Unconventional Cadherin Localization in Honey Bee Gonads Revealed Through Domain-Specific Apis mellifera E- and N-Cadherin Antibodies Indicates Alternative Functions
Abstract
:1. Introduction
2. Results and Discussion
2.1. Protein Identification by Western Blot Analysis
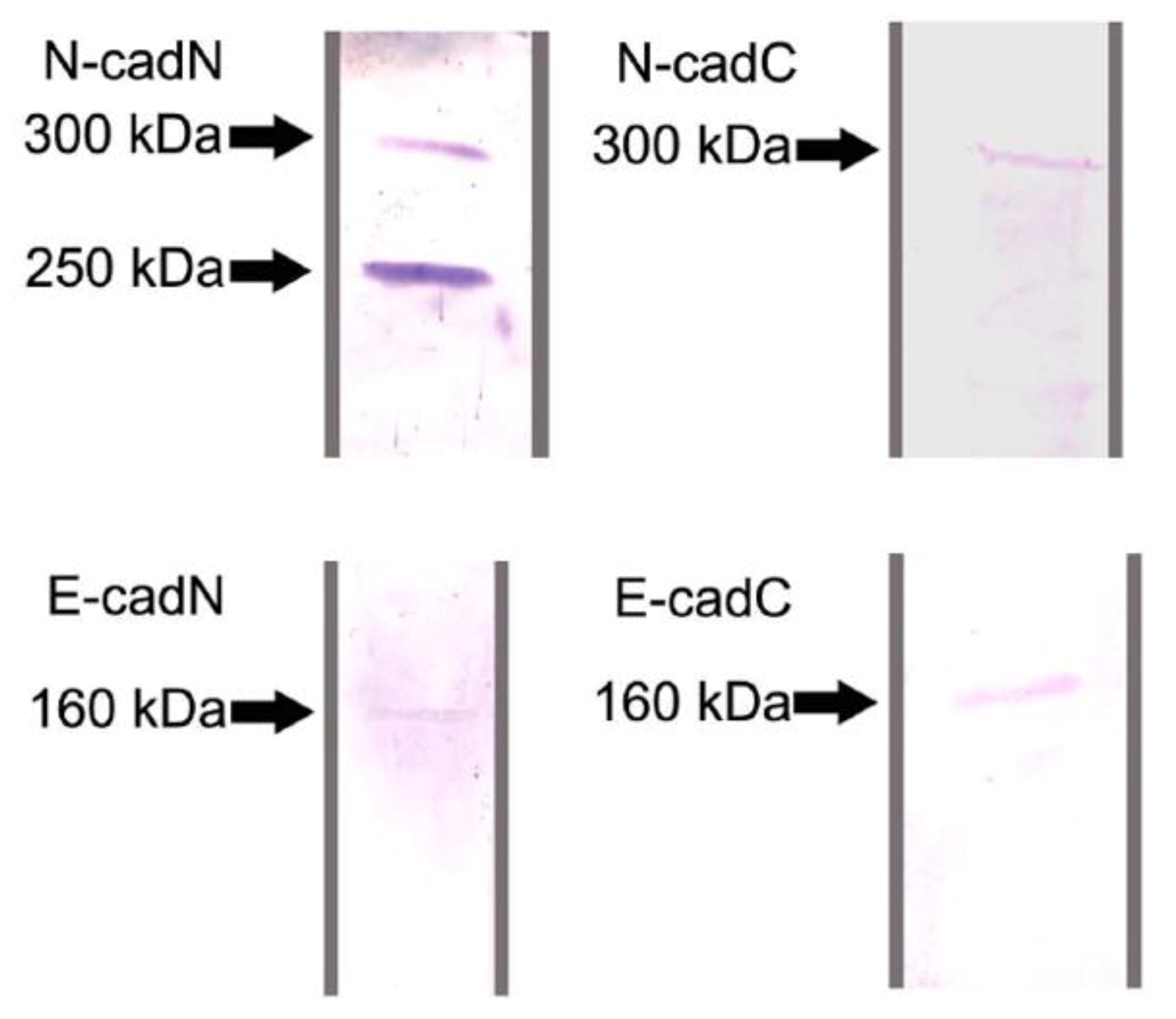
2.2. Differential Localization of Cadherin Domains in Queen and Worker Ovaries Indicates Alternative Functions Related to Fertility Status
2.2.1. Immunolocalization of the N- and C-terminal N-cadherin Domains (N-cadN and N-cadC) in Honey Bee Ovaries
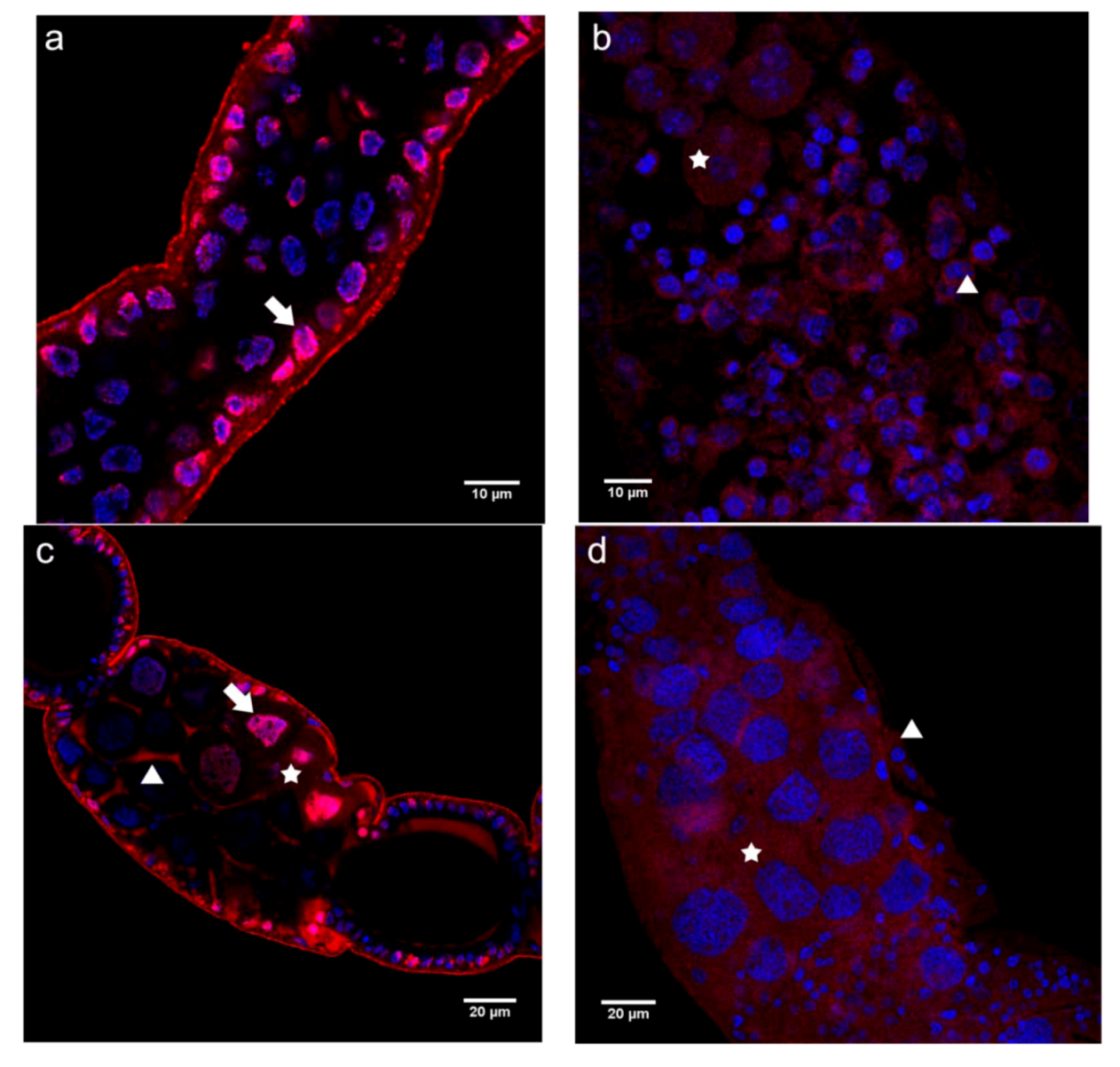


2.2.2. Immunolocalization of the N- and C-terminal E-cadherin Domains (E-cadN and E-cadC) in Honey Bee Ovaries
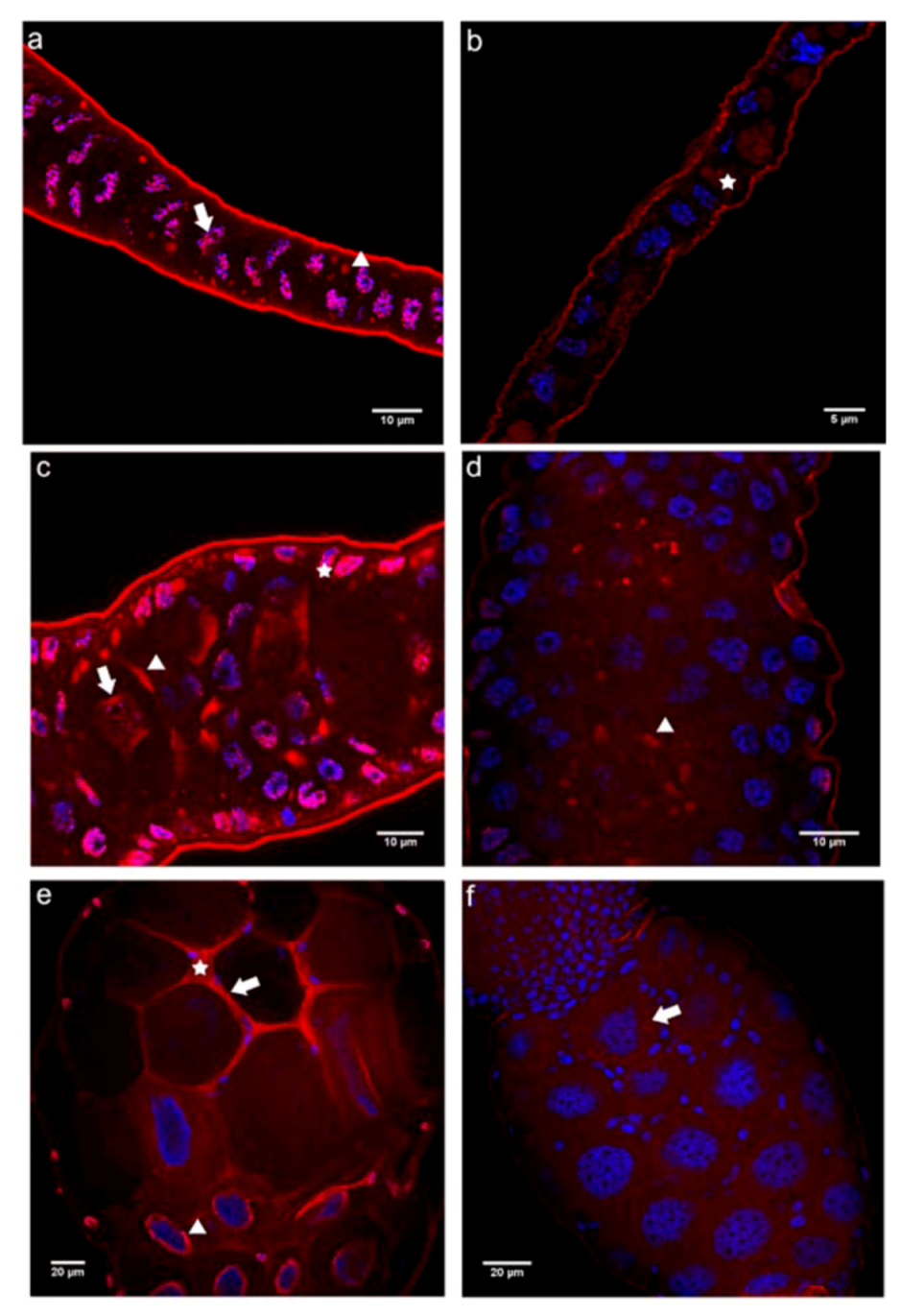
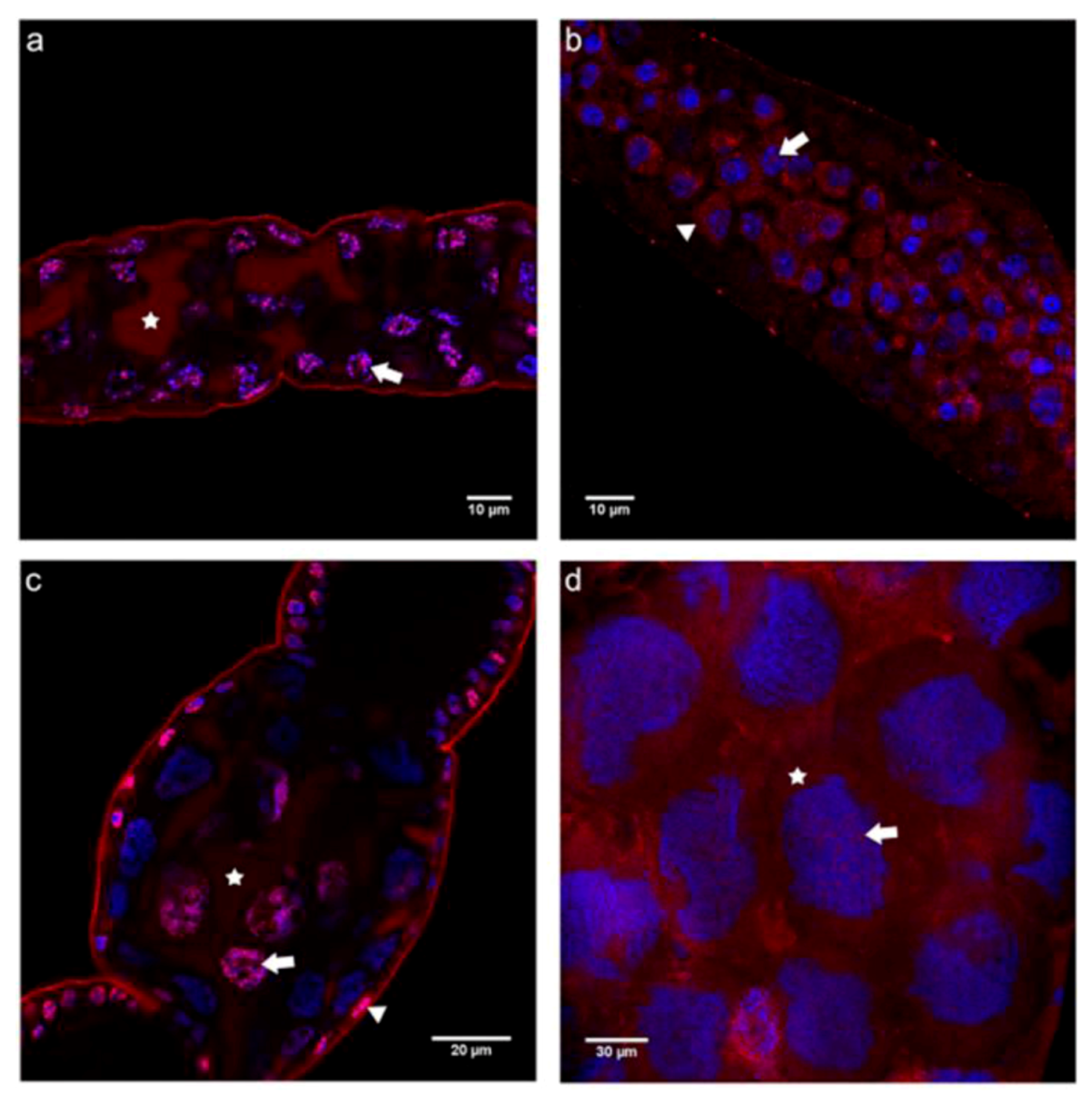
2.3. Cadherin Immunolocalization in Testes of Apis mellifera Drones - Similarities and Differences between the Sexes

3. Experimental Section
3.1. Bees
3.2. Honey Bee E- and N-cadherin Gene Predictions
3.3. RT-PCR Analysis
3.4. In vitro Expression of the N-cadherin Cytoplasmic Domain
3.5. Synthetic Peptides
3.6. Antibody Production
3.7. Dot Blot and Western Blot Analyses
3.8. Cadherin Immunolocalization in Apis mellifera Gonads
4. Conclusions
Acknowledgments
References and Notes
- Engels, W. Occurrence and significance of vitellogenins in female castes of social Hymenoptera. Am. Zool. 1974, 14, 1229–1237. [Google Scholar]
- Hartfelder, K.; Engels, W. Social insect polymorphism: Hormonal regulation of plasticity in development and reproduction in the honeybee. Curr. Topics Dev. Biol. 1998, 40, 45–77. [Google Scholar] [CrossRef]
