Long Non-Coding RNAs as Endogenous Target Mimics and Exploration of Their Role in Low Nutrient Stress Tolerance in Plants
Abstract
:1. Introduction
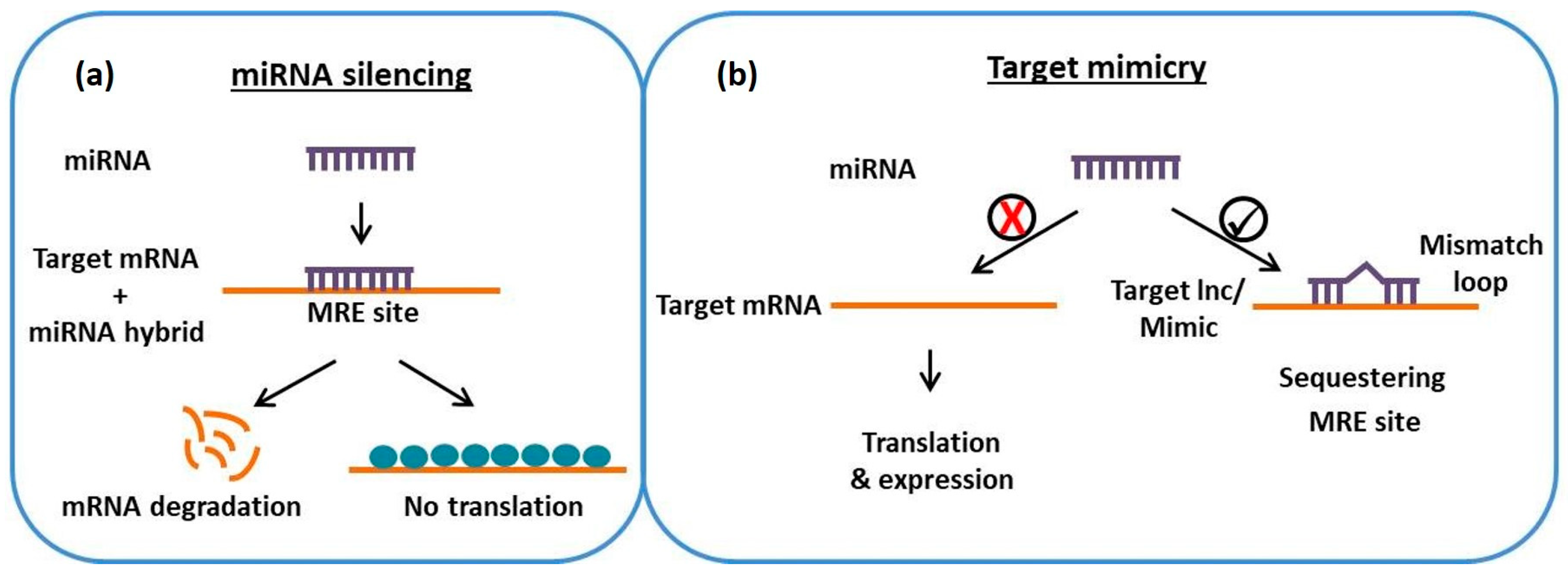
2. Role of lncRNAs as Endogenous Target Mimics for MicroRNAs
3. Long non-coding RNAs Expressed under Nitrogen and Phosphorus Deprivation
4. Putative Endogenous Target Mimics under Low Nitrogen and Phosphate Stress
5. Conclusions and Future Prospects
Funding
Conflicts of Interest
References
- Erdmann, V.A.; Barciszewska, M.Z.; Szymanski, M.; Hochberg, A.; de Groot, N.; Barciszewski, J. The non-coding RNAs as riboregulators. Nucleic Acids Res. 2001, 29, 189–193. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, J.; Meng, X.; Dobrovolskaya, O.B.; Orlov, Y.L.; Chen, M. Non-coding RNAs and their roles in stress response in plants. Genom. Proteom. Bioinform. 2017, 15, 301–312. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Jung, C.; Xu, J.; Wang, H.; Deng, S.; Bernad, L.; Arenas-Huertero, C.; Chua, N.H. Genome-wide analysis uncovers regulation of long intergenic noncoding RNAs in Arabidopsis. Plant Cell 2012, 24, 4333–4345. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.C.; Chang, H.Y. Molecular mechanisms of long noncoding RNAs. Mol. Cell 2011, 43, 904–914. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Hao, L.; Li, D.; Zhu, L.; Hu, S. Long non-coding RNAs and their biological roles in plants. Genom. Proteom. Bioinform. 2015, 3, 137–147. [Google Scholar] [CrossRef] [PubMed]
- Ravasi, T.; Suzuki, H.; Pang, K.C.; Katayama, S.; Furuno, M.; Okunishi, R.; Fukuda, S.; Ru, K.; Frith, M.C.; Gongora, M.M.; et al. Experimental validation of the regulated expression of large numbers of non-coding RNAs from the mouse genome. Genome Res. 2006, 16, 11–19. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guttman, M.; Amit, I.; Garber, M.; French, C.; Lin, M.F.; Feldser, D.; Huarte, M.; Zuk, O.; Carey, B.W.; Cassady, J.P.; et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 2009, 458, 223–227. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kung, J.T.Y.; Colognori, D.; Lee, J.T. Long noncoding RNAs: Past, present, and future. Genetics 2013, 193, 651–669. [Google Scholar] [CrossRef] [PubMed]
- Wapinski, O.; Chang, H.Y. Long noncoding RNAs and human disease. Trends Cell Biol. 2011, 21, 354–361. [Google Scholar] [CrossRef] [PubMed]
- Brannan, C.I.; Dees, E.C.; Ingram, R.S.; Tilghman, S.M. The product of the H19 gene may function as an RNA. Mol. Cell Biol. 1990, 10, 28–36. [Google Scholar] [CrossRef] [PubMed]
- Brown, C.J.; Lafreniere, R.G.; Powers, V.E.; Sebastio, G.; Ballabio, A.; Pettigrew, A.L.; Ledbetter, D.H.; Levy, E.; Craig, I.W.; Willard, H.F. Localization of the X inactivation centre on the human X chromosome in Xq13. Nature 1991, 349, 82–84. [Google Scholar] [CrossRef] [PubMed]
- Bernstein, E.; Allis, C.D. RNA meets chromatin. Gene. Dev. 2005, 19, 1635–1655. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chaumeil, J.; Le Baccon, P.; Wutz, A.; Heard, E. A novel role for Xist RNA in the formation of a repressive nuclear compartment into which genes are recruited when silenced. Genes Dev. 2006, 20, 2223–2237. [Google Scholar] [CrossRef] [PubMed]
- Duret, L.; Chureau, C.; Samain, S.; Weissenbach, J.; Avner, P. The Xist RNA gene evolved in eutherians by pseudogenization of a protein-coding gene. Science 2006, 312, 1653–1655. [Google Scholar] [CrossRef] [PubMed]
- Kapranov, P.; Cheng, J.; Dike, S.; Nix, D.A.; Duttagupta, R.; Willingham, A.T.; Stadler, P.F.; Hertel, J.; Hackermuller, J.; Hofacker, I.L.; et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 2007, 316, 1484–1488. [Google Scholar] [CrossRef] [PubMed]
- Rinn, J.L.; Kertesz, M.; Wang, J.K.; Squazzo, S.L.; Xu, X.; Brugmann, S.A.; Goodnough, L.H.; Helms, J.A.; Farnham, P.J.; Segal, E.; et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 2007, 129, 1311–1323. [Google Scholar] [CrossRef] [PubMed]
