetosis Icterohaemorrhagica). J. Exp. Med. 1917, 26, 341–353. [Google Scholar] [CrossRef]
- Thiermann, A.B.; Frank, R.R. Human leptospirosis in Detroit and the role of rats as chronic carriers. Int. J. Zoonoses 1980, 7, 62–72. [Google Scholar]
- Vinetz, J.M.; Glass, G.E.; Flexner, C.E.; Mueller, P.; Kaslow, D.C. Sporadic urban leptospirosis. Ann. Intern. Med. 1996, 125, 794–798. [Google Scholar] [CrossRef]
- Everard, C.O.R.; Tikasing, E.S. Ecology of Rodents, Proechimys-Guyannensis-Trinitatis and Oryzomys-Capito-Velutinus, on Trinida. J. Mammal. 1973, 54, 875–886. [Google Scholar] [CrossRef]
- Miller, R.I.; Ross, S.P.; Sullivan, N.D.; Perkins, N.R. Clinical and epidemiological features of canine leptospirosis in North Queensland. Aust. Vet. J. 2007, 85, 13–19. [Google Scholar] [CrossRef]
- Birnbaum, N.; Barr, S.C.; Center, S.A.; Schermerhorn, T.; Randolph, J.F.; Simpson, K.W. Naturally acquired leptospirosis in 36 dogs: serological and clinicopathological features. J. Small Anim. Pract. 1998, 39, 231–236. [Google Scholar] [CrossRef]
- Levett, P.N. Leptospirosis. Clin. Microbiol. Rev. 2001, 14, 296–326. [Google Scholar] [CrossRef]
- Acevedo-Whitehouse, K.; de la Cueva, H.; Gulland, F.M.; Aurioles-Gamboa, D.; Arellano-Carbajal, F.; Suarez-Guemes, F. Evidence of Leptospira interrogans infection in California sea lion pups from the Gulf of California. J. Wildl. Dis. 2003, 39, 145–151. [Google Scholar] [CrossRef]
- Cameron, C.E.; Zuerner, R.L.; Raverty, S.; Colegrove, K.M.; Norman, S.A.; Lambourn, D.M.; Jeffries, S.J.; Gulland, F.M. Detection of pathogenic Leptospira bacteria in pinniped populations via PCR and identification of a source of transmission for zoonotic leptospirosis in the marine environment. J. Clin. Microbiol. 2008, 46, 1728–1733. [Google Scholar] [CrossRef]
- Zuerner, R.L.; Cameron, C.E.; Raverty, S.; Robinson, J.; Colegrove, K.M.; Norman, S.A.; Lambourn, D.; Jeffries, S.; Alt, D.P.; Gulland, F. Geographical dissemination of Leptospira interrogans serovar Pomona during seasonal migration of California sea lions. Vet. Microbiol. 2009, 137, 105–110. [Google Scholar] [CrossRef]
- Colegrove, K.M.; Lowenstine, L.J.; Gulland, F.M. Leptospirosis in northern elephant seals (Mirounga angustirostris) stranded along the California coast. J. Wildl. Dis. 2005, 41, 426–430. [Google Scholar] [CrossRef]
- Matthias, M.A.; Diaz, M.M.; Campos, K.J.; Calderon, M.; Willig, M.R.; Pacheco, V.; Gotuzzo, E.; Gilman, R.H.; Vinetz, J.M. Diversity of bat-associated Leptospira in the Peruvian Amazon inferred by bayesian phylogenetic analysis of 16S ribosomal DNA sequences. Am. J. Trop. Med. Hyg. 2005, 73, 964–974. [Google Scholar]
- Nascimento, A.L.; Ko, A.I.; Martins, E.A.; Monteiro-Vitorello, C.B.; Ho, P.L.; Haake, D.A.; Verjovski-Almeida, S.; Hartskeerl, R.A.; Marques, M.V.; Oliveira, M.C.; et al. Comparative genomics of two Leptospira interrogans serovars reveals novel insights into physiology and pathogenesis. J. Bacteriol. 2004, 186, 2164–2172. [Google Scholar] [CrossRef]
- Fouts, D.E.; Mongodin, E.F.; Mandrell, R.E.; Miller, W.G.; Rasko, D.A.; Ravel, J.; Brinkac, L.M.; DeBoy, R.T.; Parker, C.T.; Daugherty, S.C.; et al. Major structural differences and novel potential virulence mechanisms from the genomes of multiple Campylobacter species. PLoS Biol. 2005, 3, e15. [Google Scholar] [CrossRef] [Green Version]