- Gutzeit, H.O.; Zissler, D.; Fleig, R. Oogenesis in the honeybee Apis mellifera: Cytological observations on the formation and differentiation of previtellogenic ovarian follicles. Rouxs Arch. Dev. Biol. 1993, 202, 181–191. [Google Scholar] [CrossRef]
- Snodgrass, R.E. Anatomy of the Honey Bee; Cornell University Press: Ithaca, NY, USA, 1956; p. XIV, 334. [Google Scholar]
- Dearden, P. Germ cell development in the honeybee (Apis mellifera); vasa and nanos expression. BMC Dev. Biol. 2006, 6, e6. [Google Scholar] [CrossRef]
- Tanaka, E.D.; Hartfelder, K. The initial stages of oogenesis and their relation to differential fertility in the honey bee Apis mellifera) castes. Arthrop. Struct. Dev. 2004, 33, 431–442. [Google Scholar] [CrossRef]
- Tanaka, E.D.; Hartfelder, K. Sequence and expression pattern of the germ line marker vasa in honey bees and stingless bees. Genet. Mol. Biol. 2009, 32, 582–583. [Google Scholar]
- Schmidt-Capella, I.C.; Hartfelder, K. Juvenile-hormone-dependent interaction of actin and spectrin is crucial for polymorphic differentiation of the larval honey bee ovary. Cell Tissue Res. 2002, 307, 265–272. [Google Scholar]
- Büning, J. The Insect Ovary; Chapman & Hall: London, UK, 1994; p. 400. [Google Scholar]
- Di Carlo, A.; De Felici, M. A role for e-cadherin in mouse primordial germ cell development. Dev. Biol. 2000, 226, 209–219. [Google Scholar] [CrossRef]
- Godt, D.; Tepass, U. Drosophila oocyte localization is mediated by differential cadherin-based adhesion. Nature 1998, 395, 387–391. [Google Scholar] [CrossRef]