- Mercer, T.R.; Dinger, M.E.; Sunkin, S.M.; Mehler, M.F.; Mattick, J.S. Specific expression of long noncoding RNAs in the mouse brain. Proc. Natl. Acad. Sci. USA 2008, 105, 716–721. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, J.; Sun, B.K.; Erwin, J.A.; Song, J.J.; Lee, J.T. Polycomb proteins targeted by a short repeat RNA to the mouse X chromosome. Science 2008, 322, 750–756. [Google Scholar] [CrossRef] [PubMed]
- Nagano, T.; Mitchell, J.A.; Sanz, L.A.; Pauler, F.M.; Ferguson-Smith, A.C.; Feil, R.; Fraser, P. The Air noncoding RNA epigenetically silences transcription by targeting G9a to chromatin. Science 2008, 322, 1717–1720. [Google Scholar] [CrossRef] [PubMed]
- Tsai, M.C.; Manor, O.; Wan, Y.; Mosammaparast, N.; Wang, J.K.; Lan, F.; Shi, Y.; Segal, E.; Chang, H.Y. Long noncoding RNA as modular scaffold of histone modification complexes. Science 2010, 329, 689–693. [Google Scholar] [CrossRef] [PubMed]
- Kaneko, S.; Li, G.; Son, J.; Xu, C.F.; Margueron, R.; Neubert, T.A.; Reinberg, D. Phosphorylation of the PRC2 component Ezh2 is cell cycle-regulated and up-regulates its binding to ncRNA. Genes Dev. 2010, 24, 2615–2620. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dinger, M.E.; Pang, K.C.; Mercer, T.R.; Crowe, M.L.; Grimmond, S.M.; Mattick, J.S. NRED: A database of long noncoding RNA expression. Nucleic Acids Res. 2009, 37, D122–D126. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, L.K.; Yang, Y.T.; Ma, X.; Han, B.; Wang, Z.S.; Zhao, Q.Y.; Wu, L.Q.; Qu, Z.Q. MicroRNA-92b promotes hepatocellular carcinoma progression by targeting Smad7 and is mediated by long non-coding RNA XIST. Cell Death Disease 2016, e2203. [Google Scholar] [CrossRef] [PubMed]
- Haag, J.R.; Pikaard, C.S. Multisubunit RNA polymerases IV and V: Purveyors of non-coding RNA for plant gene silencing. Nat. Rev. Mol. Cell Biol. 2011, 12, 483–492. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Okada, T.; Fukushima, T.; Tsudzuki, T.; Sugiura, M.; Yukawa, Y. A novel hypoxic stress-responsive long non-coding RNA transcribed by RNA polymerase III in Arabidopsis. RNA Biol. 2012, 9, 302–313. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pang, K.C.; Frithand, M.C.; Mattick, J.S. Rapid evolution of noncoding RNAs: lack of conservation does not mean lack of function. Trends Genet. 2006, 22, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Qin, T.; Zhao, H.; Cui, P.; Albesher, N.; Xiong, L. A nucleus-localized long non-coding RNA enhances drought and salt stress tolerance. Plant Physiol. 2017, 175. [Google Scholar] [CrossRef] [PubMed]
- Ding, J.; Lu, Q.; Ouyang, Y.; Mao, H.; Zhang, P.; Yao, J.; Xu, C.; Li, X.; Xiao, J.; Zhang, Q. A long noncoding RNA regulates photoperiod-sensitive male sterility, an essential component of hybrid rice. Proc. Natl. Acad. Sci. USA 2012, 109, 2654–2659. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fan, C.; Hao, Z.; Yan, J.; Li, G. Genome-wide identification and functional analysis of lincRNAs acting as miRNA targets or decoys in maize. BMC Genomics 2015, 16, 793. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.-J.; Wang, Z.-M.; Wang, M.; Wang, X.-J. Widespread long noncoding RNAs as endogenous target mimics for microRNAs in plants. Plant Physiol. 2013, 161, 1875–1884. [Google Scholar] [CrossRef] [PubMed]
- Bardou, F.; Ariel, F.; Simpson, C.G.; Romero-Barrios, N.; Laporte, P.; Balzergue, S.; Brown, J.W.S.; Crespi, M. Long noncoding RNA modulates alternative splicing regulators in Arabidopsis. Dev. Cell 2014, 30, 166–176. [Google Scholar] [CrossRef] [PubMed]
- Seo, J.S.; Sun, H.-X.; Park, B.S.; Huang, C.-H.; Yeh, S.-D.; Jung, C.; Chua, N.-H. ELF18-INDUCED LONG-NONCODING RNA associates with mediator to enhance expression of innate immune response genes in Arabidopsis. Plant Cell 2017, 29, 1024–1038. [Google Scholar] [CrossRef] [PubMed]