- Gill, S.R.; Fouts, D.E.; Archer, G.L.; Mongodin, E.F.; Deboy, R.T.; Ravel, J.; Paulsen, I.T.; Kolonay, J.F.; Brinkac, L.; Beanan, M.; et al. Insights on evolution of virulence and resistance from the complete genome analysis of an early methicillin-resistant Staphylococcus aureus strain and a biofilm-producing methicillin-resistant Staphylococcus epidermidis strain. J. Bacteriol. 2005, 187, 2426–2438. [Google Scholar] [CrossRef]
- Nelson, K.E.; Fouts, D.E.; Mongodin, E.F.; Ravel, J.; DeBoy, R.T.; Kolonay, J.F.; Rasko, D.A.; Angiuoli, S.V.; Gill, S.R.; Paulsen, I.T.; et al. Whole genome comparisons of serotype 4b and 1/2a strains of the food-borne pathogen Listeria monocytogenes reveal new insights into the core genome components of this species. Nucleic. Acids. Res. 2004, 32, 2386–2395. [Google Scholar] [CrossRef]
- Chen, Y.; Stine, O.C.; Badger, J.H.; Gil, A.I.; Nair, G.B.; Nishibuchi, M.; Fouts, D.E. Comparative genomic analysis of Vibrio parahaemolyticus: serotype conversion and virulence. BMC Genom. 2011, 12, 294. [Google Scholar] [CrossRef]
- Tettelin, H.; Masignani, V.; Cieslewicz, M.J.; Donati, C.; Medini, D.; Ward, N.L.; Angiuoli, S.V.; Crabtree, J.; Jones, A.L.; Durkin, A.S.; et al. Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: implications for the microbial “pan-genome”. Proc. Natl. Acad. Sci. USA 2005, 102, 13950–13955. [Google Scholar] [CrossRef]
- Losada, L.; Ronning, C.M.; DeShazer, D.; Woods, D.; Fedorova, N.; Kim, H.S.; Shabalina, S.A.; Pearson, T.R.; Brinkac, L.; Tan, P.; et al. Continuing evolution of Burkholderia mallei through genome reduction and large-scale rearrangements. Genome. Biol. Evol. 2010, 2, 102–116. [Google Scholar] [CrossRef]
- Pallen, M.J.; Wren, B.W. Bacterial pathogenomics. Nature 2007, 449, 835–842. [Google Scholar] [CrossRef]
- Gardiner, D.M.; McDonald, M.C.; Covarelli, L.; Solomon, P.S.; Rusu, A.G.; Marshall, M.; Kazan, K.; Chakraborty, S.; McDonald, B.A.; Manners, J.M. Comparative pathogenomics reveals horizontally acquired novel virulence genes in fungi infecting cereal hosts. PLoS Pathog. 2012, 8, e1002952. [Google Scholar] [CrossRef]
- Sudheesh, P.S.; Al-Ghabshi, A.; Al-Mazrooei, N.; Al-Habsi, S. Comparative pathogenomics of bacteria causing infectious diseases in fish. Int. J. Evol. Biol. 2012, 2012, 457264. [Google Scholar]
- Crabtree, J.; Angiuoli, S.V.; Wortman, J.R.; White, O.R. Sybil: Methods and Software for Multiple Genome Comparison and Visualization. In Gene Function Analysis; Ochs, M.F., Ed.; Humana Press Inc.: Totowa, NY, USA, 2007; pp. 93–108. [Google Scholar]
- Li, L.; Stoeckert, C.J., Jr.; Roos, D.S. OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Res. 2003, 13, 2178–2189. [Google Scholar] [CrossRef]
- Remm, M.; Storm, C.E.; Sonnhammer, E.L. Automatic clustering of orthologs and in-paralogs from pairwise species comparisons. J. Mol. Biol. 2001, 314, 1041–1052. [Google Scholar] [CrossRef]