- Gilboa, L.; Lehmann, R. How different is venus from mars? The genetics of germ-line stem cells in Drosophila females and males. Development 2004, 131, 4895–4905. [Google Scholar] [CrossRef]
- Gonzáles-Reyes, A. Stem cells, niches and cadherins: A view from Drosophila. J. Cell Sci. 2003, 116, 949–954. [Google Scholar] [CrossRef]
- Song, X.; Zhu, C.; Doan, C.; Xie, T. Germline stem cells anchored by adherens junctions in the Drosophila ovary niches. Science 2002, 296, 1855–1857. [Google Scholar] [CrossRef]
- Tazuke, S.I.; Schulz, C.; Gilboa, L.; Fogarty, M.; Mahowald, A.P.; Guichet, A.; Ephrussi, A.; Wood, C.G.; Lehmann, R.; Fuller, M.T. A germline-specific gap junction protein required for survival of differentiating early germ cells. Development 2002, 129, 2529–2539. [Google Scholar]
- Yamashita, Y.M.; Jones, D.L.; Fuller, M.T. Orientation of asymmetric stem cell division by the APC tumor suppressor and centrosome. Science 2003, 301, 1547–1550. [Google Scholar] [CrossRef]
- de Cuevas, M.; Matunis, E.L. The stem cell niche: Lessons from the Drosophila testis. Development 2011, 138, 2861–2869. [Google Scholar] [CrossRef]
- Oda, H.; Uemura, T.; Harada, Y.; Iwai, Y.; Takeichi, M. A Drosophila homolog of cadherin associated with armadillo and essential for embryonic cell-cell adhesion. Dev. Biol. 1994, 165, 716–726. [Google Scholar] [CrossRef]
- Peifer, M.; Wieschaus, E. The segment polarity gene armadillo encodes a functionally modular protein that is the Drosophila homolog of human plakoglobin. Cell 1990, 63, 1167–1178. [Google Scholar] [CrossRef]
