Simple Organics and Biomonomers Identified in HCN Polymers: An Overview
Abstract
:1. Introduction
2. Amino Acids
| Compound | Starting material, c (M) | T (°C)/t/Catalyst | Final product analysed | Hydrolysis | Method of identification | Reference |
|---|---|---|---|---|---|---|
| Glycine | HCN, 0.2 | 100/1 d/- | Raw mixture | Acid | GC-MS | [75] |
| NH4CN, 1 | 90/4 h/- | Brown precipitate | Acid | AAA | [76] | |
| HCN, 1 | 90/4 h/NH4OH (pH 9.58) | Black solid | Acid | 2D-PC | [72] | |
| HCN, 1.5 | 90/18 h/NH3 | Soluble oligomers | Acid | 2D-PC, AAA | [77] | |
| HCN, 1.5 | 70/5 d/NH4OH | Soluble oligomers | Non hydrolysis | PC | [78] | |
| HCN, 2.2 | 70/25 d/NH4OH | Soluble oligomers | Non hydrolysis | PC | [63] | |
| NaCN, 1 | 38/3–30 d/NH4Cl | Black solid | Acid | GC-MS | [46] | |
| HCN(L) | r.t./4 w/anhydrous NH3 | Dark solid residue | Acid | AAA | [51] | |
| NaCN, 1 | r.t./3 m/pH 9.2 (HCl) | Soluble oligomers | Acid, basic and neutral | GC-MS | [79] | |
| HCN, 0.1 | r.t./1–6 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | AAA, GC-MS | [80,81] | |
| HCN, 0.1 | r.t./4–12 m/NH4OH (pH 9.2) | Soluble oligomers | Acid and neutral | GC-MS | [82] | |
| HCN, 0.1 | r.t./18 m/NH4OH (pH 9.2) | Soluble oligomers | Acid, Basic and neutral | GC-MS | [79] | |
| HCN, 0.002–0.1 | r.t./UV radiation (Hg lamp)/pH (8–9) | Soluble oligomers | Acid | AAA | [69] | |
| HCN(G) | r.t./UV radiation (Hg lamp) | Solid | Acid | AAA | [83] | |
| HCN, 0.004–0.1 | r.t./γ-radiation (60Co source) | Raw mixture | Acid | PC, AAA, GC-MS | [84] | |
| HCN, 0.1 | r.t./γ-radiation (60Co source)/pH 6 | Raw mixture | Acid | AAA, GC-MS | [70] | |
| HCN, 0.1 | r.t./γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [70] | |
| NaCN, 0.1 | r.t./γ-radiation (60Co source)/pH 11.3 | Solution | Acid | AAA, GC-MS | [70] | |
| HCN, 0.1 | r.t.−40 °C/γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [71] | |
| HCN, 1.5 | Refrigerator/4 d/NH3 | Black Solid | Acid | AAA | [51] | |
| HCN, 0.1 | −20/2 m/NH3/pH 9.2 | Solution | Acid | HPLC | [67] | |
| HCN, 0.1 | −20/25 y/NH3/pH 9.2 | Solution | Acid | HPLC | [67] | |
| HCN, 0.1 | −78/25 y/NH3/pH 9.2 | Solution | Acid | HPLC | [67] | |
| Glycinamide | HCN, 1.5 | 70/5 d/NH4OH | Soluble oligomers | Non hydrolysis | PC | [78] |
| Aminomalonic acid | NaCN, 1 | 38/3–30 d/NH4Cl | Black solid | Acid | GC-MS | [46] |
| Alanine | HCN, 1.5 | 100/1 d/- | Raw mixture | Acid | GC-MS | [75] |
| NH4CN, 1 | 90/4 h/- | Brown precipitate | Acid | AAA | [76] | |
| HCN, 1 | 90/4 h/NH4OH (pH 9.58) | Black solid | Acid | 2D-PC | [72] | |
| HCN, 1.5 | 90/18 h/NH3 | Soluble oligomers | Acid | 2D-PC, AAA | [77] | |
| HCN, 1.5 | 70/5 d/NH4OH | Soluble oligomers | Non hydrolysis- | PC | [78] | |
| HCN, 2.2 | 70/25 d/NH4OH | Soluble oligomers | Non hydrolysis | PC | [63] | |
| HCN(L) | r.t./4 w/anhydrous NH3 | Dark solid residue | Acid | AAA | [51] | |
| NaCN, 1 | r.t./3 m/pH 9.2 (HCl) | Soluble oligomers | Acid, basic and neutral | GC-MS | [79] | |
| HCN, 0.1 | r.t./1–6 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | AAA, GC-MS | [80,81] | |
| HCN, 0.1 | r.t./4–12 m/NH4OH (pH 9.2) | Soluble oligomers | Acid and neutral | GC-MS | [82] | |
| HCN, 0.1 | r.t./18 m/NH4OH (pH 9.2) | Soluble oligomers | Acid, Basic and neutral | GC-MS | [79] | |
| HCN, 0.002–0.1 | r.t./UV radiation (Hg lamp)/pH (8–9) | Solution | Acid | AAA | [69] | |
| HCN(G) | r.t./UV radiation (Hg lamp) | Solid | Acid | AAA | [83] | |
| HCN, 0.004–0.1 | r.t./γ-radiation (60Co source) | Raw mixture | Acid | PC, AAA, GC-MS | [84] | |
| HCN, 0.1 | r.t./γ-radiation (60Co source)/pH 6 | Raw mixture | Acid | AAA, GC-MS | [70] | |
| HCN, 0.1 | r.t./γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [70] | |
| NaCN, 0.1 | r.t./γ-radiation (60Co source)/pH 11.3 | Solution | Acid | AAA, GC-MS | [70] | |
| HCN, 0.1 | r.t.−40 °C/γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [71] | |
| HCN, 1.5 | Refrigerator/4 d/NH3 | Black Solid | Acid | AAA | [51] | |
| HCN, 0.1 | −20/2 m/NH3/pH 9.2 | Solution | Acid | HPLC | [67] | |
| HCN, 0.1 | −20/25 y/NH3/pH 9.2 | Solution | Acid | HPLC | [67] | |
| HCN, 0.1 | −78/25 y/NH3/pH 9.2 | Solution | Acid | HPLC | [67] | |
| β-alanine | HCN, 1.5 | 90/18 h/NH3 | Soluble oligomers | Acid | 2D-PC, AAA | [77] |
| HCN, 0.1 | r.t./4–12 m/NH4OH (pH 9.2) | Soluble oligomers | Acid and neutral | GC-MS | [82] | |
| HCN, 0.1 | r.t./γ-radiation (60Co source)/pH 6 | Raw mixture | Acid | AAA, GC-MS | [70] | |
| HCN, 0.1 | r.t./γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [70] | |
| NaCN, 0.1 | r.t./γ-radiation (60Co source)/pH 11.3 | Solution | Acid | AAA, GC-MS | [70] | |
| HCN, 0.1 | r.t.−40 °C/γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [71] | |
| Sarcosine | HCN, 0.1 | r.t./γ-radiation (60Co source)/pH 6 | Raw mixture | Acid | AAA, GC-MS | [70] |
| HCN, 0.1 | r.t./γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [70] | |
| NaCN, 0.1 | r.t./γ-radiation (60Co source)/pH 11.3 | Solution | Acid | AAA, GC-MS | [70] | |
| HCN, 0.1 | r.t.−40 °C/γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [71] | |
| Serine | NH4CN, 1 | 90/4 h/- | Brown precipitate | Acid | AAA | [76] |
| HCN, 1 | 90/4 h/NH4OH (pH 9.58) | Black solid | Acid | 2D-PC | [72] | |
| HCN, 1.5 | 90/18 h/NH3 | Soluble oligomers | Acid | 2D-PC, AAA | [77] | |
| HCN(L) | r.t./.4 w/anhydrous NH3 | Dark solid residue | Acid | AAA | [51] | |