- Fouts, D.E.; Brinkac, L.; Beck, E.; Inman, J.; Sutton, G. PanOCT: Automated Clustering of Orthologs Using Conserved Gene Neighborhood for Pan-Genomic Analysis of Bacterial Strains and Closely Related Species. Nucleic. Acids. Res. 2012, 40, e172. [Google Scholar] [CrossRef]
- Rasko, D.A.; Myers, G.S.; Ravel, J. Visualization of comparative genomic analyses by BLAST score ratio. BMC Bioinformatics 2005, 6, 2. [Google Scholar] [CrossRef] [Green Version]
- Lehmann, J.S.; Fouts, D.E.; Haft, D.H.; Cannella, A.P.; Ricaldi, J.N.; Brinkac, L.; Harkins, D.; Durkin, S.; Sanka, R.; Sutton, G.; et al. Pathogenomic inference of virulence-associated genes in Leptospira interrogans. PLoS negl. Trop. Dis. 2013, 7, e2468. [Google Scholar] [CrossRef]
- Fouts, D.E. Phage_Finder: automated identification and classification of prophage regions in complete bacterial genome sequences. Nucleic Acids Res. 2006, 34, 5839–5851. [Google Scholar] [CrossRef]
- Srividhya, K.V.; Alaguraj, V.; Poornima, G.; Kumar, D.; Singh, G.P.; Raghavenderan, L.; Katta, A.V.; Mehta, P.; Krishnaswamy, S. Identification of prophages in bacterial genomes by dinucleotide relative abundance difference. PloS One 2007, 2, e1193. [Google Scholar]
- Lima-Mendez, G.; Van Helden, J.; Toussaint, A.; Leplae, R. Prophinder: a computational tool for prophage prediction in prokaryotic genomes. Bioinformatics 2008, 24, 863–865. [Google Scholar] [CrossRef]
- Zhou, Y.; Liang, Y.; Lynch, K.H.; Dennis, J.J.; Wishart, D.S. PHAST: A fast phage search tool. Nucleic Acids Res. 2011, 39, W347–W352. [Google Scholar] [CrossRef]
- Akhter, S.; Aziz, R.K.; Edwards, R.A. PhiSpy: A novel algorithm for finding prophages in bacterial genomes that combines similarity- and composition-based strategies. Nucleic Acids Res. 2012, 40, e126. [Google Scholar] [CrossRef]
- Eddy, S.R. A new generation of homology search tools based on probabilistic inference. Genome Inform. 2009, 23, 205–211. [Google Scholar] [CrossRef]
- Finn, R.D.; Clements, J.; Eddy, S.R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 2011, 39, W29–W37. [Google Scholar] [CrossRef]
- Phage_Finder version 2.1. Available online: http://sourceforge.net/projects/phage-finder/files/phage_finder_v2.1/ (Accessed on 2 April 2014).
- Langille, M.G.; Brinkman, F.S. IslandViewer: An integrated interface for computational identification and visualization of genomic islands. Bioinformatics 2009, 25, 664–665. [Google Scholar] [CrossRef]
- Langille, M.G.; Hsiao, W.W.; Brinkman, F.S. IslandViewer: An integrated interface for computational identification and visualization of genomic island predictors using a comparative genomics approachislands. BMC Bioinformatics 2009, 925, 329664–329665. [Google Scholar]
- Langille, M.G.; Hsiao, W.W.; Brinkman, F.S. Evaluation of genomic island predictors using a comparative genomics approach. BMC Bioinformatics 2008, 9, 329. [Google Scholar] [CrossRef]
- Waack, S.; Keller, O.; Asper, R.; Brodag, T.; Damm, C.; Fricke, W.F.; Surovcik, K.; Meinicke, P.; Merkl, R. Score-based prediction of genomic islands in prokaryotic genomes using hidden Markov models. BMC Bioinformatics 2006, 7, 142. [Google Scholar] [CrossRef] [Green Version]
- Aydanian, A.; Tang, L.; Morris, J.G.; Johnson, J.A.; Stine, O.C. Genetic diversity of O-antigen biosynthesis regions in Vibrio cholerae. Appl Environ. Microbiol. 2011, 77, 2247–2253. [Google Scholar] [CrossRef]
