Properties of Halococcus salifodinae, an Isolate from Permian Rock Salt Deposits, Compared with Halococci from Surface Waters
Abstract
:1. Introduction
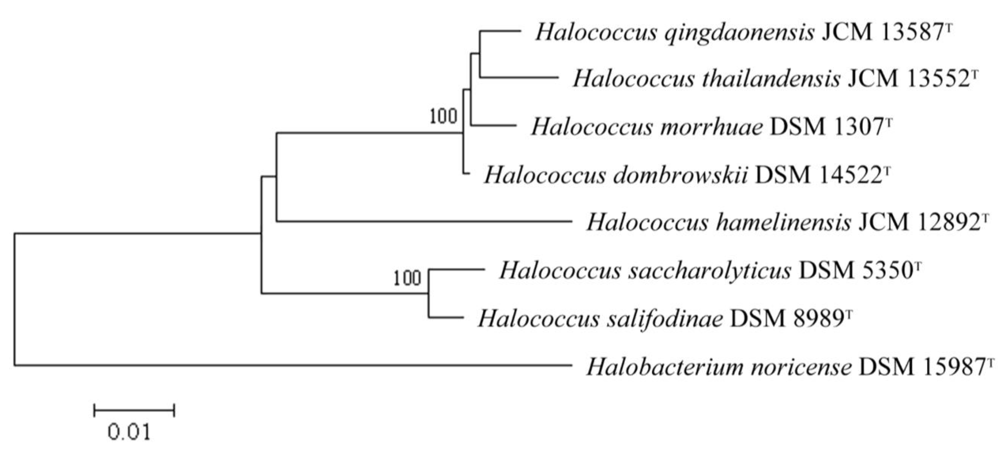
2. Results and Discussion
2.1. General Description of halococci [26]
2.2. Properties of Isolates from Permo-Triassic Salt Sediments and Surface Waters
| Characteristic* | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Oxidase | + | + | + | + | + | + | - | + | + | - | + |
| Catalase | + | + | + | + | + | + | + | + | + | + | + |
| Alkaline phosphatase | (+) | + | + | + | + | ||||||
| Esterase (C4) | + | + | + | + | + | ||||||
| Lipase esterase (C8) | + | + | + | + | + | ||||||
| Lipase (C14) | - | - | - | - | - | ||||||
| Leucine arylamidase | - | - | - | - | - | + | |||||
| Trypsin | - | - | - | - | - | - | |||||
| Acid phosphatase | - | - | + | + | + | + | - | ||||
| Cystine arylamidase | - | - | - | - | - | + | - | ||||
| Nitrate reduction | + | + | + | - | + | + | + | + | + | ||
| Gelatin liquefaction | + | - | - | v | - | - | + | - | v | ||
| Hydrolysis of starch | + | + | + | - | + | - | v | ||||
| casein | - | - | - | - | - | ||||||
| Tween 20 | - | + | + | - | |||||||
| Tween 80 | + | + | + | - | - | + | |||||
| Sensitivity to anti-biotics: Tetracycline | + | + | + | + | + | - | - | - | - | - | - |
| " : Chloramphenicol | + | + | + | + | + | - | - | - | + | - | |
| " : Novobiocin | + | + | + | + | + | - | + | + | + | + |
2.3. Cell Wall of Hcc. salifodinae

| Cell wall constituentsa | Hcc. morrhuae CCM 859b | Hcc. salifodinae BIpT DMS 8989T |