| HCN, 0.1 | r.t./1–6 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | AAA | [80,81] | |
| HCN, 0.002–0.1 | r.t./UV radiation (Hg lamp)/pH (8–9) | Solution | Acid | AAA | [69] | |
| HCN(G) | r.t./UV radiation (Hg lamp) | Solid | Acid | AAA | [83] | |
| HCN, 0.004 | r.t./γ-radiation (60Co source) | Raw mixture | Acid | PC, AAA, GC-MS | [84] | |
| HCN, 0.1 | r.t./γ-radiation (60Co source)/pH 6 | Raw mixture | Acid | AAA | [70] | |
| HCN, 0.1 | r.t./γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA | [70] | |
| NaCN, 0.1 | r.t./γ-radiation (60Co source)/pH 11.3 | Solution | Acid | AAA | [70] | |
| HCN, 0.1 | r.t.−40 °C/γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [71] | |
| HCN, 1.5 | Refrigerator/4 d/NH3 | Black Solid | Acid | AAA | [51] | |
| 2,3-Aminopropioinic acid | HCN, 1.5 | 90/18 h/NH3 | Soluble oligomers | Acid | 2D-PC, AAA | [77] |
| Aspartic acid | HCN, 0.2 | 100/1 d/- | Raw mixture | Acid | GC-MS | [75] |
| NH4CN, 1 | 90/4 h/- | Brown precipitate | Acid | AAA | [76] | |
| HCN, 1 | 90/4 h/NH4OH (pH 9.58) | Black solid | Acid | 2D-PC | [72] | |
| HCN, 1.5 | 90/18 h/NH3 | Soluble oligomers | Acid | 2D-PC, AAA | [77] | |
| HCN, 1.5 | 70/5 d/NH4OH | Soluble oligomers | Non hydrolysis | PC | [78] | |
| HCN, 2.2 | 70/25 d/NH4OH | Soluble oligomers | Non hydrolysis | PC | [63] | |
| NaCN, 1 | 38/3–30 d/NH4Cl | Black solid | Acid | GC-MS | [46] | |
| HCN(L) | r.t./4 w/anhydrous NH3 | Dark solid residue | Acid | AAA | [51] | |
| NaCN, 1 | r.t./3 m/pH 9.2 (HCl) | Soluble oligomers | Acid, basic and neutral | GC-MS | [79] | |
| HCN, 0.1 | r.t./1–6 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | AAA, GC-MS | [80,81] | |
| HCN, 0.1 | r.t./4–12 m/NH4OH (pH 9.2) | Soluble oligomers | Acid and neutral | GC-MS | [82] | |
| HCN, 0.1 | r.t./18 m/NH4OH (pH 9.2) | Soluble oligomers | Acid, Basic and neutral | GC-MS | [79] | |
| HCN, 0.002–0.1 | r.t./UV radiation (Hg lamp)/pH (8–9) | Solution | Acid | AAA | [69] | |
| HCN(G) | r.t./UV radiation (Hg lamp) | Solid | Acid | AAA | [83] | |
| HCN, 0.004–0.1 | r.t./γ-radiation (60Co source) | Raw mixture | Acid | PC, AAA, GC-MS | [84] | |
| HCN, 0.1 | r.t./γ-radiation (60Co source)/pH 6 | Raw mixture | Acid | AAA, GC-MS | [70] | |
| HCN, 0.1 | r.t./γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [70] | |
| NaCN, 0.1 | r.t./γ-radiation (60Co source)/pH 11.3 | Solution | Acid | AAA, GC-MS | [70] | |
| HCN, 0.1 | r.t.−40 °C/γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [71] | |
| HCN, 1.5 | Refrigerator/4 d/NH3 | Black Solid | Acid | AAA | [51] | |
| HCN, 0.1 | −20/2 m/NH3/pH 9.2 | Solution | Acid | HPLC | [67] | |
| HCN, 0.1 | −20/25 y/NH3/pH 9.2 | Solution | Acid | HPLC | [67] | |
| HCN, 0.1 | −78/25 y/NH3/pH 9.2 | Solution | Acid | HPLC | [67] | |
| Diaminosuccinic acid | NaCN, 1 | r.t./3 m/pH 9.2 (HCl) | Soluble oligomers | Acid, basic and neutral | GC-MS | [79] |
| HCN, 0.1 | r.t./1–6 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | AAA, GC-MS | [80,81] | |
| HCN, 0.1 | r.t./4–12 m/NH4OH (pH 9.2) | Soluble oligomers | Acid and neutral | GC-MS | [82] | |
| HCN, 0.1 | r.t./18 m/NH4OH (pH 9.2) | Soluble oligomers | Acid, Basic and neutral | GC-MS | [79] | |
| 2-aminoisobutyric acid | HCN, 0.2 | 100/1 d/- | Raw mixture | Acid | GC-MS | [75] |
| NaCN, 1 | r.t./3 m/pH 9.2 (HCl) | Soluble oligomers | Acid, basic and neutral | GC-MS | [79] | |
| HCN, 0.1 | r.t./1–6 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | GC-MS | [80,81] | |
| HCN, 0.1 | r.t./4–12 m/NH4OH (pH 9.2) | Soluble oligomers | Acid and neutral | GC-MS | [82] | |
| HCN, 0.1 | r.t./18 m/NH4OH (pH 9.2) | Soluble oligomers | Acid, Basic and neutral | GC-MS | [79] | |
| 2-amino-n-butyric acid | HCN, 1.5 | 90/18 h/NH3 | Soluble oligomers | Acid | 2D-PC, AAA | [77] |
| HCN, 0.1 | r.t./γ-radiation (60Co source)/pH 6 | Raw mixture | Acid | AAA, GC-MS | [70] | |
| HCN, 0.1 | r.t./γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [70] | |
| NaCN, 0.1 | r.t./γ-radiation (60Co source)/pH 11.3 | Solution | Acid | AAA, GC-MS | [70] | |
| HCN, 0.1 | r.t.−40 °C/γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [71] | |
| 4-amino-n-butyric acid | NH4CN, 1 | 90/4 h/- | Brown precipitate | Acid | AAA | [76] |
| HCN, 1 | 90/4 h/NH4OH (pH 9.58) | Black solid | Acid | 2D-PC | [72] | |
| Threonine | NH4CN, 1 | 90/4 h/- | Brown precipitate | Acid | AAA | [76] |
| HCN, 1 | 90/4 h/NH4OH (pH 9.58) | Black solid | Acid | 2D-PC | [72] | |
| HCN, 1.5 | 90/18 h/NH3 | Soluble oligomers | Acid | 2D-PC, AAA | [77] | |
| HCN(L) | r.t./4 w/anhydrous NH3 | Dark solid residue | Acid | AAA | [51] | |
| HCN, 0.1 | r.t./1–6 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | AAA | [80,81] | |
| HCN(G) | r.t./UV radiation (Hg lamp) | Solid | Acid | AAA | [83] | |
| HCN, 0.004–0.1 | r.t./γ-radiation (60Co source) | Raw mixture | Acid | PC, AAA, GC-MS | [84] | |
| HCN, 0.1 | r.t.−40 °C/γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [71] | |
| HCN, 1.5 | Refrigerator/4 d/NH3 | Black Solid | Acid | AAA | [51] | |
| Glutamic acid | HCN, 0.2 | 100/1 d/- | Raw mixture | Acid | GC-MS | [75] |
| NH4CN, 1 | 90/4 h/- | Brown precipitate | Acid | AAA | [76] | |
| HCN, 1.5 | 90/18 h/NH3 | Soluble oligomers | Acid | 2D-PC, AAA | [77] | |
| HCN(L) | r.t./4 w/anhydrous NH3 | Dark solid residue | Acid | AAA | [51] | |