- Heo, J.B.; Sung, S. Vernalization-mediated epigenetic silencing by a long intronic noncoding RNA. Science 2011, 331, 76–79. [Google Scholar] [CrossRef] [PubMed]
- Gendall, A.R.; Levy, Y.Y.; Wilson, A.; Dean, C. The VERNALIZATION 2 gene mediates the epigenetic regulation of vernalization in Arabidopsis. Cell 2001, 107, 525–535. [Google Scholar] [CrossRef]
- Wood, C.C.; Robertson, M.; Tanner, G.; Peacock, W.J.; Dennis, E.S.; Helliwell, C.A. The Arabidopsis thaliana vernalization response requires a polycomb-like protein complex that also includes VERNALIZATION INSENSITIVE3. Proc. Natl. Acad. Sci. USA 2006, 103, 14631–14636. [Google Scholar] [CrossRef] [PubMed]
- Nischal, L.; Mohsin, M.; Khan, I.; Kardam, H.; Wadhwa, A.; Abrol, Y.P.; Iqbal, M.; Ahmad, A. Identification and comparative analysis of microRNAs associated with low-N tolerance in rice genotypes. PLoS ONE 2012, 7. [Google Scholar] [CrossRef] [PubMed]
- Elanchezhian, R.; Krishnapriya, V.; Pandey, R.; Rao, A.S.; Abrol, Y.P. Physiological and molecular approaches for improving phosphorus uptake efficiency of crops. Curr. Sci. India 2015, 108, 1271–1279. [Google Scholar]
- Frink, C.R.; Waggoner, P.E.; Ausubel, J.H. Nitrogen fertilizer: retrospect and prospect. Proc. Natl. Acad. Sci. USA 1999, 96, 1175–1180. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Wang, C.; Bao, H.; Chen, H.; Wang, Y. Genome-wide identification and characterization of novel lncRNAs in Populus under nitrogen deficiency. Mol. Genet. Genomics 2016, 291, 1663–1680. [Google Scholar] [CrossRef] [PubMed]
- Lv, Y.; Liang, Z.; Ge, M.; Qi, W.; Zhang, T.; Lin, F.; Peng, Z.; Zhao, H. Genome-wide identification and functional prediction of nitrogen-responsive intergenic and intronic long non-coding RNAs in maize (Zea mays L.). BMC Genomics 2016, 17, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Amaral, N.S.D.; Melo, N.C.; Maia, B.d.M.; Rocha, R.M. Noncoding RNA profiles in tobacco- and alcohol-associated diseases. Genes 2017, 8, 6. [Google Scholar] [CrossRef] [PubMed]
- Steinkraus, B.R.; Toegel, M.; Fulga, T.A. Tiny giants of gene regulation: Experimental strategies for microRNA functional studies. Wiley Interdiscip. Rev. Dev. Biol. 2016, 5, 311–362. [Google Scholar] [CrossRef] [PubMed]
- Dogini, D.B.; Pascoal, V.D.B.; Avansini, S.H.; Vieira, A.S.; Pereira, T.C.; Lopes-Cendes, I. The new world of RNAs. Genet. Mol. Biol. 2014, 37, 285–293. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lozada-Chávez, I.; Stadler, P.F.; Prohaska, S.J. Hypothesis for the modern RNA world: A pervasive non-coding RNA-based genetic regulation is a prerequisite for the emergence of multicellular complexity. Orig. Life Evol. Biosph. 2011, 41, 587–607. [Google Scholar] [CrossRef] [PubMed]
- Ameres, S.L.; Zamore, P.D. Diversifying microRNA sequence and function. Nat. Rev. Mol. Cell Biol. 2013, 14, 475–488. [Google Scholar] [CrossRef] [PubMed]
- Franco-Zorrilla, J.M.; Valli, A.; Todesco, M.; Mateos, I.; Puga, M.I.; Rubio-Somoza, I.; Leyva, A.; Weigel, D.; García, J.A.; Paz-Ares, J. Target mimicry provides a new mechanism for regulation of microRNA activity. Nat. Genet. 2007, 39, 1033–1037. [Google Scholar] [CrossRef] [PubMed]
- Salmena, L.; Poliseno, L.; Tay, Y.; Kats, L.; Pandolfi, P.P. A ceRNA hypothesis: The Rosetta stone of a hidden RNA language? Cell 2011, 146, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Kartha, R.V.; Subramanian, S. Competing endogenous RNAs (ceRNAs): New entrants to the intricacies of gene regulation. Front. Genet. 2014, 5, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Todesco, M.; Rubio-Somoza, I.; Paz-Ares, J.; Weigel, D. A collection of target mimics for comprehensive analysis of MicroRNA function in Arabidopsis thaliana. PLoS Genet. 2010, 6, e1001031. [Google Scholar] [CrossRef] [PubMed]
- Reichel, M.; Li, Y.; Li, J.; Millar, A.A. Inhibiting plant microRNA activity: Molecular SPONGEs, target MIMICs and STTMs all display variable efficacies against target microRNAs. Plant Biotechnol. J. 2015, 13, 915–926. [Google Scholar] [CrossRef] [PubMed]