| Glucose | 440 | 470 |
| Mannose | 350 | 220 |
| Galactose | 270 | 360 |
| Ribose | n.d. | 60 |
| Glucosamine | 380 | 180 |
| Galactosamine | 200 | 80 |
| Glucuronic acid | 470 | 60 |
| Galacturonic acid | 200 | 20 |
| Gulosaminuronic acid | 110 | n.d. |
| Acetate | 620 | 660 |
| Sulfate | 1470 | 1580 |
| Phosphate | 120 | 130 |
| Glycine | 100 | 7 |
| Lysine | n.d. | 1 |
2.4. Production of polyhydroxyalkanotes (PHA)
2.5. BLAST Search of Genes in the Genome of Hcc. hamelinensis.
2.5.1. Polyhydroxyalkanaote Synthase (phaC)
2.5.2. Subunit A (atpA) of the Archaeal ATP Synthase
2.6. Are Permo-Triassic Isolates Suitable for Evolutionary Studies?
2.6.1. What Type of Results Can be Expected?
2.6.2. Which Strains Should be Used for Comparative Studies?
3. Experimental Section
4. Conclusions
Acknowledgments
References
- Denner, E.B.M.; McGenity, T.J.; Busse, H.-J.; Wanner, G.; Grant, W.D.; Stan-Lotter, H. Halococcus salifodinae sp. nov., an archaeal isolate from an Austrian salt mine. Int. J. System. Bacteriol. 1994, 44, 774–780. [Google Scholar] [CrossRef]
- Radax, C.; Gruber, G.; Stan-Lotter, H. Novel haloarchaeal 16S rRNA gene sequences from Alpine Permo-Triassic rock salt. Extremophiles 2001, 5, 221–228. [Google Scholar] [CrossRef]
- Stan-Lotter, H.; McGenity, T.J.; Legat, A.; Denner, E.B.M.; Glaser, K.; Stetter, K.O.; Wanner, G. Very similar strains of Halococcus salifodinae are found in geographically separated Permo-Triassic salt deposits. Microbiol. 1999, 145, 3565–3574. [Google Scholar]
- Schoop, G. Halococcus litoralis, ein obligat halophiler Farbstoffbildner. Dtsch Tierärztl Wochens 1935, 43, 817–820. [Google Scholar]
- Oren, A.; Arahal, D.R.; Ventosa, A. Emended descriptions of genera of the family Halobacteriaceae. Int. J. Syst. Evol. Microbiol. 2009, 59, 637–642. [Google Scholar] [CrossRef]
- Kocur, M.; Hodgkiss, W. Taxonomic status of the genus Halococcus Schoop. Int. J. Syst. Bacteriol. 1973, 23, 151–156. [Google Scholar] [CrossRef]
- Montero, C.G.; Ventosa, A.; Rodriguez-Valera, F.; Kates, M.; Moldoveanu, N.; Ruiz-Berraquero, F. Halococcus saccharolyticus sp. nov., a new species of extremely halophilic non-alkaliphilic cocci. Syst. Appl. Microbiol. 1989, 12, 167–171. [Google Scholar] [CrossRef]
- Stan-Lotter, H.; Pfaffenhuemer, M.; Legat, A.; Busse, H.-J.; Radax, C.; Gruber, C. Halococcus dombrowskii sp. nov., an archaeal isolate from a Permian alpine salt deposit. Int. J. Syst. Evol. Microbiol. 2002, 52, 1807–1814. [Google Scholar] [CrossRef]
- Goh, F.; Leuko, S.; Allen, M.A.; Bowman, J.P.; Kamekura, M.; Neilan, B.A.; Burns, B.P. Halococcus hamelinensis sp. nov., a novel halophilic archaeon isolated from stromatolites in Shark Bay, Australia. Int. J. Syst. Evol. Microbiol. 2006, 56, 1323–1329. [Google Scholar] [CrossRef]
- Wang, Q.-F.; Li, W.; Yang, H.; Liu, Y.-L.; Cao, H.-H.; Dornmayr-Pfaffenhuemer, M.; Stan-Lotter, H.; Guo, G.-Q. Halococcus qingdaonensis sp. nov., a halophilic archaeon isolated from a crude sea-salt sample. Int. J. Syst. Evol. Microbiol. 2007, 57, 600–604. [Google Scholar] [CrossRef]