| NaCN, 1 | r.t./3 m/pH 9.2 (HCl) | Soluble oligomers | Acid, basic and neutral | GC-MS | [79] | |
| HCN, 0.1 | r.t./1–6 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | AAA, GC-MS | [80,81] | |
| HCN, 0.1 | r.t./18 m/NH4OH (pH 9.2) | Soluble oligomers | Acid, Basic and neutral | GC-MS | [79] | |
| HCN(G) | r.t./UV radiation (Hg lamp) | Solid | Acid | AAA | [83] | |
| HCN, 0.004–0.1 | r.t./γ-radiation (60Co source) | Raw mixture | Acid | PC, AAA, GC-MS | [84] | |
| HCN, 0.1 | r.t./γ-radiation (60Co source)/pH 6 | Raw mixture | Acid | AAA, GC-MS | [70] | |
| HCN, 0.1 | r.t./γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [70] | |
| NaCN, 0.1 | r.t./γ-radiation (60Co source)/pH 11.3 | Solution | Acid | AAA, GC-MS | [70] | |
| HCN, 0.1 | r.t.−40 °C/γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [71] | |
| HCN, 1.5 | Refrigerator/4 d/NH3 | Black Solid | Acid | AAA | [51] | |
| 2-methyl aspartic acid | HCN, 0.1 | r.t./γ-radiation (60Co source)/pH 6 | Raw mixture | Acid | AAA, GC-MS | [70] |
| HCN, 0.1 | r.t./γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [70] | |
| NaCN, 0.1 | r.t./γ-radiation (60Co source)/pH 11.3 | Solution | Acid | AAA, GC-MS | [70] | |
| HCN, 0.1 | r.t.−40 °C/γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [71] | |
| Ornithine | NH4CN, 1 | 90/4 h/- | Brown precipitate | Acid | AAA | [76] |
| HCN, 1 | 90/4 h/NH4OH (pH 9.58) | Black solid | Acid | 2D-PC | [72] | |
| Histidine | NH4CN, 1 | 90/4 h/- | Brown precipitate | Acid | AAA | [76] |
| HCN, 1 | 90/4 h/NH4OH (pH 9.58) | Black solid | Acid | 2D-PC | [72] | |
| HCN(L) | r.t./4 w/anhydrous NH3 | Dark solid residue | Acid | AAA | [51] | |
| HCN, 0.1 | r.t./1–6 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | AAA | [80,81] | |
| HCN(G) | r.t./UV radiation (Hg lamp) | Solid | Acid | AAA | [83] | |
| HCN, 0.1 | r.t.−40 °C/γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [71] | |
| HCN, 1.5 | Refrigerator/4 d/NH3 | Black Solid | Acid | AAA | [51] | |
| Valine | HCN(L) | r.t./4 w/anhydrous NH3 | Dark solid residue | Acid | AAA | [51] |
| HCN, 0.1 | r.t./1–6 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | AAA | [80,81] | |
| HCN(G) | r.t./UV radiation (Hg lamp) | Solid | Acid | AAA | [83] | |
| HCN, 0.1 | r.t./γ-radiation (60Co source) | Raw mixture | Acid | PC, AAA, GC-MS | [84] | |
| HCN, 1.5 | Refrigerator/4 d/NH3 | Black Solid | Acid | AAA | [51] | |
| Isoleucine | HCN, 1.5 | 90/18 h/NH3 | Soluble oligomers | Acid | 2D-PC, AAA | [77] |
| HCN (L) | r.t./4 w/anhydrous NH3 | Dark solid residue | Acid | AAA | [51] | |
| NaCN, 1 | r.t./3 m/pH 9.2 (HCl) | Soluble oligomers | Acid, basic and neutral | GC-MS | [79] | |
| HCN, 0.1 | r.t./1–6 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | AAA, GC-MS | [80,81] | |
| HCN, 0.1 | r.t./18 m/NH4OH (pH 9.2) | Soluble oligomers | Acid, Basic and neutral | GC-MS | [79] | |
| HCN(G) | r.t./UV radiation (Hg lamp) | Solid | Acid | AAA | [83] | |
| HCN, 1.5 | Refrigerator/4 d/NH3 | Black Solid | Acid | AAA | [51] | |
| Leucine | HCN, 1.5 | 90/18 h/NH3 | Soluble oligomers | Acid | 2D-PC, AAA | [77] |
| HCN (L) | r.t./4 w/anhydrous NH3 | Dark solid residue | Acid | AAA | [51] | |
| HCN, 0.1 | r.t./1–6 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | AAA | [80,81] | |
| HCN(G) | r.t./UV radiation (Hg lamp) | Solid | Acid | AAA | [83] | |
| HCN, 1.5 | Refrigerator/4 d/NH3 | Black Solid | Acid | AAA | [51] | |
| Citrulline | HCN, 0.1 | r.t./1–6 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | AAA | [80,81] |
| HCN, 0.1 | r.t.−40 °C/γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [71] | |
| Lysine | NH4CN, 1 | 90/4 h/- | Brown precipitate | Acid | AAA | [76] |
| HCN, 1 | 90/4 h/NH4OH (pH 9.58) | Black solid | Acid | 2D-PC | [72] | |
| HCN(L) | r.t./4 w/anhydrous NH3 | Dark solid residue | Acid | AAA | [51] | |
| HCN, 0.1 | r.t./1–6 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | AAA | [80,81] | |
| HCN(G) | r.t./UV radiation (Hg lamp) | Solid | Acid | AAA | [83] | |
| HCN, 0.1 | r.t.−40 °C/γ-radiation (60Co source)/NH3/pH 9 | Solution | Acid | AAA, GC-MS | [71] | |
| HCN, 1.5 | Refrigerator/4 d/NH3 | Black Solid | Acid | AAA | [51] | |
| Arginine | HCN(L) | r.t./4 w/anhydrous NH3 | Dark solid residue | Acid | AAA | [51] |
| HCN, 1.5 | Refrigerator/4 d/NH3 | Black Solid | Acid | AAA | [51] |
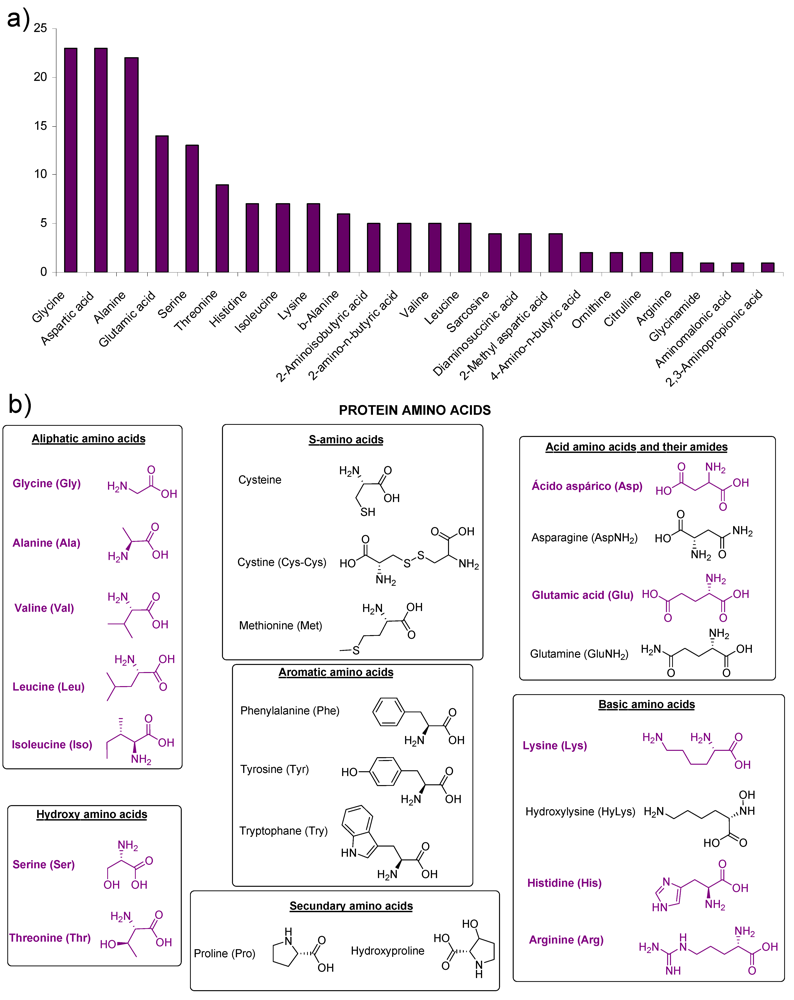
Hydantoins

3. Nucleic-Acid Bases

3.1. Purines
| Compound | Starting material, c (M) | T (ºC)/t/Catalyser | Final product analysed | Hydrolysis | Yield (%) | Method of identification | Reference |
|---|---|---|---|---|---|---|---|
| Xhanthine | HCN, 0.1 | −78/27 y/NH3 (pH 9.2) | Solution + black solid | Acid | 0.022 | HPLC-UV, GC-MS | [66] |
| Neutral | 0.022 | ||||||
| Hypoxanthine | HCN, 1.5 | 90/18 h/NH3 | Solution + black solid | Acid | ~1 μmol/L | PC, UV-spectrum, PE | [77] |
| NaCN, 1 | 38/3–30 d/NH4Cl | Black solid | Acid | D | GC-MS | [46] | |
| HCN, 0.1 | −78/27 y/NH3 (pH 9.2) | Solution + black solid | Acid | 0.0041 | HPLC-UV, GC-MS | [66] | |
| Neutral | 0.0058 | ||||||
| Adenine | HCN, (s) | 90/24 h/NH4OH (1.5 M) | Solution | Acid | D | PC, UV-spectrum | [6] |
| HCN, 1.5 | 90/18 h/NH3 | Solution + black solid | Acid | ~1 μmol/L | PC, UV-spectrum, PE | [77] | |
| HCN, 9.9 | 90/8 d/NH4OH | Solution | Non hydrolysis | 60 mg/L | 2D-PC, UV-spectrum | [78] | |