- Liu, Q.; Wang, F.; Axtell, M.J. Analysis of complementarity requirements for plant microRNA targeting using a Nicotiana benthamiana quantitative transient assay. Plant Cell 2014, 26, 741–753. [Google Scholar] [CrossRef] [PubMed]
- Strachan, T.; Read, A.P. Human Molecular Genetics; Garland Science, Taylor & Francis Group: New York, NY, USA, 2011. [Google Scholar]
- Quek, X.C.; Thomson, D.W.; Maag, J.L.V.; Bartonicek, N.; Signal, B.; Clark, M.B.; Gloss, B.S.; Dinger, M.E. lncRNAdb v2.0: Expanding the reference database for functional long noncoding RNAs. Nucleic Acids Res. 2014, 43, D168–D173. [Google Scholar] [CrossRef] [PubMed]
- Gupta, P.K. MicroRNAs and target mimics for crop improvement. Curr. Sci. India 2015, 108, 1624–1633. [Google Scholar]
- Bonnet, E.; He, Y.; Billiau, K.; Van de Peer, Y. TAPIR, a web server for the prediction of plant microRNA targets, including target mimics. Bioinformatics 2010, 26, 1566–1568. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Karakulah, G.; Yucebilgili-Kurtoglu, K.; Unver, T. PeTMbase: A Database of Plant Endogenous Target Mimics (eTMs). PLoS ONE 2016, 11, e0167698. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Zhi, H.; Zhang, Y.; Liu, Y.; Zhang, J.; Gao, Y.; Guo, M.; Ning, S.; Li, X. miRSponge: A manually curated database for experimentally supported miRNA sponges and ceRNAs. Database (Oxford) 2015, bav0982015. [Google Scholar] [CrossRef] [PubMed]
- Yuan, C.; Meng, X.; Li, X.; Illing, N.; Ingle, R.A.; Wang, J.; Chen, M. PceRBase: A database of plant competing endogenous RNA. Nucleic Acids Res. 2017, 45, D1009–D1014. [Google Scholar] [CrossRef] [PubMed]
- Bari, R.; Pant, B.D.; Stitt, M.; Scheible, W.R. PHO2, MicroRNA399, and PHR1 define a phosphate-signaling pathway in plants. Plant Physiol. 2006, 141, 988–999. [Google Scholar] [CrossRef] [PubMed]
- Pant, B.D.; Buhtz, A.; Kehr, J.; Scheible, W.R. MicroRNA399 is a long-distance signal for the regulation of plant phosphate homeostasis. Plant J. 2008, 53, 731–738. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- López-Arredondo, D.L.; Leyva-Gonzalez, M.A.; Gonzalez-Morales, S.I.; Lopez-Bucio, J.; Herrera-Estrella, L. Phosphatenutrition: Improvinglow-phosphatetolerance in crops. Annu. Rev. Plant Biol. 2014, 65. [Google Scholar] [CrossRef]
- Vidal, E.A.; Moyano, T.C.; Canales, J.; Gutiérrez, R.A. Nitrogen control of developmental phase transitions in Arabidopsis thaliana. J. Exp. Bot. 2014, 65, 5611–5618. [Google Scholar] [CrossRef] [PubMed]
- Alvarez, J.M.; Riveras, E.; Vidal, E.A.; Gras, D.E.; Contreras-López, O.; Tamayo, K.P.; Aceituno, F.; Gómez, I.; Ruffel, S.; LejayChandra, L.; et al. Systems approach identifies TGA1 and TGA4 transcription factors as important regulatory components of the nitrate response of Arabidopsis thaliana roots. Plant J. 2014, 80, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Ganie, A.H.; Ahmad, A.; Pandey, R.; Aref, I.M.; Yousuf, P.Y.; Ahmad, S.; Iqbal, M. Metabolite profiling of low-P tolerant and low-P sensitive maize genotypes under phosphorus starvation and restoration conditions. PLoS ONE 2015, 10, e0129520. [Google Scholar] [CrossRef] [PubMed]
- Nazir, M.; Pandey, R.; Siddiqi, T.O.; Ibrahim, M.M.; Qureshi, I.M.; Vengavasi, K.; Abraham, G.; Ahmad, A. Nitrogen-deficiency stress induces protein expression differentially in low-N tolerant and low-N sensitive maize genotypes. Front. Plant Sci. 2016, 7, 298. [Google Scholar] [CrossRef] [PubMed]
- Giri, J.; Bhosale, R.; Huang, G.; Pandey, B.K.; Parker, H.; Zappala, S.; Yang, J.; Dievart, A.; Bureau, C.; Ljung, K.; et al. Rice auxin influx carrier OsAUX1 facilitates root hair elongation in response to low external phosphate. Nat. Commun. 2018, 9, 1–7. [Google Scholar] [CrossRef]