- Jacobsen, A.; Hendriksen, R.S.; Aaresturp, F.M.; Ussery, D.W.; Friis, C. The Salmonella enterica pan-genome. Microb. Ecol. 2011, 62, 487–504. [Google Scholar] [CrossRef]
- Rodriguez-Valera, F.; Ussery, D.W. Is the pan-genome also a pan-seletome? F1000 Research 2012, 1, 16. [Google Scholar]
- Wright, A. Mechanism of Conversion of the Salmonella O antigen by Bacteriophage Epsilon 34. J. Bacteriol. 1971, 105, 927–936. [Google Scholar]
- Kropinski, A.M.; Prangishvili, D.; Lavigne, R. Position paper: the creation of a rational scheme for the nomenclature of viruses of Bacteria and Archaea. Environ. Microbiol. 2009, 11, 2775–2777. [Google Scholar] [CrossRef]
- Ricaldi, J.N.; Fouts, D.E.; Selengut, J.D.; Harkins, D.M.; Patra, K.P.; Moreno, A.; Lehmann, J.S.; Purushe, J.; Sanka, R.; Torres, M.; et al. Whole Genome Analysis of Leptospira licerasiae Provides Insight into Leptospiral Evolution and Pathogenicity. PLoS negl. Trop. Dis. 2012, 6, e1853. [Google Scholar] [CrossRef]
- Qin, J.H.; Zhang, Q.; Zhang, Z.M.; Zhong, Y.; Yang, Y.; Hu, B.Y.; Zhao, G.P.; Guo, X.K. Identification of a novel prophage-like gene cluster actively expressed in both virulent and avirulent strains of Leptospira interrogans serovar Lai. Infect. Immun. 2008, 76, 2411–2419. [Google Scholar] [CrossRef]
- Bourhy, P.; Frangeul, L.; Couve, E.; Glaser, P.; Saint Girons, I.; Picardeau, M. Complete nucleotide sequence of the LE1 prophage from the spirochete Leptospira biflexa and characterization of its replication and partition functions. J. Bacteriol. 2005, 187, 3931–3940. [Google Scholar] [CrossRef]
- Girons, I.S.; Bourhy, P.; Ottone, C.; Picardeau, M.; Yelton, D.; Hendrix, R.W.; Glaser, P.; Charon, N. The LE1 bacteriophage replicates as a plasmid within Leptospira biflexa: construction of an L. biflexa-Escherichia coli shuttle vector. J. Bacteriol. 2000, 182, 5700–5705. [Google Scholar] [CrossRef]
- He, P.; Sheng, Y.Y.; Shi, Y.Z.; Jiang, X.G.; Qin, J.H.; Zhang, Z.M.; Zhao, G.P.; Guo, X.K. Genetic diversity among major endemic strains of Leptospira interrogans in China. BMC Genomics 2007, 8, 204. [Google Scholar] [CrossRef]
- Van Melderen, L. Toxin-antitoxin systems: why so many, what for? Curr. Opin. Microbiol. 2010, 13, 781–785. [Google Scholar] [CrossRef]
- Gerdes, K. Toxin-antitoxin modules may regulate synthesis of macromolecules during nutritional stress. J. Bacteriol. 2000, 182, 561–572. [Google Scholar] [CrossRef]
- Engelberg-Kulka, H.; Amitai, S.; Kolodkin-Gal, I.; Hazan, R. Bacterial programmed cell death and multicellular behavior in bacteria. PLoS Genet. 2006, 2, e135. [Google Scholar] [CrossRef]
- Hazan, R.; Engelberg-Kulka, H. Escherichia coli mazEF-mediated cell death as a defense mechanism that inhibits the spread of phage P1. Mol. Genet. Genom.: MGG 2004, 272, 227–234. [Google Scholar]
- Pecota, D.C.; Wood, T.K. Exclusion of T4 phage by the hok/sok killer locus from plasmid R1. J. Bacteriol. 1996, 178, 2044–2050. [Google Scholar]
- Ramage, H.R.; Connolly, L.E.; Cox, J.S. Comprehensive functional analysis of Mycobacterium tuberculosis toxin-antitoxin systems: implications for pathogenesis, stress responses, and evolution. PLoS Genet. 2009, 5, e1000767. [Google Scholar] [CrossRef]