| HCN, 10 | 80/24 h/NH4OH | Solution | Acid | 0.027 | HPLC-UV | [99] | |
| HCN, 1.5 | 70/2 d/NH4OH (3 M) | Solution | Non hydrolysis | D | 2D-PC, UV-spectrum | [78] | |
| HCN, 11.1 | 70/5 d/NH4OH (12.8 M) | Solution | Non hydrolysis | 110 mg/L | 2D-PC, UV-spectrum | [78] | |
| Acid | 700 mg/L | ||||||
| NaCN, 1 | 38/3–30 d/NH4Cl | Black solid | Acid | D | GC-MS | [46] | |
| HCN, 14.6 | r.t./26 h/NH4OH (7 M) | Solution | Non hydrolysis | D | 2D-PC, UV-spectrum | [78] | |
| HCN, 8.25 | r.t./26 h/NH4OH (13 M) | Solution | Non hydrolysis | D | 2D-PC, UV-spectrum | [78] | |
| HCN, 0.1 | r.t./1 w/NH4OH (pH 9.2) | Solution + black solid | Acid | 0.000013 | HPLC-UV, GC-MS | [66] | |
| HCN, 0.1 | r.t./4 w/NH4OH (pH 9.2) | Solution + black solid | Acid | 0.00031 | HPLC-UV, GC-MS | [66] | |
| HCN, 0.1 | r.t./8 w/NH4OH (pH 9.2) | Solution + black solid | Acid | 0.00062 | HPLC-UV, GC-MS | [66] | |
| HCN, 0.1 | r.t./4–12 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | 0.003–0.004 | GC-MS | [82] | |
| NaCN, 1 | r.t./1 y/pH 9.2 (HCl) | Soluble oligomers | Acid | D | TLC, UV-spectrum | [82] | |
| NaCN, 2 (+HCOH) | r.t./9 m/pH 9.2 (HCl) | Non hydrolysis | 3 μmol/L | HPLC, UV spectrum, MS | [96] | ||
| Neutral (2) | 0.06 | ||||||
| HCN, 0.2 | r.t.−40 °C/γ-radiation/pH 6 | Solution | Acid (3) | t | HPLC, GC-MS | [97] | |
| HCN, 0.01 (+glyconitrile) | −2/60 d/NH4OH (pH 9.2) | Solution | Acid (2) | 0.02 | HPLC-UV | [93] | |
| HCN, 0.01 | −2/98 d/NH4OH (pH 9.2) | Solution | Acid (2) | 0.004 | HPLC-UV | [93] | |
| HCN, 0.1 | −20/2 m/NH3 (pH 9.2) | Solution | Acid | 0.005 | HPLC-UV, ESI-MS | [67] | |
| HCN, 0.001 | −20/3 m/NH4OH (pH 9.2) | Solution + black solid | Acid | 0.0042 | HPLC-UV, GC-MS | [66] | |
| HCN, 0.01 | −20/3 m/NH4OH (pH 9.2) | Solution + black solid | Acid | 0.01 | HPLC-UV, GC-MS | [66] | |
| HCN | −20/3 m/NH4OH (pH 9.2) | Solution + black solid | Acid | 0.0094 | HPLC-UV, GC-MS | [66] | |
| HCN, 0.1 | −20/25 y/NH3 (pH 9.2) | Solution | Acid | 0.038 | HPLC-UV | [99] | |
| HCN, 0.1 | −20/25 y/NH3 (pH 9.2) | Solution | Acid | 0.035 | HPLC-UV, ESI-MS | [67] | |
| NaCN, 0.1 | −30/2 m/NH4Cl | Solution | Acid | 0.0004 | HPLC-UV | [99] | |
| HCN, 0.1 | −78/25 y/NH3 (pH 9.2) | Solution | Acid | 0.04 | HPLC-UV, ESI-MS | [67] | |
| HCN, 0.1 | −78/27 y/NH3 (pH 9.2) | Solution + black solid | Acid | 0.029 | HPLC-UV, GC-MS | [66] | |
| Neutral | 0.012 | ||||||
| Non hydrolysis | 0.00016 | ||||||
| Guanine | HCN, 10 | 80/24 h/NH4OH | Solution | Acid | 0.0007 | HPLC-UV | [99] |
| NaCN, 1 | 38/3–30 d/NH4Cl | Black solid | Acid | D | GC-MS | [46] | |
| HCN, 0.1 | −20/25 y/NH3 (pH 9.2) | Solution | Acid | 0.0035 | HPLC-UV | [99] | |
| HCN, 0.1 | −20/25 y/NH3 (pH 9.2) | Solution | Acid | 0.0004 | HPLC-UV, ESI-MS | [67] | |
| NaCN, 0.1 | −30/2 m/NH4Cl | Solution | Acid | 0.000014 | HPLC-UV | [99] | |
| HCN, 0.1 | −78/27 y/NH3 (pH 9.2) | Solution + black solid | Acid | 0.0067 | HPLC-UV, GC-MS | [66] | |
| Neutral | 0.0033 | ||||||
| Non hydrolysis | 0.00011 | ||||||
| 2,6-diaminopurine | HCN, 0.1 | −78/27 y/NH3 (pH 9.2) | Solution + black solid | Neutral | 0.0091 | HPLC-UV, GC-MS | [66] |
| 8-hydroxymethyladenine | NaCN, 2 (+HCOH) | r.t./9 m/pH 9.2 (HCl) | Non hydrolysis | 47 μmol/L | HPLC, UV spectrum, MS | [96] | |
| Neutral (2)> | 0.06 |
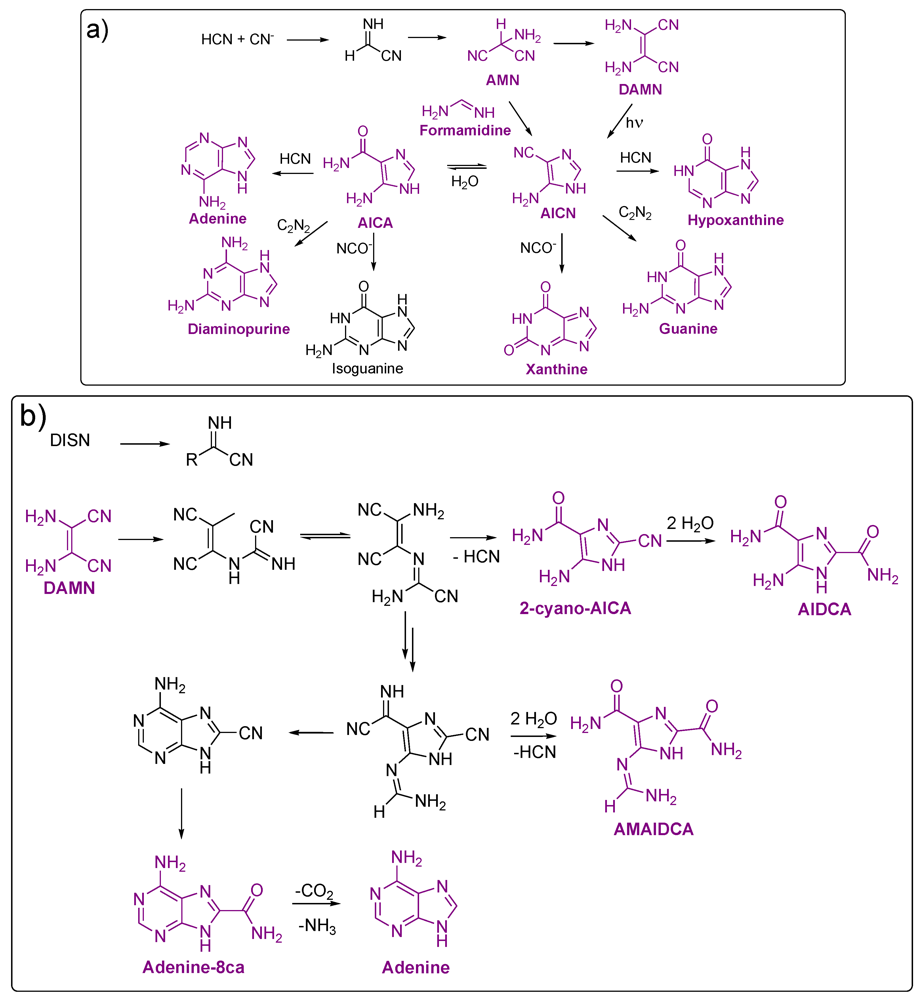
3.2. Pyrimidines
| Compound | Starting material, c(M) | T (ºC)/t/Catalyst | Final product analysed | Hydrolysis | Yield (%) | Method of identification | Reference |
|---|---|---|---|---|---|---|---|
| 4,5-Dihydroxypyrimidine | HCN, 0.1 | r.t./4–12 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | 0.62 | TLC, UV spectrum | [102] |
| HCN, 0.1 | r.t./4–12 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | 0.7–0.9 | GC-MS | [82] | |
| NaCN, 1 | r.t./1 y/pH 9.2 (HCl) | Soluble oligomers | Acid | D | TLC, UV spectrum | [82] | |
| HCN, 0.1 | −78/27 y/NH3 (pH 9.2) | Solution + black solid | Acid | 0.65 | HPLC-UV, GC-MS | [66] | |
| Uracil | NaCN, 1 | 38/3–30 d/NH4Cl | Black solid | Acid | D | GC-MS | [46] |
| HCN, 1 | r.t./6 m/NH4OH (pH 9.2) | Solution | Acid (2) | 0.001 | HPLC-UV | [103] | |
| HCN, 0.1 | r.t./6 m/NH4OH (pH 9.2) | Solution | Acid (2) | 0.005 | HPLC-UV | [103] | |
| NaCN, 1 | r.t./6 m/pH 9.2 (HCl) | Solution | Acid (2) | 0.001 | HPLC-UV | [103] | |
| HCN, 0.1 | −78/27 y/NH3 (pH 9.2) | Solution + black solid | Acid | 0.00026 | HPLC-UV, GC-MS | [66] | |
| Neutral | 0.0017 | ||||||
| HCN, 0.2 | r.t.−40 °C/γ-rad (60Co)/pH 6 | Solution | Acid (3) | t | HPLC, GC-MS | [97] | |
| 5-Hydroxyuracil | NaCN, 1 | 38/3–30 d/NH4Cl | Black solid | Acid | D | GC-MS | [46] |
| HCN, 0.1 | r.t./4–12 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | 0.002–0.004 | TLC, UV spectrum | [102] | |
| HCN, 0.1 | r.t./4–12 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | 0.003 | GC-MS | [82] | |
| HCN, 0.1 | −78/27 y/NH3 (pH 9.2) | Solution + black solid | Acid | 0.0015 | HPLC-UV, GC-MS | [66] | |
| Cytosine | HCN, 0.2 | r.t.−40 °C/γ-rad (60Co)/pH 6 | Solution | Acid (3) | t | HPLC, GC-MS | [97] |
| 5-Aminouracil | NaCN, 1 | 38/3–30 d/NH4Cl | Black solid | Acid | D | GC-MS | [46] |
| HCN, 0.1 | −78/27 y/NH3 (pH 9.2) | Solution + black solid | Acid | 0.0058 | HPLC-UV, GC-MS | [66] | |
| Neutral | 0.0038 | ||||||
| Orotic acid | NaCN, 1 | 38/3–30 d/NH4Cl | Black solid | Acid | D | GC-MS | [46] |