- Gutierrez, R.A. Systems biology for enhanced plant nitrogen nutrition. Science 2012, 336, 1673–1675. [Google Scholar] [CrossRef] [PubMed]
- Ehrhardt, D.W.; Frommer, W.B. New Technologies for 21st Century Plant Science. Plant Cell 2012, 24, 374–394. [Google Scholar] [CrossRef] [PubMed]
- Agrama, H.A.S.; Zakaria, A.G.; Said, F.B.; Tuinstra, M. Identification of quantitative trait loci for nitrogen use efficiency in maize. Mol. Breeding 1999, 5, 187–195. [Google Scholar] [CrossRef]
- Garnett, T.; Conn, V.; Kaiser, B.N. Root based approaches to improving nitrogen use efficiency in plants. Plant Cell Environ. 2009, 32, 1272–1283. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Curci, P.L.; AieseCigliano, R.; Zuluaga, D.L.; Janni, M.; Sanseverino, W.; Sonnante, G. Transcriptomic response of durum wheat to nitrogen starvation. Sci. Rep. 2017, 7, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Gelli, M.; Duo, Y.; Konda, A.R.; Zhang, C.; Holding, D.; Dweikat, I. Identification of differentially expressed genes between sorghum genotypes with contrasting nitrogen stress tolerance by genome-wide transcriptional profiling. BMC Genomics 2014, 15, 179. [Google Scholar] [CrossRef] [PubMed]
- Quan, X.; Zeng, J.; Ye, L.; Chen, G.; Han, Z.; Shah, J.M.; Zhang, G. Transcriptome profiling analysis for two Tibetan wild barley genotypes in responses to low nitrogen. BMC Plant Biol. 2016. [Google Scholar] [CrossRef] [PubMed]
- Sinha, S.K.; Amitha Mithra, S.V.; Chaudhary, S.; Tyagi, P.; Venkadesan, S.; Rani, M.; Mandal, P.K. Transcriptome analysis of two rice varieties contrasting for nitrogen use efficiency under chronic N starvation reveals differences in chloroplast and starch metabolism-related genes. Genes 2018. [Google Scholar] [CrossRef] [PubMed]
- Marschner, H.; Marschner, P. Mineral Nutrition of Higher Plants, 3rd ed.; Elsevier/Academic Press: London, UK; Waltham, MA, USA, 2012. [Google Scholar]
- Pandey, R.; Zinta, G.; AbdElgawad, H.; Ahmad, A.; Jain, V.; Janssens, I.A. Physiological and molecular alterations in plants exposed to high [CO2] under phosphorus stress. Biotechnol. Adv. 2015, 33, 303–316. [Google Scholar] [CrossRef] [PubMed]
- López-Bucio, J.; Herńandez-Abreu, E.; Sánchez-Calderón, L.; Nieto-Jacobo, M.F.; Simpson, J.; Herrera-Estrella, L. Phosphate availability alters architecture and causes changes hormone sensitivity in the Arabidopsis root system. Plant Physiol. 2002, 129, 244–256. [Google Scholar] [CrossRef] [PubMed]
- Nacry, P.; Canivenc, G.; Muller, B.; Azmi, A.; Van Onckelen, V.; Rossignol, M.; Doumas, P. A role for auxin redistribution in the responses of the root system architecture to phosphate starvation in Arabidopsis. Plant Physiol. 2005, 138, 2061–2074. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Calderón, L.; López-Bucio, J.; Chacón-López, A.; Cruz-Ramírez, A.; Nieto-Jacobo, F.; Dubrovsky, J.G.; Herrera-Estrella, J. Phosphate starvation induces a determinate developmental program in the roots of Arabidopsis thaliana. Plant Cell Physiol. 2006, 46, 174–184. [Google Scholar] [CrossRef] [PubMed]
- Svistoonoff, S.; Creff, A.; Reymond, M.; Sigoillot-Claude, C.; Ricaud, L.; Blanchet, A.; Nussaume, L.; Desnos, T. Root tip contact with low-phosphate media reprograms plant root architecture. Nat. Genet. 2007, 39, 792–796. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wang, Y.-L.; Almvik, M.; Clarke, N.; Eich-Greatorex, S.; Øgaard, A.F.; Krogstad, T.; Lambers, H.; Clarke, J.L. Contrasting responses of root morphology and root-exuded organic acids to low phosphorus availability in three important food crops with divergent root traits. AoB Plants 2015, 7, plv097. [Google Scholar] [CrossRef] [PubMed]
- Vengavasi, K.; Pandey, R. Root exudation index: Screening organic acid exudation and phosphorus acquisition efficiency in soybean genotypes. Crop Pasture Sci. 2016, 67, 1096–1109. [Google Scholar] [CrossRef]