- Picardeau, M.; Ren, S.; Girons, I.S. Killing effect and antitoxic activity of the Leptospira interrogans toxin-antitoxin system in Escherichia coli. J. Bacteriol. 2001, 183, 6494–6497. [Google Scholar] [CrossRef]
- Picardeau, M.; Le Dantec, C.; Richard, G.F.; Saint Girons, I. The spirochetal chpK-chromosomal toxin-antitoxin locus induces growth inhibition of yeast and mycobacteria. FEMS Microbiol. Lett. 2003, 229, 277–281. [Google Scholar] [CrossRef]
- Picardeau, M.; Bulach, D.M.; Bouchier, C.; Zuerner, R.L.; Zidane, N.; Wilson, P.J.; Creno, S.; Kuczek, E.S.; Bommezzadri, S.; Davis, J.C.; et al. Genome sequence of the saprophyte Leptospira biflexa provides insights into the evolution of Leptospira and the pathogenesis of leptospirosis. PloS One 2008, 3, e1607. [Google Scholar] [CrossRef]
- Bourhy, P.; Salaun, L.; Lajus, A.; Medigue, C.; Boursaux-Eude, C.; Picardeau, M. A genomic island of the pathogen Leptospira interrogans serovar Lai can excise from its chromosome. Infect. Immun. 2007, 75, 677–683. [Google Scholar] [CrossRef]
- Boursaux-Eude, C.; Saint Girons, I.; Zuerner, R. IS1500, an IS3-like element from Leptospira interrogans. Microbiology 1995, 141, 2165–2173. [Google Scholar] [CrossRef]
- Zuerner, R.L.; Trueba, G.A. Characterization of IS1501 mutants of Leptospira interrogans serovar pomona. FEMS Microbiol. Lett. 2005, 248, 199–205. [Google Scholar] [CrossRef]
- Zuerner, R.L. Nucleotide sequence analysis of IS1533 from Leptospira borgpetersenii: identification and expression of two IS-encoded proteins. Plasmid 1994, 31, 1–11. [Google Scholar] [CrossRef]
- Zhong, Y.; Chang, X.; Cao, X.J.; Zhang, Y.; Zheng, H.; Zhu, Y.; Cai, C.; Cui, Z.; Zhang, Y.; Li, Y.Y.; et al. Comparative proteogenomic analysis of the Leptospira interrogans virulence-attenuated strain IPAV against the pathogenic strain 56601. Cell. Res. 2011, 21, 1210–1229. [Google Scholar] [CrossRef]
- Bulach, D.M.; Kalambaheti, T.; de la Pena-Moctezuma, A.; Adler, B. Lipopolysaccharide biosynthesis in Leptospira. J. Mol. Microbiol. Biotechnol. 2000, 2, 375–380. [Google Scholar]
- Bulach, D.M.; Zuerner, R.L.; Wilson, P.; Seemann, T.; McGrath, A.; Cullen, P.A.; Davis, J.; Johnson, M.; Kuczek, E.; Alt, D.P.; et al. Genome reduction in Leptospira borgpetersenii reflects limited transmission potential. Proc. Natl. Acad. Sci. USA 2006, 103, 14560–14565. [Google Scholar] [CrossRef]
- Chandler, M.G.; Pritchard, R.H. The effect of gene concentration and relative gene dosage on gene output in Escherichia coli. Molecular & general genetics: MGG 1975, 138, 127–141. [Google Scholar]
- Schmid, M.B.; Roth, J.R. Gene location affects expression level in Salmonella typhimurium. J. Bacteriol. 1987, 169, 2872–2875. [Google Scholar]
- Sousa, C.; de Lorenzo, V.; Cebolla, A. Modulation of gene expression through chromosomal positioning in Escherichia coli. Microbiology 1997, 143, 2071–2078. [Google Scholar] [CrossRef]
- Thompson, A.; Gasson, M.J. Location effects of a reporter gene on expression levels and on native protein synthesis in Lactococcus lactis and Saccharomyces cerevisiae. Appl. Environ. Microbiol. 2001, 67, 3434–3439. [Google Scholar]