| HCN, 0.1 | r.t./4–12 m/NH4OH (pH 9.2) | Soluble oligomers | Neutral (2) | 0.009 | TLC, UV spectrum | [102] | |
| HCN, 0.1 | r.t./4–12 m/NH4OH (pH 9.2) | Soluble oligomers | Acid | 0.009 | GC-MS | [82] | |
| HCN, 0.1 | −78/27 y/NH3 (pH 9.2) | Solution + black solid | Acid | 0.0025 | HPLC-UV, GC-MS | [66] | |
| Neutral | 0.1 | ||||||
| 5-Aminoorotic acid | HCN, 0.1 | −78/27 y/NH3 (pH 9.2) | Solution + black solid | Neutral | 0.019 | HPLC-UV, GC-MS | [66] |
| HCN, 0.1 | −78/27 y/NH3 (pH 9.2) | Solution + black solid | Non hydrolysis | 0.00028 | HPLC-UV, GC-MS | [66] | |
| Thymine | HCN, 0.2 | r.t.−40 °C/γ-rad (60Co)/pH 6 | Solution | Acid (3) | t | HPLC, GC-MS | [97] |
| 1,2,5,6-Tetrahydropyrimidine | NaCN, 1 | 38/3–30 d/NH4Cl | Black solid | Acid | D | GC-MS | [46] |
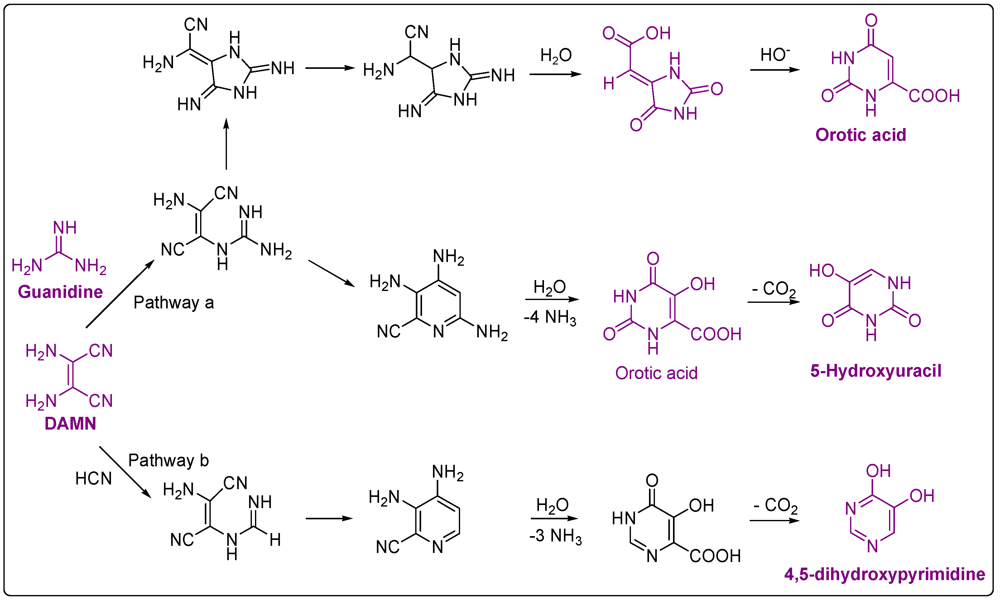
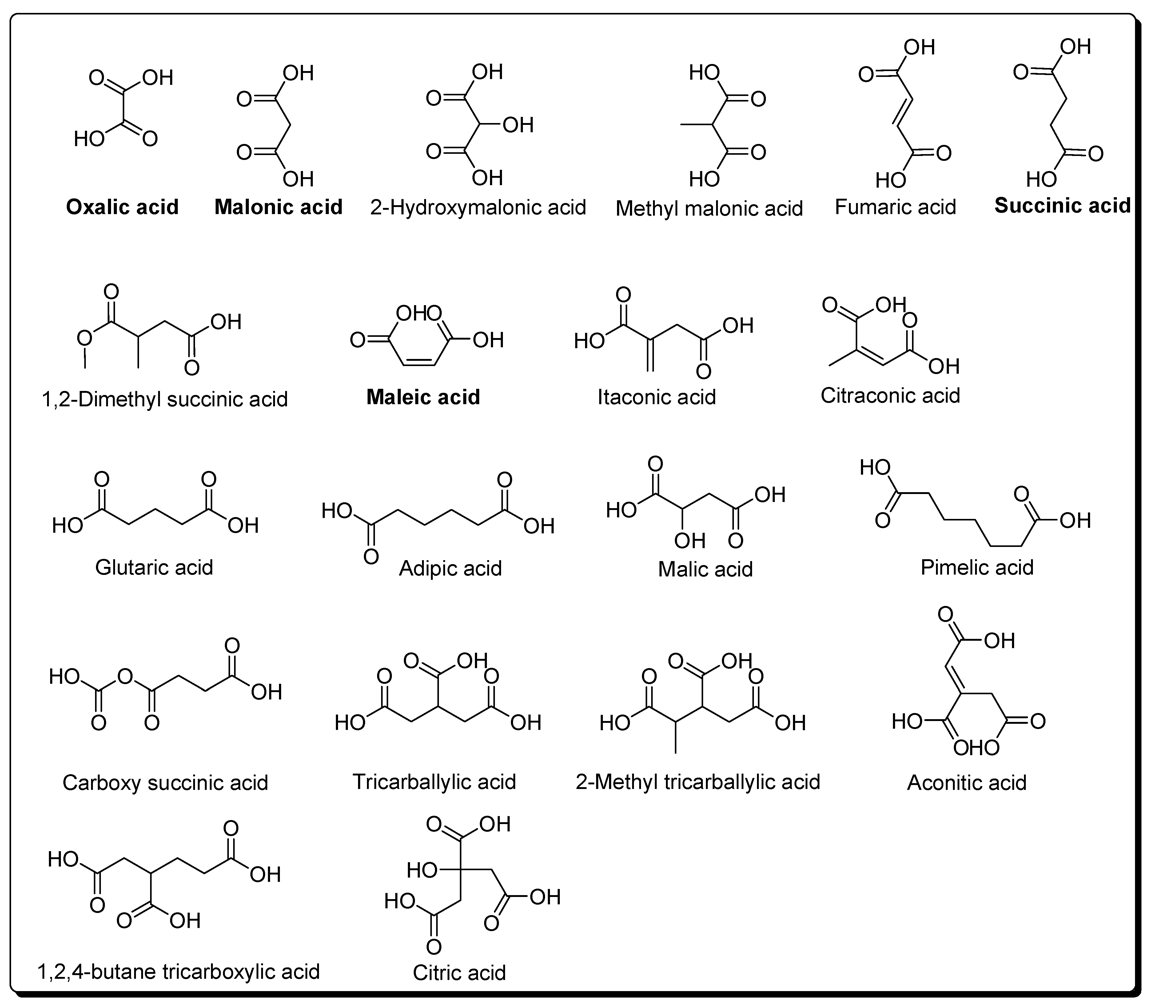
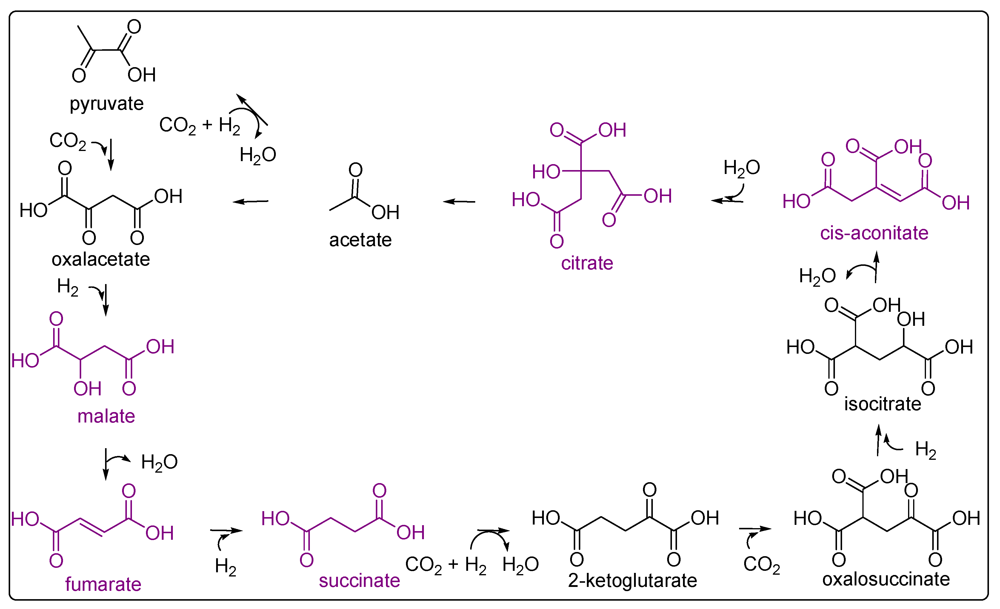
4. Carboxylic Acids
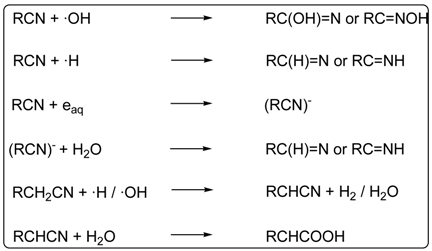
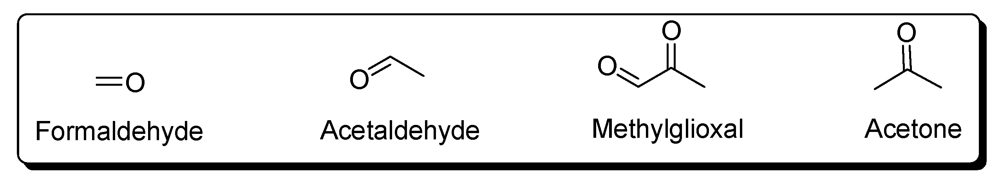
5. Carbonyl Compounds
6. Pteridines
7. Others

8. Summary

Acknowledgments
References
- Miller, S.L.; Cleaves, H.J. Systems Biology: Geomics; Rigoutsos, I., Stephanopoulos, G., Eds.; Oxford Univessity Press: New York, NY, USA, 2007; Volume 1, pp. 3–56. [Google Scholar]
- Bauer, H. Die ersten organisch-chemischen Synthesen. Naturwissenschaften 1980, 67, 1–6. [Google Scholar] [CrossRef]
- Proust, J.L. Contributions on Cyanides. Ann. Chim. Phys. 1806, 60, 233. [Google Scholar]
- Wipperman, R. Ueber Tricyanwasserstoff, eine der blausaure polymere verbindung. Ber. Deustchen Chem. Ges. 1874, 7, 767–772. [Google Scholar] [CrossRef]
- Pflüger, E. Beitragë zur Lehre von der Respiration. I. Ueber die physiologische Verbrennung in den lebendigen organismen. Arch. Ges. Physiol. 1875, 10, 641–644. [Google Scholar] [CrossRef]
- Oró, J. Synthesis of adenine from ammonium cyanide. Biochem. Biophys. Res. Commun. 1960, 2, 407–412. [Google Scholar] [CrossRef]
- Gao, Y.; Solomon, P.M. HCN survey of normal spiral, infrared-luminous and ultraluminous galaxies. Astrophys. J. Suppl. Ser. 2004, 152, 63–80. [Google Scholar] [CrossRef]
- Buhl, D. Chemical constituents of interstellar clouds. Nature 1971, 234, 332–334. [Google Scholar] [CrossRef]
- Boger, G.I.; Sternberg, A. CN and HCN in dense interstellar clouds. Astrophys. J. 2005, 632, 302–315. [Google Scholar] [CrossRef]
- Greaves, J.S.; Church, S.E. Photodissociation and the CN: HCN ratio: Observations of a ‘Third Bar’ in OMC1. Mon. Not. R. Astron. Soc. 1996, 283, 1179–1183. [Google Scholar] [CrossRef]
- Simon, R.; Stutzki, J.; Sternberg, A.; Winnewisser, G. Chemical stratification in the orion bar region: CN and CS submillimeter observations. Astron. Astrophys. 1997, 327, L9–L12. [Google Scholar]
- Young Owl, R.C.; Meixner, M.M.; Wolfire, M.; Tielens, A.G.G.M.; Tauber, J. HCN and HCO+ images of the orion bar photodissociation region. Astrophys. J. 2000, 540, 886–906. [Google Scholar] [CrossRef]
- Savage, C.; Apponi, A.J.; Ziurys, L.M.; Wyckoff, S. Galactic 12C/13C Ratios from millimeter-wave observations of interstellar CN. Astrophys. J. 2002, 578, 211–223. [Google Scholar] [CrossRef]
- Schneider, N.; Simon, R.; Kramer, C.; Kraemer, K.; Stutzki, J.; Mookerjea, B. A multiwavelength study of the S 106 region-II. Characteristics of the photon dominated region. Astron. Astrophys. 2003, 406, 915–935. [Google Scholar] [CrossRef]
- Fuente, A.; Martin-Pintado, J.; Cernicharo, J.; Bachiller, R. A chemical study of the photodissociation region NGC 7023. Astron. Astrophys. 1993, 276, 473–488. [Google Scholar]
- Fuente, A.; Martin-Pintado, J.; Gaume, R. High-density CN filaments in NGC 2023. Astrophys. J. 1995, 442, L33. [Google Scholar] [CrossRef]
- Jansen, D.J.; van Dishoeck, E.F.; Black, J.H.; Spaans, M.; Sosin, C. Physical and chemical structure of the IC 63 nebula. II. Chemical models. Astron. Astrophys. 1995, 302, 223–242. [Google Scholar]
- Fuente, A.; Rodriguez-Franc, A.; Garcia-Burillo, S.; Martin-Pintado, J.; Black, J.H. Observational study of reactive ions and radicals in PDRs. Astron. Astrophys. 2003, 406, 899–913. [Google Scholar] [CrossRef]
- Bachiller, R.; Forveille, T.; Huggins, P.J.; Cox, P. The chemical evolution of planetary nebulae. Astron. Astrophys. 1997, 324, 1123–1134. [Google Scholar]
- McKay, C.P.; Borucki, W.J. Organic synthesis in experimental impact shocks. Science 1997, 276, 390–392. [Google Scholar] [CrossRef]
- Magee-Sauer, K.; Mumma, M.J.; DiSanti, M.A.; Russo, N.J.; Retting, T.W. Infrared spectroscopy of the ν3 band of hydrogen cyanide in Comet C/1995 O1 hale-bopp. Icarus 1999, 142, 498–508. [Google Scholar] [CrossRef]
- Wootten, A.; Lichten, S.M.; Sahai, R.; Wannier, P.G. CN abundance variations in the shell of IRC + 10216. Astrophys. J. 1982, 257, 151–160. [Google Scholar] [CrossRef]
- Truong-Bach, A.; Nguyen-Q-Rieu, A.; Omont, O.H.; Johansson, L.E.B. The circumstellar shell of IRC + 10216-Photo-chemistry of C2H and CN. Astron. Astrophys. 1987, 176, 285–293. [Google Scholar]
- Bachiller, R.; Fuente, A.; Bujarrabal, V.; Colomar, F.; Loup, C.; Omont, A.; de Jong, T. A survey of CN in circumstellar envelopes. Astron. Astrophys. 1997, 319, 235–243. [Google Scholar]
- Lindqvist, M.; Schçier, F.L.; Lucas, R.; Olofsson, H. Molecular envelopes around carbon stars Interferometric observations and models of HCN and CN emission. Astron. Astrophys. 2000, 361, 1036–1057. [Google Scholar]