- Vengavasi, K.; Pandey, R.; Abraham, G.; Yadav, R.K. Comparative analysis of soybean root proteome reveals molecular basis of differential carboxylate efflux under low phosphorus stress. Genes 2017, 8, 341. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Krogstad, T.; Clarke, N.; Øgaard, A.F.; Clarke, J.L. Impact of phosphorus on rhizosphere organic anions of wheat at different growth stages under field conditions. AoB Plants 2017, 9, plx008. [Google Scholar] [CrossRef]
- Vengavasi, K.; Pandey, R. Root exudation potential in contrasting soybean genotypes in response to low soil phosphorus availability is determined by photo-biochemical processes. Plant Physiol. Biochem. 2018, 124, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Palma, D.A.; Blumwald, E.; Plaxton, W.C. Upregulation of vascular H+-translocating pyrophosphatase by phosphate starvation of Brassica napus (rapeseed) suspension cell cultures. FEBS Lett. 2000, 486, 155–158. [Google Scholar] [CrossRef]
- Theodorou, M.E.; Plaxton, W.C. Purification and characterization of pyrophosphate dependent phosphofructokinase from phosphate-starved Brassica nigra suspension cells. Plant Physiol. 1996, 112, 343–351. [Google Scholar] [CrossRef] [PubMed]
- Plaxton, W.C.; Tran, H.T. Metabolic adaptations of phosphate-starved plants. Plant Physiol. 2011, 156, 1006–1015. [Google Scholar] [CrossRef] [PubMed]
- Vengavasi, K.; Kumar, A.; Pandey, R. Transcript abundance, enzyme activity and metabolite concentration regulates differential carboxylate efflux in soybean under low phosphorus stress. Indian J. Plant Physi. 2016, 21, 179–188. [Google Scholar] [CrossRef]
- Chiou, T.J.; Aung, K.; Lin, S.-I.; Wu, C.-C.; Chiang, S.-F.; Su, C.-L. Regulation of phosphate homeostasis by microRNA in Arabidopsis. Plant Cell 2006, 18, 412–421. [Google Scholar] [CrossRef] [PubMed]
- Doerner, P. Phosphate starvation signaling: A threesome controls systemic Pi homeostasis. Curr. Opin. Plant Biol. 2008, 11, 536–540. [Google Scholar] [CrossRef] [PubMed]
- Lin, W.Y.; Lin, S.I.; Chiou, T.J. Molecular regulators of phosphate homeostasis in plants. J. Exp. Bot. 2009, 60, 1427–1438. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shin, H.; Shin, H.-S.; Chen, R.; Harrison, M.J. Loss of At4 function impacts phosphate distribution between the roots and the shoots during phosphate starvation. Plant J. 2006, 45, 712–726. [Google Scholar] [CrossRef] [PubMed]
- Kuo, H.F.; Chiou, T.J. The role of microRNAs in phosphorus deficiency signaling. Plant Physiol. 2011, 156, 1016–1024. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Zhao, M.; Zhang, X.; Liu, M.; Yang, C.; Chen, Y.; Chen, R.; Wen, J.; Mysore, K.S.; Zhang, W.H. Novel phosphate deficiency-responsive long non-coding RNAs in the legume model plant Medicago truncatula. J. Exp. Bot. 2017, 68, 5937–5948. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Muchhal, U.S.; Raghothama, K.G. Differential expression of TPSI1, a phosphate starvation-induced gene in tomato. Plant Mol. Biol. 1997, 33, 867–874. [Google Scholar] [CrossRef] [PubMed]
- Burleigh, S.M.; Harrison, M.J. Characterization of the Mt4 gene from Medicago truncatula. Gene 1998, 216, 47–53. [Google Scholar] [CrossRef]
- Burleigh, S.H.; Harrison, M.J. The down-regulation of Mt4-like genes by phosphate fertilization occurs systemically and involves phosphate translocation to the shoots. Plant Physiol. 1999, 119, 241–248. [Google Scholar] [CrossRef] [PubMed]
- Di, C.; Yuan, J.; Wu, Y.; Li, J.; Lin, H.; Hu, L.; Zhang, T.; Qi, Y.; Gerstein, M.B.; Guo, Y.; et al. Characterization of stress-responsive lncRNAs in Arabidopsis thaliana by integrating expression, epigenetic and structural features. Plant J. 2014, 80, 848–861. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.; Zhang, Y.; Dong, J.; Sun, Y.; Lim, B.L.; Liu, D.; Lu, Z.J. Systematic characterization of novel lncRNAs responding to phosphate starvation in Arabidopsis thaliana. BMC Genomics 2016, 17, 655. [Google Scholar] [CrossRef] [PubMed]