- Namwong, S.; Tanasupawat, S.; Visessanguan, W.; Kudo, T.; Itoh, T. Halococcus thailandensis sp. nov., from fish sauce in Thailand. Int. J. Syst. Evol. Microbiol. 2007, 57, 2199–2203. [Google Scholar] [CrossRef]
- Wright, A.-D.G. Phylogenetic relationships within the order Halobacteriales inferred from16S rRNA gene sequences. Int. J. Syst. Evol. Microbiol. 2006, 56, 1223–1227. [Google Scholar] [CrossRef]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar]
- Kumar, S. Molecular clocks: four decades of evolution. Nat. Rev. Genetics 2005, 6, 654–662. [Google Scholar] [CrossRef]
- Dennis, P.P.; Shimmin, L.C. Evolutionary divergence and salinity-mediated selection in halophilic Archaea. Microb. Mol. Biol. Rev. 1997, 61, 90–104. [Google Scholar]
- Gogarten, J.P.; Townsend, J.P. Horizontal gene transfer, genome innovation and evolution. Nat. Rev. Microbiol. 2005, 3, 679–687. [Google Scholar] [CrossRef]
- McGenity, T.J.; Gemmell, R.T.; Grant, W.D.; Stan-Lotter, H. Origins of halophilic microorganisms in ancient salt deposits. Environ. Microbiol. 2000, 2, 243–250. [Google Scholar] [CrossRef]
- Schubert, B.A.; Lowenstein, T.K.; Timofeeff, M.N.; Parker, M.A. Halophilic Archaea cultured from ancient halite, Death Valley, California. Environ. Microbiol. 2010, 12, 440–454. [Google Scholar] [CrossRef]
- Fendrihan, S.; Dornmayr-Pfaffenhuemer, M.; Gerbl, F.W.; Holzinger, A.; Grösbacher, M.; Briza, P.; Erler, A.; Gruber, C.; Plätzer, K.; Stan-Lotter, H. Spherical particles of halophilic Archaea correlate with exposure to low water activity - implications for microbial survival in fluid inclusions of ancient halite. Geobiology 2012, 10, 424–433. [Google Scholar] [CrossRef]
- Schubert, B.A.; Lowenstein, T.K.; Timofeeff, M.N. Microscopic identification of prokaryotes in modern and ancient halite, Saline Valley and Death Valley, California. Astrobiology 2009, 9, 467–482. [Google Scholar] [CrossRef]
- Norton, C.F.; Grant, W.D. Survival of halobacteria within fluid inclusions in salt crystals. J. Gen. Microbiol. 1988, 134, 1365–1373. [Google Scholar]
- Mormile, M.R.; Biesen, M.A.; Gutierrez, M.C.; Ventosa, A.; Pavlovich, J.B.; Onstott, T.C.; Fredrickson, J.K. Isolation of Halobacterium salinarum retrieved directly from halite brine inclusions. Environ. Microbiol. 2003, 5, 1094–1102. [Google Scholar] [CrossRef]
- Fendrihan, S.; Legat, A.; Pfaffenhuemer, M.; Gruber, C.; Weidler, G.; Gerbl, F.; Stan-Lotter, H. Extremely halophilic archaea and the issue of long-term microbial survival. Rev. Environ. Sci. Biotech. 2006, 5, 203–218. [Google Scholar] [CrossRef]
- Gramain, A.; Chong Díaz, G.C.; Demergasso, C.; Lowenstein, T.K.; McGenity, T.J. Archaeal diversity along a subterranean salt core from the Salar Grande (Chile). Environ. Microbiol. 2011, 13, 2105–2121. [Google Scholar] [CrossRef]
- Burns, B.P.; Gudhka, R.K.; Neilan, B.A. Genome sequence of the halophilic archaeon Halococcus hamelinensis. J. Bacteriol. 2012, 194, 2100–2101. [Google Scholar] [CrossRef]