- Van Zadelhoff, G.-J.; Aikawa, Y.; Hogerheijde, M.R.; van Dishoeck, E.F. Axi-symmetric models of ultraviolet radiative transfer with applications to circumstellar disk chemistry. Astron. Astrophys. 2003, 397, 789–802. [Google Scholar] [CrossRef]
- Thi, W.-F.; van Zadelhoff, G.-J.; van Dishoeck, E.F. Organic molecules in protoplanetary disks around TTauri and HerbigAe stars. Astron. Astrophys. 2004, 425, 955–972. [Google Scholar] [CrossRef]
- Rank, M.D.; Townes, C.H.; Welch, W.J. Interstellar molecules and dense clouds. Science 1971, 174, 1083–1101. [Google Scholar] [CrossRef]
- Donn, B. Comets: Chemistry and chemical evolution. J. Mol. Evol. 1982, 18, 157–160. [Google Scholar] [CrossRef]
- Fray, N.; Bénilan, Y.; Cottin, H.; Gazeau, M.-C.; Crovisier, J. The origin of the CN radical in comets: A review from observations and models. Planet Space Sci. 2005, 53, 1243–1262. [Google Scholar] [CrossRef]
- Matthews, C.N. Hydrogen cyanide polymers from the impact of comet P/Shoemaker-Levy 9 on Jupiter. Adv. Space Res. 1997, 19, 1087–1091. [Google Scholar] [CrossRef]
- Pizzarello, S. Hydrogen cyanide in Murchinson meteorite. Astrophys. J. Lett. 2012, 754, L27. [Google Scholar] [CrossRef]
- Hanel, R.A.; Conrath, B.; Flaser, F.M.; Kunde, V.; Maguire, W.; Pearl, J.; Pirraglia, J.; Samuelson, R.; Herath, L.; Allison, M.; et al. Infrared observations of the saturnian system from voyager 1. Science 1981, 212, 192–200. [Google Scholar]
- Tokunaga, A.T.; Beck, S.C.; Geballe, T.R.; Lacey, J.H.; Serabyn, E. The detection of HCN on Jupiter. Icarus 1981, 48, 283–289. [Google Scholar] [CrossRef]
- Owen, T. The atmosphere of Titan. J. Mol. Evol. 1982, 18, 150–156. [Google Scholar] [CrossRef]
- Irvine, W.M. The composition of interstellar molecular clouds. Space Sci. Rev. 1999, 90, 203–218. [Google Scholar] [CrossRef]
- Hidayat, T.; Marten, A.; Bézard, B.; Gautier, D.; Owen, T.; Matthwes, H.E.; Paubert, G. Millimeter and submillimeter heterodyne observations of Titan: Retrieval of the vertical profile of HCN and the12C/13C ratio. Icarus 1997, 126, 170–182. [Google Scholar] [CrossRef]
- Hards, V. Volcanic contributions to the global carbon cycle. Sustainable and renewable energy. Br. Geol. Surv. Occas. Publ. 2005, 10, 16–17. [Google Scholar]
- Ferris, J. Marine hydrothermal systems and the origin of life: Chemical markers of prebiotic chemistry in hydrothermal systems. Orig. Life Evol. Biosph. 1992, 22, 109–134. [Google Scholar] [CrossRef]
- Miller, S.L. The mechanism of synthesis of amino acids by electric discharges. Biochim. Biophys. Acta 1957, 23, 480–489. [Google Scholar] [CrossRef]
- Greenberg, J.M.; Mendoza-Gomez, C.X.; Pirronello, V. The Chemistry of life’s origins. NATO ASI Ser. Ser. C Math. Phys. Sci. 1993, 416, 259–299. [Google Scholar]
- Bar Nun, A.; Bar-Nun, N.; Bauer, S.H.; Sagan, C. Shock synthesis of amino acids in simulated primitive environments. Science 1970, 168, 470–473. [Google Scholar]
- Ferris, J.P.; Chen, C.T. Photosynthesis of organic compounds in the atmosphere of Jupiter. Nature 1975, 258, 587–588. [Google Scholar] [CrossRef]
- Stribling, R.; Miller, S.L. Electric discharge synthesis of HCN in simulated Jovian Atmospheres. Orig. Life 1987, 17, 261. [Google Scholar] [CrossRef]
- Cataldo, F.; Lilla, E.; Ursini, O.; Angelini, G. TGA-FT-IR study of pirólisis of poly(hydrogen cyanide) synthesized from termal decomposition of formamide. Implications in cometary emissions. J. Anal. Appl. Pyrolysis 2010, 87, 34–44. [Google Scholar] [CrossRef]
- Ruiz-Bermejo, M.; de la Fuente, J.L.; Rogero, C.; Menor-Salván, C.; Osuna-Esteba, S.; Martín-Gago, J.A. New insights into the characterization of ‘Insoluble Black HCN polymers’. Chem. Biodiver. 2012, 9, 25–40. [Google Scholar] [CrossRef]
- Mamajanov, I.; Herzfeld, J. HCN polymers characterized by SSNMR: Solid state reaction of crystalline tetramer (diaminomaleonitrile). J. Chem. Phys. 2009, 130, 134504. [Google Scholar] [CrossRef]
- Mamajanov, I.; Herzfeld, J. HCN polymers characterized by solid state NMR: Chains and sheets formed in the neat liquid. J. Chem. Phys. 2009, 130, 134503. [Google Scholar] [CrossRef]
- Umemoto, K.; Takahashi, M.; Yokota, K. Studies on structure of HCN oligomers. Orig. Life 1987, 17, 283–293. [Google Scholar] [CrossRef]
- Ferris, J.P.; Edelson, E.H.; Auyeung, J.M.; Joshi, P.C. Structural studies on HCN oligomers. J. Mol. Evol. 1981, 17, 69–77. [Google Scholar] [CrossRef]
- Matthews, C.N.; Moser, R.E. Peptide synthesis from hydrogen cyanide and water. Nature 1967, 215, 1230–1234. [Google Scholar] [CrossRef]
- Völker, T. Polymere blausäure. Angew. Chem. 1960, 72, 379–384. [Google Scholar] [CrossRef]
- Pernot, P.; Carrasco, N.; Thissen, R.; Schmitz-Afonso, I. Tholinomics-chemical analysis of nitrogen-rich polymers. Anal. Chem. 2010, 82, 1371–1380. [Google Scholar] [CrossRef]
- Vuitton, V.; Bonnet, J.Y.; Frisari, M.; Thissen, R.; Quirico, E.; Dutuit, O.; Schmitt, B.; Le Roy, L.; Fray, N.; Cottin, H.; et al. Very high resolution mass spectrometry of HCN polymers and tholins. Faraday Discuss. 2010, 147, 495–508. [Google Scholar] [CrossRef]
- Hanczyc, M.M. Metabolism and motility in prebiotic structures. Phylosophi. Trans. B 2011, 366, 2885–2895. [Google Scholar] [CrossRef] [Green Version]
- De la Fuente, J.L.; Ruiz-Bermejo, M.; Menor-Salván, C.; Osuna-Esteban, S. Thermal characterization of HCN polymers by TG-MS, TG, DTA and DSC methods. Polym. Degrad. Stab. 2011, 96, 943–948. [Google Scholar] [CrossRef]