- Xie, T.; Spradling, A.C. Decapentaplegic is essential for the maintenance and division of germline stem cells in the Drosophila ovary. Cell 1998, 94, 251–260. [Google Scholar] [CrossRef]
- King, J.J.; Szakmary, A.; Cox, D.N.; Lin, H. Yb modulates the division of both germline and somatic cells through piwi and hh-mediated mechanisms in the Drosophila ovary. Mol. Cell 2001, 7, 497–508. [Google Scholar] [CrossRef]
- Niewiadomska, P.; Godt, D.; Tepass, U. DE-cadherin is required for intercellular motility during Drosophila oogenesis. J. Cell. Biol. 1999, 144, 533–547. [Google Scholar] [CrossRef]
- Zartman, J.J.; Kanodia, J.S.; Yakoby, N.; Schafer, X.; Watson, C.; Schlichting, K.; Dahmann, C.; Shvartsman, S.Y. Expression patterns of cadherin genes in Drosophila oogenesis. Gene Expr. Patterns 2009, 9, 31–36. [Google Scholar] [CrossRef]
- Schlichting, K.; Wilsch-Bräuninger, M.; Demontis, F.; Dahmann, C. Cadherin Cad99c is required for normal microvilli morphology in Drosophila follicle cells. J. Cell Sci. 2006, 119, 1184–1195. [Google Scholar]
- Le Bras, S.; Van Doren, M. Development of the male germline stem cell niche in Drosophila. Dev. Biol. 2006, 294, 92–103. [Google Scholar] [CrossRef]
- Renkawitz-Pohl, R.; Hempel, L.; Hollmann, M.; Schäfer, M.S. Spermatogenesis. In Comprehensive Insect Molecular Science; Gilbert, L.I., Iatrou, K., Gill, S., Eds.; Elsevier: Oxford, UK, 2005; Volume 1, pp. 157–177. [Google Scholar]