- Grant, W.D.; Genus, I.V. Halococcus Schoop 1935a, 817AL. In Bergey's Manual of Systematic Bacteriology, 2nd ed.; Boone, D.R., Castenholz, R.W., Garrity, G.M., Eds.; Springer-Verlag: New York, NY, USA, 2001; Volume 1, pp. 311–314. [Google Scholar]
- Stackebrandt, E.; Goebel, B.M. Taxonomic note: a place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species definition in bacteriology. Int. J. Syst. Bacteriol. 1994, 44, 846–849. [Google Scholar] [CrossRef]
- Wayne, L.G.; Brenner, D.J.; Colwell, R.R.; Grimont, P.A.D.; Kandler, O.; Krichevsky, M.I.; Moore, L.H.; Moore, W.E.C.; Murray, R.G.E.; Stackebrandt, E.; Starr, M.P.; Trüper, H.G. International Committee on Systematic Bacteriology. Report of the ad hoc committee on reconciliation of approaches to bacterial systematics. Int. J. Syst. Bacteriol. 1987, 37, 463–464. [Google Scholar] [CrossRef]
- Zharkov, M.A. History of Paleozoic Salt Accumulation; Springer Verlag: Berlin, Germany, 1981. [Google Scholar]
- Stan-Lotter, H.; Radax, C.; McGenity, T.J.; Legat, A.; Pfaffenhuemer, M.; Wieland, H.; Gruber, C.; Denner, E.B.M. From intraterrestrials to extraterrestrials - viable haloarchaea in ancient salt deposits. In Halophilic Microorganisms; Ventosa, A., Ed.; Springer Verlag: New York, NY, USA, 2004; pp. 89–102. [Google Scholar]
- Kocur, M.; Smid, B.; Martinec, T. The fine structure of extreme halophilic cocci. Microbios 1972, 5, 101–107. [Google Scholar]
- Sumper, M.; Berg, E.; Mengele, R.; Strobel, I. Primary structure and glycosylation of the S-Layer protein of Haloferax volcanii. J. Bacteriol. 1990, 172, 7111–7118. [Google Scholar]
- Schleifer, K.H.; Steber, J.; Mayer, H. Chemical composition and structure of the cell wall of Halococcus morrhuae. Zbl. Bakt. Hyg. 1. Abt. Orig. 1982, 3, 171–178. [Google Scholar]
- Niemetz, R.; Kärcher, U.; Kandler, O.; Tindall, B.J.; König, H. The cell wall polymer of the extremely halophilic archaeon Natronococcus occultus. Eur. J. Biochem. 1997, 249, 905–911. [Google Scholar]
- Kandler, O.; König, H. Cell envelopes of Archaebacteria. In The Bacteria vol. VII; Woese, C.R., Wolfe, R.S., Eds.; Academic Press: New York, NY, USA, 1985; pp. 413–457. [Google Scholar]
- König, H.; Rachel, R.; Claus, H. Proteinaceous surface layers of Archaea: ultrastructure and biochemistry. In Archaea. Molecular Cell Biology; Cavicchioli, R., Ed.; ASM Press: Washington, DC, USA, 2007; pp. 315–340. [Google Scholar]
- Claus, H.; Akca, E.; Debaerdemaeker, T.; Evrard, C.; Deqlercq, J.P.; Harris, J.R.; Schlott, B.; König, H. Molecular organization of selected prokaryoticS-layer proteins. Can. J. Microbiol. 2005, 51, 731–743. [Google Scholar] [CrossRef]
- Kärcher, U.; Schröder, H.; Haslinger, E.; Allmeier, G.; Schreiner, R.; Wieland, F.; Haselbeck, A.; König, H. Primary structure of the heterosaccharide of the surface glycoprotein of Methanothermus fervidus. J. Biol. Chem. 1993, 268, 26821–26826. [Google Scholar]