- He, C.; Lin, G.; Upton, K.T.; Imanaka, H.; Smith, M.A. Structural investigation of HCN polymer isotopomers by solution-state multidimensional NMR. J. Phys. Chem. A 2012, 116, 4751–4759. [Google Scholar] [CrossRef]
- Ferris, J.P.; Hagan, W.J. HCN and Chemicals evolution: The possible role of cyano compounds in prebiotic síntesis. Tetrahedron 1984, 40, 1093–1120. [Google Scholar] [CrossRef]
- Schwartz, A.W.; Voet, A.B. Recent progress in the prebiotic chemistry of HCN. Orig. Life 1984, 14, 91–98. [Google Scholar] [CrossRef]
- Saladino, R.; Crestini, C.; Costanzo, G.; DiMauro, E. Advances in the prebiotic synthesis of nucleic acids bases: Implications for the origin of Life. Curr. Org. Chem. 2004, 8, 1425–1443. [Google Scholar] [CrossRef]
- Matthews, C.N.; Minard, R.D. Hydrogen cyanide polymers, comets and the origin of life. Faraday Discuss. 2006, 133, 393–401. [Google Scholar] [CrossRef]
- Miller, S.L. A production of amino acids under possible primitive Earth conditions. Science 1953, 117, 528–529. [Google Scholar]
- Oró, J.; Kamat, S.S. Amino-acids synthesis from hydrogen cyanide under possible pritive Earth conditions. Nature 1961, 190, 442–443. [Google Scholar] [CrossRef]
- Sanchez, R.A.; Ferris, J.P.; Orgel, L.E. Studies in prebiotic synthesis II, Synthesis of purine precursors anda mino acids from aqueous hydrogen cyanide. J. Mol. Biol. 1967, 30, 223–252. [Google Scholar]
- Stribling, R.; Miller, S.L. Energy yields for hydrogen cyanide and formaldehyde syntheses: The HCN and amino acid concentrations in the primitive ocean. Orig. Life 1987, 17, 261–273. [Google Scholar] [CrossRef]
- Miyakawa, S.; Cleaves, H.J.; Miller, S.L. The cold origin of life: B. Implications based on pyrimidines and purines produced from frozen ammonium cyanide solutions. Orig. Life Evol. Biosph. 2002, 32, 209–218. [Google Scholar] [CrossRef]
- Levy, M.; Miller, S.L.; Brinton, K.; Bada, J.L. Prebiotic synthesis of adenine and amino acids under Europa-like conditions. Icarus 2000, 145, 609–123. [Google Scholar] [CrossRef]
- Garzón, L.; Garzón, M.L. Radioactivity as a significant energy source in prebiotic synthesis. Orig. Life Evol. Biosph. 2001, 31, 3–13. [Google Scholar] [CrossRef]
- Abelson, P.H. Chemical events on the primitive Earth. Proc. Natl. Acad. Sci.USA 1966, 55, 1365–1372. [Google Scholar] [CrossRef]
- Draganic, Z.; Draganic, I. Evidence for amino acids in hydrolysates of compounds formed by ionizing radiations. Orig. Life 1977, 8, 371–376. [Google Scholar] [CrossRef]
- Draganic, Z.D.; Niketic, V.; Jovanovic, S.; Draganic, I.G. The radiolysis of aqueous ammonium cyanide: Compounds of interest to chemical evolution studies. J. Mol. Evol. 1980, 15, 239–260. [Google Scholar] [CrossRef]
- Labadie, M.; Jensen, R.; Neuzil, E. Recherches sur I’évolution pré-biologique III. Les acides azulmiques noirs formés à partir du cyanure d’ammonium. Biochim. Biophys. Acta 1968, 165, 525–533. [Google Scholar] [CrossRef]
- Moser, R.E.; Claggett, A.R.; Matthews, C.N. Peptide formation from aminomalononitrile (HCN trimer). Tetrahedron Lett. 1968, 9, 1605–1068. [Google Scholar] [CrossRef]
- Moser, R.E.; Claggett, A.R.; Matthews, C.N. Peptide formation from diaminomaleonitrile (HCN tetramer). Tetrahedron Lett. 1968, 9, 1599–1603. [Google Scholar] [CrossRef]
- Yuasa, S.; Flory, D.; Basile, B.; Oró, J. On the abiotic formation of amino acids I. HCN as precursors of amino acids detected in extracts of lunar samples II. Formation of HCN and amino acids from simulated mixtures of gases released from lunar samples. J. Mol. Evol. 1984, 20, 52–58. [Google Scholar] [CrossRef]
- Labadie, M.; Jensen, R.; Neuzil, E. Recherches sur l’eévolution pré-biologique. I. Composition en amino-acides des microsphérules obtenues a partir du cyanure d’ammonium. Bull. Soc. Chim. Biol. 1967, 49, 673–682. [Google Scholar]
- Lowe, C.U.; Ress, A.; Markham, F.R.S. Synthesis of complex organic compounds from simple precursors: Formation of amino-acids, amino-acid polymer, fatty acids and purines from ammonium cyanide. Nature 1963, 19, 219–222. [Google Scholar] [CrossRef]
- Oró, J.; Kimball, P. Synthesis of purines under possible primitive Earth conditions. I. Adenine from hydrogen cyanide. Arch. Biochem. Biophys. 1962, 94, 217–227. [Google Scholar]
- Ferris, J.P.; Wos, J.D.; Nooner, D.W.; Oró, J. Chemical evolution XXI. The amino acids released on hydrolysis f HCN oligomers. J. Mol. Evol.o 1974, 3, 225–231. [Google Scholar] [CrossRef]
- Ferris, J.P.; Donner, D.B.; Lobo, A.P. Possible role of hydrogen cyanide in chemical evolution: Investigation of prposed direct synthesis of peptides from hydrogen cyanide. J. Mol. Biol. 1973, 74, 499–510. [Google Scholar] [CrossRef]
- Ferris, J.P.; Wos, J.D.; Ryan, T.J.; Lobo, A.P.; Donner, D.B. Biomolecules from HCN. Orig. Life 1974, 5, 153–157. [Google Scholar] [CrossRef]
- Ferris, J.P.; Joshi, P.C.; Edelson, E.H.; Lawless, J.G. HCN: A plausible source of purines, pyrimidines and amino acids on the primitive Earth. J. Mol. Evol. 1978, 11, 293–311. [Google Scholar] [CrossRef]
- Mizutani, H.; Mikuni, H.; Takahasi, M.; Noda, H. Study on the photochemical reaction of HCN and its polymers products relating to primary chemical evolution. Orig. Life 1975, 6, 513–525. [Google Scholar] [CrossRef]