- Beye, M.; Hasselmann, M.; Fondrk, M.K.; Page, R.E., Jr; Omholt, S.W. The gene csd is the primary signal for sexual development in the honeybee and encodes an SR-type protein. Cell 2003, 114, 419–429. [Google Scholar] [CrossRef]
- Meves, F. Die Spermatocytenteilung bei der Honigbiene (Apis mellifica L.) nebst Bemerkungen über die Chromatinreduktion. Arch. Mikroskop. Anat. 1997, 70, 414–491. [Google Scholar] [CrossRef]
- Cruz-Landim, C.; Beig, D.; Silva de Moraes, R.L.M. The process of differentiation during spermatogenesis in bees (Hymenoptera, Apidae). Caryologia 1980, 33, 1–15. [Google Scholar]
- Hoage, T.R.; Kessel, R.G. An electron microscope study of the process of differentiation during spermatogenesis in the drone honey bee (Apis mellifera L.) with special reference to centriole replication and elimination. J. Ultrastruct. 1968, 24, 6–32. [Google Scholar]
- Kerr, W.E.; Silveira, Z.V. A note on the formation of honeybee spermatozoa. J. Apic. Res. 1974, 13, 121–126. [Google Scholar]
- Nachtsheim, H. Cytologische Studien über die Geschlechtsbestimmung bei der Honigbiene (Apis mellifica L.). Arch. Zellforsch. 1913, 11, 169–241. [Google Scholar]
- Florecki, M.M.; Hartfelder, K. Cytoplasmic and nuclear localization of cadherin in honey bee (Apis mellifera L.) gonads. Cell Biol. Int. 2011, 35, 45–49. [Google Scholar] [CrossRef]
- Hsu, S.N.; Yonekura, S.; Ting, C.Y.; Robertson, H.M.; Iwai, Y.; Uemura, T.; Lee, C.H.; Chiba, A. Conserved alternative splicing and expression patterns of arthropod N-cadherin. PLoS Genetics 2009, 5, e1000441. [Google Scholar] [CrossRef]