- Brown, A.D.; Cho, K.Y. The walls of extremely halophilic cocci: Gram-positive bacteria lacking muramic acid. J. Gen. Microbiol. 1970, 62, 267–270. [Google Scholar]
- Reistad, R. Cell wall of an extremely halophilic coccus. Investigation of ninhydrin-positive compounds. Arch. Microbiol. 1972, 82, 24–30. [Google Scholar]
- Steber, J.; Schleifer, K.H. N-Glycyl-glucosamine, a novel constituent in the cell wall of Halococcus morrhuae. Arch. Microbiol. 1979, 123, 209–212. [Google Scholar] [CrossRef]
- Steber, J. Untersuchungen zur chemischen Zusammensetzung und Struktur der Zellwand von Halococcus morrhuae. PhD thesis, Technical University, Munich, Germany, 1976. [Google Scholar]
- Koch, A.L. What size should a bacterium be? A question of scale. Annu. Rev. Microbiol. 1996, 50, 317–334. [Google Scholar] [CrossRef]
- Koch, A.L. Were Gram-positive rods the first bacteria? Trends Microbiol. 2003, 11, 166–170. [Google Scholar] [CrossRef]
- Rehm, H.A. Biogenesis of microbial polyhydroxyalkanoate granules: A platform technology for the production of tailor-made bioparticles. Curr. Issues Mol. Biol. 2007, 9, 41–62. [Google Scholar]
- Fernandez-Castillo, R.; Rodriguez-Valera, F.; Gonzales-Ramos, J.; Ruiz-Berraquero, F. Accumulation of poly(β-hydroxybutyrate) by halobacteria. Appl. Environ. Microbiol. 1986, 51, 214–216. [Google Scholar]
- Quillaguamán, J.; Guzmán, H.; Van-Thuoc, D.; Hatti-Kaul, R. Synthesis and production of polyhydroxyalkanoates by halophiles: current potential and future prospects. Appl. Microbiol. Biotechnol. 2010, 85, 1687–1696. [Google Scholar] [CrossRef]
- Legat, A.; Gruber, C.; Zangger, K.; Wanner, G.; Stan-Lotter, H. Identification of polyhydroxyalkanoates in Halococcus and other haloarchaeal species. Appl. Microbiol. Biotechn. 2010, 87, 1119–1127. [Google Scholar] [CrossRef]
- Baliga, N.S.; Bonneau, R.; Facciotti, M.T.; Pan, M.; Glusman, G.; Deutsch, E.W.; Shannon, P.; Chiu, Y.; Weng, R.S.; Gan, R.R.; Hung, P.; Date, S.V.; Marcotte, E.; Hood, L.; Ng, W.V. Genome sequence of Haloarcula marismortui: A halophilic archaeon from the Dead Sea. Genome Res. 2004, 14, 2221–2234. [Google Scholar] [CrossRef]
- Bolhuis, H.; Palm, P.; Wende, A.; Falb, M.; Rampp, M.; Rodriguez-Valera, F.; Pfeiffer, F.; Oesterhelt, D. The genome of the square archaeon Haloquadratum walsbyi: life at the limits of water activity. BMC Genomics 2006, 7, 169. [Google Scholar]
- Han, J.; Lu, Q.; Zhou, L.; Zhou, J.; Xiang, H. Molecular characterization of the phaECHm genes, required for biosynthesis of poly(3-hydroxybutyrate) in the extremely halophilic archaeon Haloarcula marismortui. Appl. Environ. Microbiol. 2007, 73, 6058–6065. [Google Scholar]
- Lu, Q.; Han, J.; Zhou, L.; Zhou, J.; Xiang, H. Genetic and biochemical characterization of the poly(3-hydroxybutyrate-co-3-hydroxyvalerate) synthase in Haloferax mediterranei. J. Bacteriol. 2008, 190, 4173–4180. [Google Scholar] [CrossRef]
- Kalia, V.C.; Lal, S.; Cheema, S. Insight into the phylogeny of polyhydroxyalkanoate biosynthesis: horizontal gene transfer. Gene 2007, 389, 19–26. [Google Scholar] [CrossRef]