- Carvalho, M.H.C.; Sun, H.; Bowler, C.; Chua, N. Noncoding and coding transcriptome responses of a marine diatom to phosphate fluctuations. New Phytol. 2016, 210, 497–510. [Google Scholar] [CrossRef] [PubMed]
- Griffiths-Jones, S.; Grocock, R.J.; van Dongen, S.; Bateman, A.; Enright, A.J. miRbase: microRNA sequences, targets and gene nomenclature. Nucleic Acid Res. 2006, 34, D140–D144. [Google Scholar] [CrossRef] [PubMed]
- Paul, S.; Datta, S.K.; Datta, K. miRNA regulation of nutrient homeostasis in plants. Front. Plant Sci. 2015, 6, 232. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, G.N.; Rothstein, S.J.; Spangenberg, G.; Kant, S. Role of microRNAs involved in plant response to nitrogen and phosphorous limiting conditions. Front. Plant Sci. 2015, 6, 629. [Google Scholar] [CrossRef] [PubMed]
- Chen, X. A MicroRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 2004, 303, 2022–2025. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Bernatavichute, Y.V.; Cokus, S.; Pellegrini, M.; Jacobsen, S.E. Genome-wide analysis of mono-, di- and trimethylation of histone H3 lysine 4 in Arabidopsis thaliana. Genome Biol. 2009, 10, R62. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Ding, H.; Zhu, J.K.; Zhang, F.; Li, W.X. Involvement of miR169 in the nitrogen-starvation responses in Arabidopsis. New Phytol. 2011, 190, 906–915. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Ying, S.; Huang, H.; Li, K.; Wu, P.; Shou, H. Involvement of OsSPX1 in phosphate homeostasis in rice. Plant J. 2009, 57, 895–904. [Google Scholar] [CrossRef] [PubMed]
- Duan, K.; Yi, K.; Dang, L.; Huang, H.; Wu, W.; Wu, P. Characterization of a sub-family of Arabidopsis genes with the SPX domain reveals their diverse functions in plant tolerance to phosphorus starvation. Plant J. 2008, 54, 965–975. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Huang, W.; Ying, Y.; Li, S.; Secco, D.; Tyerman, S.; Whelan, J.; Shou, H. Functional characterization of the rice SPX-MFS family reveals a key role of OsSPX-MFS1 in controlling phosphate homeostasis in leaves. New Phytol. 2012, 196, 139–148. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ligaba, A.; Yamaguchi, M.; Shen, H.; Sasaki, T.; Yamamoto, Y.; Matsumoto, H. Phosphorus deficiency enhances plasma membrane HC-ATPase activity and citrate exudation in greater purple lupin (Lupinuspilosus). Funct. Plant Biol. 2004, 31, 1075–1083. [Google Scholar] [CrossRef]
- Hsieh, L.C.; Lin, S.I.; Shih, A.C.C.; Chen, J.W.; Lin, W.Y.; Tseng, C.Y.; Li, W.H.; Chiou, T.J. Uncovering small RNA-mediated responses to phosphate deficiency in Arabidopsis by deep sequencing. Plant Physiol. 2009, 151, 2120–2132. [Google Scholar] [CrossRef] [PubMed]
- Liang, G.; He, H.; Yu, D. Identification of nitrogen starvation-responsive MicroRNAs in Arabidopsis thaliana. PLoS ONE 2012, 7, e48951. [Google Scholar] [CrossRef] [PubMed]
- Zeng, H.Q.; Zhu, Y.Y.; SQ, H.; Yang, Z.M. Analysis of phosphorus-deficient responsive miRNAs and cis-elements from soybean (Glycine max L.). J. Plant Physiol. 2010, 167, 1289–1297. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Liu, X.; Guo, C.; Gu, J.; Xiao, K. Identification and characterization of micro RNAs from wheat (Triticum aestivum L.) under phosphorus deprivation. J. Plant Biochem. Biot. 2013, 22, 113–123. [Google Scholar] [CrossRef]
- Xu, Z.; Zhong, S.; Li, X.; Li, W.; Rothstein, S.J.; Zhang, S.; Bi, Y.; Xie, C. Genome wide identification of microRNAs in response to low nitrate availability in maize leaves and roots. PLoS ONE 2011, 6, e28009. [Google Scholar] [CrossRef] [PubMed]
- Trevisan, S.; Nonis, A.; Begheldo, M.; Manoli, A.; Palme, K.; Caporale, G.; Ruperti, B.; Quaggiotti, S. Expression and tissue-specific localization of nitrate-responsive miRNAs in roots of maize seedlings. Plant Cell Environ. 2012, 35, 1137–1155. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhang, C.; Hao, Q.; Sha, A.; Zhou, R.; Zhou, X.; Yuan, L. Elucidation of miRNAs-mediated responses to low nitrogen stress by deep sequencing of two soybean genotypes. PLoS ONE 2013, 8, e67423. [Google Scholar] [CrossRef] [PubMed]