- Uemura, K.; Kihara, T.; Kuzuya, A.; Okawa, K.; Nishimoto, T.; Bito, H.; Ninomyia, H.S.; Sugimoto, H.; Kinoshita, A.; Shimohama, S. Activity-dependent regulation of β-catenin via ε-cleavage of N-cadherin. Biochem. Biophys. Res. Commun. 2006a, 345, 951–958. [Google Scholar] [CrossRef]
- Uemura, K.; Takeshi, K.; Kuzuya, A.; Okawa, K.; Nishimoto, T.; Ninomyia, H.S.; Sugimoto, H.; Kinoshita, A.; Shimohama, S. Characterization of sequential N-cadherin cleavage by ADAM10 and PS1. Neurosci. Lett. 2006b, 402, 278–283. [Google Scholar] [CrossRef]
- Marambaud, P.; Wen, P.H.; Dutt, A.; Shioi, J.; Takashima, A.; Siman, R.; Robakis, N.K. A CBP binding transcriptional repressor produced by the PS1/epsilon-cleavage of N-cadherin is inhibited by PS1 FAD mutations. Cell 2003, 114, 635–645. [Google Scholar] [CrossRef]
- Theurkauf, W.; Alberts, B.M.; Jan, Y.N.; Jongens, T.A. A central role for microtubules in the differentiation of Drosophila oocytes. Development 1993, 118, 1169–1180. [Google Scholar]
- Bilinski, S.; Jaglarz, M. Organization and possible functions of microtubule cytoskeleton in hymenopteran nurse cells. Cell Motil. Cytoskeleton 1999, 43, 213–220. [Google Scholar]
- Marambaud, P.; Shioi, J.; Serban, G.; Georgakopoulos, A.; Sarner, S.; Nagy, V.; Baki, L.; Wen, P.; Efthimiopoulos, S.; Shao, Z.; et al. A presenilin-1/gammasecretase cleavage releases the E-cadherin intracellular domain and regulates disassembly of adherens junctions. EMBO J. 2002, 21, 1948–1956. [Google Scholar] [CrossRef]