- Altschul, S.A.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar]
- Muench, S.P.; Trinick, J.; Harrison, M.A. Structural divergence of the rotary ATPases. Q Rev. Biophys. 2011, 44, 311–356. [Google Scholar] [CrossRef]
- Stan-Lotter, H.; Sulzner, M.; Egelseer, E.; Norton, C.F.; Hochstein, L.I. Comparison of membrane ATPases from extreme halophiles isolated from ancient salt deposits. Origins Life Evol. Biosphere. 1993, 23, 53–64. [Google Scholar] [CrossRef]
- Hochstein, L.I.; Kristjansson, H.; Altekar, W. The purification and subunit structure of a membrane-bound ATPase from the archaebacterium Halobacterium saccharovorum. Biochem. Biophys. Res. Commun. 1987, 147, 295–300. [Google Scholar] [CrossRef]
- Stan-Lotter, H.; Hochstein, L.I. A comparison of an ATPase from the archaebacterium Halobacterium saccharovorum with the F1 moiety from the Escherichia coli ATP synthase. Eur. J. Biochem. 1989, 179, 155–160. [Google Scholar] [CrossRef]
- Ochman, H.; Wilson, A. Evolution in bacteria: evidence for a universal rate in cellular genomes. J. Mol. Evol. 1987, 26, 74–86. [Google Scholar] [CrossRef]
- Park, J.S.; Vreeland, R.H.; Cho, B.C.; Lowenstein, T.K.; Timofeeff, M.N.; Rosenzweig, W.D. Haloarchaeal diversity in 23, 121 and 419 MYA salts. Geobiology 2009, 7, 515–523. [Google Scholar] [CrossRef]
- Mani, K.; Salgaonkar, B.B.; Braganca, J.M. Culturable halophilic archaea at the initial and crystallization stages of salt production in a natural solar saltern of Goa, India. Aquat. Biosyst. 2012, 8, 15. [Google Scholar] [CrossRef]
- Yildiz, E.; Ozcan, B.; Caliskan, M. Isolation, characterization and phylogenetic analysis of halophilic Archaea from a salt mine in central Anatolia (Turkey). Polish J. Microbiol. 2012, 61, 111–117. [Google Scholar]
- Gruber, C.; Legat, A.; Pfaffenhuemer, M.; Radax, C.; Weidler, G.; Busse, H.-J.; Stan-Lotter, H. Halobacterium noricense sp. nov., an archaeal isolate from a bore core of an alpine Permian salt deposit, classification of Halobacterium sp. NRC-1 as a strain of H. salinarum and emended description of H. salinarum. Extremophiles 2004, 8, 431–439. [Google Scholar] [CrossRef]
- Lillo, J.; Rodriguez-Valera, F. Effects of culture conditions on poly(ß -hydroxybutyric acid) production by Haloferax mediterranei. Appl. Environ. Microbiol. 1990, 56, 2517–2521. [Google Scholar]
- Bergmeyer, H.U. Methoden der enzymatischen Analyse; Verlag Chemie: Weinheim, Germany, 1974. [Google Scholar]
- Dodgston, K.S.; Price, R.G. A note on the determination of the ester sulphate content of sulphated polysaccharides. Biochem. J. 1962, 84, 106–110. [Google Scholar]
- Chen, P.S.; Toribara, T.Y.; Warner, H. Microdetermination of phosphorus. Analyt. Chem. 1956, 28, 1756–1758. [Google Scholar]
- Humble, M.W.; King, A.; Phillips, I. API ZYM: A simple rapid system for the detection of bacterial enzymes. J. Clin. Pathol. 1977, 30, 275–277. [Google Scholar] [CrossRef]