- Dryselius, R.; Izutsu, K.; Honda, T.; Iida, T. Differential replication dynamics for large and small Vibrio chromosomes affect gene dosage, expression and location. BMC Genomics 2008, 9, 559. [Google Scholar] [CrossRef]
- Block, D.H.; Hussein, R.; Liang, L.W.; Lim, H.N. Regulatory consequences of gene translocation in bacteria. Nucleic Acids Res. 2012, 40, 8979–8992. [Google Scholar] [CrossRef]
- de la Pena-Moctezuma, A.; Bulach, D.M.; Kalambaheti, T.; Adler, B. Comparative analysis of the LPS biosynthetic loci of the genetic subtypes of serovar Hardjo: Leptospira interrogans subtype Hardjoprajitno and Leptospira borgpetersenii subtype Hardjobovis. FEMS Microbiol. Lett. 1999, 177, 319–326. [Google Scholar] [CrossRef]
- Alikhan, N.F.; Petty, N.K.; Ben Zakour, N.L.; Beatson, S.A. BLAST Ring Image Generator (BRIG): simple prokaryote genome comparisons. BMC Genomics 2011, 12, 402. [Google Scholar] [CrossRef]
- Faruque, S.M.; Sack, D.A.; Sack, R.B.; Colwell, R.R.; Takeda, Y.; Nair, G.B. Emergence and evolution of Vibrio cholerae O139. Proc. Natl. Acad. Sci. USA 2003, 100, 1304–1309. [Google Scholar] [CrossRef]
- Blokesch, M.; Schoolnik, G.K. Serogroup conversion of Vibrio cholerae in aquatic reservoirs. PLoS Pathog. 2007, 3, e81. [Google Scholar] [CrossRef]
- Ricaldi, N.J.; Matthias, M.A.; Vinetz, J.M.; Lewis, A.L. Expression of sialic acids and other nonulosonic acids in Leptospira. BMC Microbiol. 2012, 12, 161. [Google Scholar] [CrossRef]
- Fernandez-Moreira, E.; Helbig, J.H.; Swanson, M.S. Membrane vesicles shed by Legionella pneumophila inhibit fusion of phagosomes with lysosomes. Infect. Immun. 2006, 74, 3285–3295. [Google Scholar] [CrossRef]
- Meri, T.; Murgia, R.; Stefanel, P.; Meri, S.; Cinco, M. Regulation of complement activation at the C3-level by serum resistant leptospires. Microb. Pathog. 2005, 39, 139–147. [Google Scholar] [CrossRef]
- Hathaway, S.C.; Marshall, R.B.; Little, T.W.; Headlam, S.A.; Winter, P.J. Differentiation of reference strains of leptospires of the Pomona serogroup by cross-agglutination absorption and restriction endonuclease analysis. Res. Vet. Sci. 1985, 39, 145–150. [Google Scholar]
- Lo, M.; Bulach, D.M.; Powell, D.R.; Haake, D.A.; Matsunaga, J.; Paustian, M.L.; Zuerner, R.L.; Adler, B. Effects of temperature on gene expression patterns in Leptospira interrogans serovar Lai as assessed by whole-genome microarrays. Infect. Immun. 2006, 74, 5848–5859. [Google Scholar] [CrossRef]
- Xue, F.; Dong, H.; Wu, J.; Wu, Z.; Hu, W.; Sun, A.; Troxell, B.; Yang, X.F.; Yan, J. Transcriptional responses of Leptospira interrogans to host innate immunity: significant changes in metabolism, oxygen tolerance, and outer membrane. PLoS Neglect. Trop. Dis. 2010, 4, e857. [Google Scholar] [CrossRef]
- Wozniak, R.A.; Fouts, D.E.; Spagnoletti, M.; Colombo, M.M.; Ceccarelli, D.; Garriss, G.; Dery, C.; Burrus, V.; Waldor, M.K. Comparative ICE Genomics: Insights into the Evolution of the SXT/R391 Family of ICEs. PLoS Genet. 2009, 5, e1000786. [Google Scholar] [CrossRef]
- Patarakul, K.; Lo, M.; Adler, B. Global transcriptomic response of Leptospira interrogans serovar Copenhageni upon exposure to serum. BMC Microbiol. 2010, 10, 31. [Google Scholar] [CrossRef]