- Ferber, E.C.; Kajita, M.; Wadlow, A.; Tobiansky, L.; Niessen, C.; Ariga, H.; Daniel, J.; Fujita, Y. A role for the cleaved cytoplasmic domain of E-cadherin in the nucleus. J. Biol. Chem. 2008, 283, 12691–12700. [Google Scholar]
- Baki, L.; Marambaud, P.; Efthimiopoulos, S.; Georgakopoulos, A.; Wen, P.; Cui, W.; Shioi, J.; Koo, E.; Ozawa, M.; Friedrich, V.L.; et al. Presenilin-1 binds cytoplasmic epithelial cadherin, inhibits cadherin/p120 association, and regulates stability and function of the cadherin/catenin adhesion complex. Proc. Natl. Acad. Sci. USA 2001, 98, 2381–2386. [Google Scholar]
- Lin, H.; Yue, L.; Spradling, A.C. The Drosophila fusome, a germline-specific organelle, contains membrane skeletal proteins and functions in cyst formation. Development 1994, 120, 947–956. [Google Scholar]
- The Honey Bee Genome Sequencing Consortium. Insights into social insects from the genome of the honeybee Apis mellifera. Nature 2006, 443, 931–948. [Google Scholar] [CrossRef]
- Rutherford, K.M.; Parkhill, J.; Crook, J.; Horsnell, T.; Rice, P.; Rajandream, M.-A.; Barrell, B. Artemis: Sequence visualisation and annotation. Bioinformatics 2000, 16, 944–945. [Google Scholar] [CrossRef]
- ExPASy Bioinformatics Resource Portal. Available online: http://www.expasy.org (accessed on 6 November 2012).
- Smith, P.K.; Krohn, R.; Hermanson, G.T.; Mallia, A.K.; Gartner, F.H.; Provenzano, M.D.; Fujimoto, E.K.; Goeke, N.M.; Olson, B.J.; Klenk, D.C. Measurement of protein using bicinchoninic acid. Anal. Biochem. 1985, 150, 76–85. [Google Scholar]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage t4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef]