- Oren, A.; Ventosa, A.; Grant, W.D. Proposed minimalstandards for description of new taxa in the order Halobacteriales. Int. J. Syst. Bacteriol. 1997, 47, 233–238. [Google Scholar] [CrossRef]
- Cashion, P.; Hodler-Franklin, M.A.; McCully, J.; Franklin, M. A rapid method for the base ratio determination of bacterial DNA. Anal. Biochem. 1977, 81, 461–466. [Google Scholar] [CrossRef]
- De Ley, J.; Cattoir, H.; Reynaerts, A. The quantitative measurement of DNA hybridisation from renaturation rates. Eur. J. Biochem. 1970, 12, 133–142. [Google Scholar] [CrossRef]
- Huß, V.A.R.; Festl, H.; Schleifer, K.H. Studies on the spectrometric determination of DNA hybridisation from renaturation rates. System. Appl. Microbiol. 1983, 4, 184–192. [Google Scholar] [CrossRef]
- Escara, J.F.; Hutton, J.R. Thermal stability and renaturation of DNA in dimethylsulphoxide solutions: acceleration of renaturation rate. Biopolymers 1980, 19, 1315–1327. [Google Scholar] [CrossRef]
- Jahnke, K.-D. BASIC computer program for evaluation of spectroscopic DNA renaturation data from GILFORD System 2600 spectrometer on a PC/XT/AT type personal computer. J. Microbiol. Meth. 1992, 15, 61–73. [Google Scholar] [CrossRef]
- Maidak, B.L.; Cole, J.R.; Lilburn, T.G.; Parker, C.T., Jr.; Saxman, P.R.; Farris, R.J.; Garrity, G.M.; Olsen, G.J.; Schmidt, T.M.; Tiedje, J.M. The RDP-II (Ribosomal Database Project). Nucleic Acids Res. 2001, 29, 173–174. [Google Scholar]
- Jukes, T.H.; Cantor, R.R. Evolution of protein molecules. In Mammalian Protein Metabolism; Munro, H.N, Ed.; Academic Press: New York, NY, USA, 1969; Volume 3, pp. 21–132. [Google Scholar]
- Kumar, S.; Dudley, J.; Nei, M.; Tamura, K. MEGA: A biologist-centric software for evolutionary analysis of DNA and protein sequences. Brief Bioinform. 2008, 9, 299–306. [Google Scholar]
- Thompson, J.D.; Gibson, T.J.; Plewniak, F.; Jeanmougin, F.; Higgins, D.G. The CLUSTAL_X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997, 25, 4876–4882. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Legat, A.; Denner, E.B.M.; Dornmayr-Pfaffenhuemer, M.; Pfeiffer, P.; Knopf, B.; Claus, H.; Gruber, C.; König, H.; Wanner, G.; Stan-Lotter, H. Properties of Halococcus salifodinae, an Isolate from Permian Rock Salt Deposits, Compared with Halococci from Surface Waters. Life 2013, 3, 244-259. https://doi.org/10.3390/life3010244
Legat A, Denner EBM, Dornmayr-Pfaffenhuemer M, Pfeiffer P, Knopf B, Claus H, Gruber C, König H, Wanner G, Stan-Lotter H. Properties of Halococcus salifodinae, an Isolate from Permian Rock Salt Deposits, Compared with Halococci from Surface Waters. Life. 2013; 3(1):244-259. https://doi.org/10.3390/life3010244
Chicago/Turabian StyleLegat, Andrea, Ewald B. M. Denner, Marion Dornmayr-Pfaffenhuemer, Peter Pfeiffer, Burkhard Knopf, Harald Claus, Claudia Gruber, Helmut König, Gerhard Wanner, and Helga Stan-Lotter. 2013. "Properties of Halococcus salifodinae, an Isolate from Permian Rock Salt Deposits, Compared with Halococci from Surface Waters" Life 3, no. 1: 244-259. https://doi.org/10.3390/life3010244