- Lo, M.; Murray, G.L.; Khoo, C.A.; Haake, D.A.; Zuerner, R.L.; Adler, B. Transcriptional response of Leptospira interrogans to iron limitation and characterization of a PerR homolog. Infect. Immun. 2010, 78, 4850–4859. [Google Scholar] [CrossRef]
- Matsunaga, J.; Lo, M.; Bulach, D.M.; Zuerner, R.L.; Adler, B.; Haake, D.A. Response of Leptospira interrogans to physiologic osmolarity: relevance in signaling the environment-to-host transition. Infect. Immun. 2007, 75, 2864–2874. [Google Scholar] [CrossRef]
- Ko, A.I.; Goarant, C.; Picardeau, M. Leptospira: the dawn of the molecular genetics era for an emerging zoonotic pathogen. Nat. Rev. Microbiol. 2009, 7, 736–747. [Google Scholar] [CrossRef]
- Adler, B.; Lo, M.; Seemann, T.; Murray, G.L. Pathogenesis of leptospirosis: The influence of genomics. Vet. Microbiol. 2011, 153, 73–81. [Google Scholar] [CrossRef]
- Narayanavari, S.A.; Sritharan, M.; Haake, D.A.; Matsunaga, J. Multiple leptospiral sphingomyelinases (or are there?). Microbiology 2012, 158, 1137–1146. [Google Scholar] [CrossRef]
- Bourhy, P.; Louvel, H.; Girons, I.S.; Picardeau, M. Random insertional mutagenesis of Leptospira interrogans, the agent of leptospirosis, using a mariner transposon. J. Bacteriol. 2005, 187, 3255–3258. [Google Scholar] [CrossRef]
- Murray, G.L.; Srikram, A.; Henry, R.; Hartskeerl, R.A.; Sermswan, R.W.; Adler, B. Mutations affecting Leptospira interrogans lipopolysaccharide attenuate virulence. Mol. Microbiol. 2010, 78, 701–709. [Google Scholar] [CrossRef]
- Nally, J.E.; Whitelegge, J.P.; Bassilian, S.; Blanco, D.R.; Lovett, M.A. Characterization of the outer membrane proteome of Leptospira interrogans expressed during acute lethal infection. Infect. Immun. 2007, 75, 766–773. [Google Scholar] [CrossRef]
- Lin, Y.P.; Chang, Y.F. A domain of the Leptospira LigB contributes to high affinity binding of fibronectin. Biochem. Biophys. Res. Comm. 2007, 362, 443–448. [Google Scholar] [CrossRef]
- Murray, G.L.; Morel, V.; Cerqueira, G.M.; Croda, J.; Srikram, A.; Henry, R.; Ko, A.I.; Dellagostin, O.A.; Bulach, D.M.; Sermswan, R.W.; et al. Genome-wide transposon mutagenesis in pathogenic Leptospira species. Infect. Immun. 2009, 77, 810–816. [Google Scholar] [CrossRef]
- Ahuja, N.; Kumar, P.; Bhatnagar, R. The adenylate cyclase toxins. Crit. Rev. Microbiol. 2004, 30, 187–196. [Google Scholar] [CrossRef]
- Li, L.; Ojcius, D.M.; Yan, J. Comparison of invasion of fibroblasts and macrophages by high- and low-virulence Leptospira strains: colonization of the host-cell nucleus and induction of necrosis by the virulent strain. Arch. Microbiol. 2007, 188, 591–598. [Google Scholar] [CrossRef]
- Bensing, B.A.; Rubens, C.E.; Sullam, P.M. Genetic loci of Streptococcus mitis that mediate binding to human platelets. Infect. Immun. 2001, 69, 1373–1380. [Google Scholar] [CrossRef]
- Bensing, B.A.; Siboo, I.R.; Sullam, P.M. Proteins PblA and PblB of Streptococcus mitis, which promote binding to human platelets, are encoded within a lysogenic bacteriophage. Infect. Immun. 2001, 69, 6186–6192. [Google Scholar] [CrossRef]