- Rachinsky, A.; Hartfelder, K. In vitro biosynthesis of juvenile hormone in larval honey bees: Comparison of six media. In Vitro Cell Dev. Biol. Anim. 1998, 34, 646–648. [Google Scholar] [CrossRef]
- Arnold, G.; Leconte, Y.; Trouiller, J.; Hervet, H.; Chappe, B.; Masson, C. Inhibition of worker honeybee ovaries development by a mixture of fatty acid esters from larvae. C. R. Acad. Sci. III 1994, 317, 511–515. [Google Scholar]
- Oldroyd, B.P.; Smolenski, A.J.; Cornuet, J.-M.; Crozier, R.H. Anarchy in the beehive. Nature 1994, 371, 749. [Google Scholar]
- Ratnieks, F.L.W.; Visscher, P.K. Worker policing in the honeybee. Nature 1989, 342, 796–797. [Google Scholar] [CrossRef]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Florecki, M.M.; Hartfelder, K. Unconventional Cadherin Localization in Honey Bee Gonads Revealed Through Domain-Specific Apis mellifera E- and N-Cadherin Antibodies Indicates Alternative Functions. Insects 2012, 3, 1200-1219. https://doi.org/10.3390/insects3041200
Florecki MM, Hartfelder K. Unconventional Cadherin Localization in Honey Bee Gonads Revealed Through Domain-Specific Apis mellifera E- and N-Cadherin Antibodies Indicates Alternative Functions. Insects. 2012; 3(4):1200-1219. https://doi.org/10.3390/insects3041200
Chicago/Turabian StyleFlorecki, Mônica M., and Klaus Hartfelder. 2012. "Unconventional Cadherin Localization in Honey Bee Gonads Revealed Through Domain-Specific Apis mellifera E- and N-Cadherin Antibodies Indicates Alternative Functions" Insects 3, no. 4: 1200-1219. https://doi.org/10.3390/insects3041200