- Sweeney, M.A.; Toste, A.P.; Ponnamperuma, C. Formation of amino acids by Cobalt-60 irradiation of hydrogen cyanide solutions. Orig. Life 1976, 7, 187–189. [Google Scholar] [CrossRef]
- Ferris, J.P.; Wos, J.D.; Lobo, A.P. Chemical Evolution. XXII. The hydantoins released on hydrolysis of HCN oligomers. J. Mol. Evol. 1974, 3, 311–316. [Google Scholar] [CrossRef]
- Ferris, J.P.; Donner, D.B.; Lobo, A.P. Possible role of hydrogen cyanide in Chemicals evolution: The oligomerization and condensation of hydrogen cyanide. J. Mol. Biol. 1973, 74, 511–518. [Google Scholar] [CrossRef]
- Miller, S.L.; Orgel, L.E. The Origins of Life on the Earth; Prentice-Hall: New York, NY, USA, 1974. [Google Scholar]
- Gilbert, W. The origin of life: The RNA world. Nature 1986, 319, 618. [Google Scholar] [CrossRef]
- Sheppard, T.P.; Ordoukhanian, P.; Joyce, G.F. A DNA enzyme with N-glycosylase activity. Proc. Natl. Acad. Sci. USA 2000, 97, 7802–7807. [Google Scholar] [CrossRef]
- Santoro, S.W.; Joyce, G.F.; Sakthivel, K.; Gramatikova, S.; Barbas, C.F., III. RNA cleavage by a DNA enzymewith extended chemical functionality. J. Am. Chem. Soc. 2000, 122, 2433–2439. [Google Scholar] [CrossRef]
- Pullman, B. Electronic Factors in Biochemical Evolution. In Exobiology; Ponnamperuma, C., Ed.; North Holland Publishing Company: Amsterdam, The Netherlands and London, UK, 1972; p. 140. [Google Scholar]
- Sanchez, R.A.; Ferris, J.P.; Orgel, L.E. Conditions for purine synthesis: Did prebiotic synthesis occur at low temperatures? Science 1966, 153, 72–73. [Google Scholar]
- Schwartz, A.W.; Joosten, H.; Voet, A.B. Prebiotic adenine synthesis via HCN oligomerization in ice. BioSystems 1982, 15, 191–193. [Google Scholar] [CrossRef]
- Schwartz, A.W.; Goverde, M. Acceleration of HCN oligomerization by formaldehyde and related compounds: Implications for prebiotic synthesis. J. Mol. Evol. 1982, 18, 351–353. [Google Scholar] [CrossRef]
- Voet, A.B.; Schwartz, A.W. Prebiotic adenine synthesis from HCN—Evidence for a newly discorvered major pathway. Bioorg. Chem. 1983, 12, 8–17. [Google Scholar] [CrossRef]
- Schwartz, A.W.; Bakker, C.G. Was adenine the first purine? Science 1989, 245, 1102–1104. [Google Scholar]
- Negrón-Mendoza, A.; Draganic, Z.D. Search for heterocyclic radiolytic products in aqueous Solutions of cyanide. Adv. Space Res. 1984, 4, 121–124. [Google Scholar] [CrossRef]
- Borquez, E.; Cleaves, H.J.; Lazcano, A.; Miller, S.L. An investigation of prebiotic purine synthesis from the hydrolysis of HCN polymers. Orig. Life Evol. Biosph. 2005, 35, 79–90. [Google Scholar] [CrossRef]
- Levy, M.; Miller, S.L.; Oró, J. Production of guanine from NH4CN polymerizations. J. Mol. Evol. 1999, 49, 165–168. [Google Scholar] [CrossRef]
- Ferris, J.P.; Orgel, L.E. An inusual photochemical rearrangement in the shyntesis of adenine from hydrogen cyanide. J. Am. Chem. Soc. 1966, 88, 1074–1074. [Google Scholar] [CrossRef]
- Ferris, J.P.; Orgel, L.E. Studies in Prebiotic Synthesis. I. Aminomalononitrile and 4-Amino-5-cyanoimidazole. J. Am. Chem. Soc. 1966, 88, 3829–3831. [Google Scholar] [CrossRef]
- Ferris, J.P.; Joshi, P.C.; Lawless, J.G. Chemical evolution XXIX. Pyrimidines from hydrogen cyanide. BioSystems 1977, 9, 81–86. [Google Scholar] [CrossRef]
- Voet, A.B.; Schwartz, A.W. Uracil synthesis via HCN oligomerization. Orig. Life 1982, 12, 45–49. [Google Scholar] [CrossRef]
- Negrón-Mendoza, A.; Draganic, Z.D.; Navarro-Gonzalez, R.; Draganic, I.G. Aldehydes, ketones, and carboxylic acids formed radiolytically in aqueous Solutions of cyanides and simple nitriles. Rad. Res. 1983, 95, 248–261. [Google Scholar] [CrossRef]
- Negrón-Mendoza, A.; Ramos-Bernal, S.; Cruz, E.; Juárez, J.M. Radiolysis of HCN in heterogeneous phase. Rad. Phys. Chem. 2001, 61, 771–772. [Google Scholar] [CrossRef]
- Eschenmoser, A. On a hypothetical generational relationship between HCN and constituents of the reductive citric acid cycle. Chem. Biodiver. 2007, 4, 554–573. [Google Scholar] [CrossRef]
- Smith, E.; Morowitz, H.J. Universality in intermediary metabolism. Proc. Natl. Acad. Sci. USA 2004, 101, 13168–13173. [Google Scholar] [CrossRef]
- Eschenmoser, A.; Loewenthal, E. Chemistry of potentially prebiological natural products. Chem. Soc. Rev. 1992, 21, 1–16. [Google Scholar] [CrossRef]
- Eschenmosr, A. Vitamin BI2: Experiments concerning the origin of its molecular structure. Angew. Chem. Int. Ed. 1988, 27, 5–39. [Google Scholar] [CrossRef]
- Visser, C.M. Evolutionary roots of catalysis by nicotinamida and flavins in C-H oxidoreductases and in photosynthesis. Orig. Life 1982, 12, 165–179. [Google Scholar] [CrossRef]
- Schimpl, A.; Lemmon, R.M.; Calvin, M. Cyanamide formation under primitive Earth conditions. Science 1965, 147, 149–150. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Ruiz-Bermejo, M.; Zorzano, M.-P.; Osuna-Esteban, S. Simple Organics and Biomonomers Identified in HCN Polymers: An Overview. Life 2013, 3, 421-448. https://doi.org/10.3390/life3030421
Ruiz-Bermejo M, Zorzano M-P, Osuna-Esteban S. Simple Organics and Biomonomers Identified in HCN Polymers: An Overview. Life. 2013; 3(3):421-448. https://doi.org/10.3390/life3030421
Chicago/Turabian StyleRuiz-Bermejo, Marta, María-Paz Zorzano, and Susana Osuna-Esteban. 2013. "Simple Organics and Biomonomers Identified in HCN Polymers: An Overview" Life 3, no. 3: 421-448. https://doi.org/10.3390/life3030421