- Mitchell, J.; Sullam, P.M. Streptococcus mitis phage-encoded adhesins mediate attachment to {alpha}2–8-linked sialic acid residues on platelet membrane gangliosides. Infect. Immun. 2009, 77, 3485–3490. [Google Scholar] [CrossRef]
- Edwards, C.N.; Nicholson, G.D.; Hassell, T.A.; Everard, C.O.; Callender, J. Thrombocytopenia in leptospirosis: the absence of evidence for disseminated intravascular coagulation. Am. J. Trop. Med. Hyg. 1986, 35, 352–354. [Google Scholar]
- Marotto, P.C.; Marotto, M.S.; Santos, D.L.; Souza, T.N.; Seguro, A.C. Outcome of leptospirosis in children. Am. J. Trop. Med. Hyg. 1997, 56, 307–310. [Google Scholar]
- Nicodemo, A.C.; Del Negro, G.; Amato Neto, V. Thrombocytopenia and leptospirosis. Rev. Inst. Med. Trop. Sao Paulo 1990, 32, 252–259. [Google Scholar]
- Marcsisin, R.A.; Bartpho, T.; Bulach, D.M.; Srikram, A.; Sermswan, R.W.; Adler, B.; Murray, G.L. Use of a high-throughput screen to identify Leptospira mutants unable to colonize the carrier host or cause disease in the acute model of infection. J. Med. Microbiol. 2013, 62, 1601–1608. [Google Scholar] [CrossRef]
- Guy, L.; Nystedt, B.; Toft, C.; Zaremba-Niedzwiedzka, K.; Berglund, E.C.; Granberg, F.; Naslund, K.; Eriksson, A.S.; Andersson, S.G. A gene transfer agent and a dynamic repertoire of secretion systems hold the keys to the explosive radiation of the emerging pathogen Bartonella. PLoS Genet 2013, 9, e1003393. [Google Scholar] [CrossRef]
- Choy, H.A.; Kelley, M.M.; Chen, T.L.; Moller, A.K.; Matsunaga, J.; Haake, D.A. Physiological osmotic induction of Leptospira interrogans adhesion: LigA and LigB bind extracellular matrix proteins and fibrinogen. Infect. Immun. 2007, 75, 2441–2450. [Google Scholar] [CrossRef]
- Perez, J.; Goarant, C. Rapid Leptospira identification by direct sequencing of the diagnostic PCR products in New Caledonia. BMC Microbiol. 2010, 10, 325. [Google Scholar] [CrossRef]
- Oliveira, R.; de Morais, Z.M.; Goncales, A.P.; Romero, E.C.; Vasconcellos, S.A.; Nascimento, A.L. Characterization of novel OmpA-like protein of Leptospira interrogans that binds extracellular matrix molecules and plasminogen. PloS One 2011, 6, e21962. [Google Scholar]
- Yan, W.; Faisal, S.M.; McDonough, S.P.; Divers, T.J.; Barr, S.C.; Chang, C.F.; Pan, M.J.; Chang, Y.F. Immunogenicity and protective efficacy of recombinant Leptospira immunoglobulin-like protein B (rLigB) in a hamster challenge model. Microbes Infect. 2009, 11, 230–237. [Google Scholar] [CrossRef]
- Verma, A.; Rathinam, S.R.; Priya, C.G.; Muthukkaruppan, V.R.; Stevenson, B.; Timoney, J.F. LruA and LruB antibodies in sera of humans with leptospiral uveitis. Clin. Vaccine. Immunol. CVI 2008, 15, 1019–1023. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Lehmann, J.S.; Matthias, M.A.; Vinetz, J.M.; Fouts, D.E. Leptospiral Pathogenomics. Pathogens 2014, 3, 280-308. https://doi.org/10.3390/pathogens3020280
Lehmann JS, Matthias MA, Vinetz JM, Fouts DE. Leptospiral Pathogenomics. Pathogens. 2014; 3(2):280-308. https://doi.org/10.3390/pathogens3020280
Chicago/Turabian StyleLehmann, Jason S., Michael A. Matthias, Joseph M. Vinetz, and Derrick E. Fouts. 2014. "Leptospiral Pathogenomics" Pathogens 3, no. 2: 280-308. https://doi.org/10.3390/pathogens3020280