- Yan, Y.; Wang, H.; Hamera, H.; Chen, X.; Fang, R. miR444a has multiple functions in the rice nitrate-signaling pathway. Plant J. 2014, 78, 44–55. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rogers, E.D.; Jackson, J.; Moussaieff, A.; Aharoni, A.; Benfey, P.N. Cell types specific transcriptional profiling: Implications for metabolite profiling. Plant J. 2012, 70, 5–17. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.C.; Liao, J.; Li, Z.; Yu, Y.; Zhang, J.; Li, Q.; Qu, L.; Shu, W.; Chen, Y. Genome-wide screening and functional analysis identify a large number of long noncoding RNAs involved in the sexual reproduction of rice. Genome Biol. 2014, 15, 512. [Google Scholar] [CrossRef] [PubMed] [Green Version]

| Name | Features | Links | Reference |
|---|---|---|---|
| TAPIR | Only tool for the TM prediction in plants; applies Franco-Zorrilla rule for target mimicry; contains data for 10 plant species; RNA hybrid and miRBase are the data sources | http://bioinformatics.psb.ugent.be/webtools/tapir/ | [55] |
| miRSponge | 1.6% data is from plant and others are of non-plant; experimentally validated; literature mining is the data source | http://www.bio-bigdata.net/miRSponge/ | [57] |
| PeTMbase | Contains 2728 TMs for 11 species; uses Wu et al. (2013) target mimicry rule; GreeNC, PNRD, miRBase, NCBI SRA are the data sources | http://petmbase.org | [56] |
| PceRBase | First database for plant TMs; 167608 TMs from 26 plant species; Phytozomev10, TAIR10, MSU RGGP & miRBase and literature are the data sources | http://bis.zju.edu.cn/pcernadb/ | [58] |
| miRNA ID | eTM ID | lncRNA ID | Low N | Low P | References |
|---|---|---|---|---|---|
| ath-miR156a-5p | ath_eTM_miR156a-5p-2 | GRNC_ath_lcl|Athaliana_AT1G52347.1 gene = AT1G52347 | A. thaliana (P↑) | [112] | |
| ath_eTM_miR156a-5p-1 | PNRD_ath_NONATHT000580 | ||||
| ath-miR156b-5p | ath_eTM_miR156b-5p-2 | GRNC_ath_lcl|Athaliana_AT1G52347.1 gene = AT1G52347 | A. thaliana (N↑) | [113] | |
| ath_eTM_miR156b-5p-1 | PNRD_ath_NONATHT000580 | ||||
| ath-miR156c-5p | ath_eTM_miR156c-5p-2 | GRNC_ath_lcl|Athaliana_AT1G52347.1 gene = AT1G52347 | A. thaliana (N↑) | [113] | |
| ath_eTM_miR156c-5p-1 | PNRD_ath_NONATHT000580 | ||||
| ath-miR156d-5p | ath_eTM_miR156d-5p-2 | GRNC_ath_lcl|Athaliana_AT1G52347.1 gene = AT1G52347 | A. thaliana (N↑) | [113] | |
| ath_eTM_miR156d-5p-1 | PNRD_ath_NONATHT000580 | ||||
| ath-miR156e | ath_eTM_miR156e-2 | GRNC_ath_lcl|Athaliana_AT1G52347.1 gene = AT1G52347 | A. thaliana (N↑) | [113] | |
| ath_eTM_miR156e-1 | PNRD_ath_NONATHT000580 | ||||
| gma-miR156e | gma_eTM_miR156e-3 | GRNC_gma_lcl|Gmax_Glyma.18G293400.2 | Soybean (P↑) | [114] | |
| gma_eTM_miR156e-2 | GRNC_gma_lcl|Gmax_Glyma.18G293400.1 | ||||
| gma_eTM_miR156e-1 | GRNC_gma_lcl|Gmax_Glyma.05G242200.1 | ||||
| ath-miR156f-5p | ath_eTM_miR156f-5p-2 | GRNC_ath_lcl|Athaliana_AT1G52347.1 gene = AT1G52347 | A. thaliana (N↑) | [113] | |
| ath_eTM_miR156f-5p-1 | PNRD_ath_NONATHT000580 | ||||
| ath-miR156g | ath_eTM_miR156g-2 | GRNC_ath_lcl|Athaliana_AT1G52347.1 gene = AT1G52347 | A. thaliana (N↑) | [113] | |
| ath_eTM_miR156g-1 | PNRD_ath_NONATHT000580 | ||||
| ath-miR156h | ath_eTM_miR156h-3 | GRNC_ath_lcl|Athaliana_AT3G18217.1 gene = AT3G18217 | A. thaliana (N↑) | [113] | |
| ath_eTM_miR156h-2 | GRNC_ath_lcl|Athaliana_AT1G52347.1 gene = AT1G52347 | ||||
| ath_eTM_miR156h-1 | PNRD_ath_NONATHT000580 | ||||
| gma-miR159a-3p | gma_eTM_miR159a-3p-1 | gma_TCONS_00088249 | Soybean (P↑) | [114] | |
| tae-miR159b | tae_eTM_miR159b-2 | GRNC_tae_lcl|Taestivum_Traes_2DS_9A9CAF0B0.1 | Wheat(P↑) | [115] | |
| tae_eTM_miR159b-1 | GRNC_tae_lcl|Taestivum_Traes_1AL_8C8E43898.1 | ||||
| zma-miR160a-3p | zma_eTM_miR160a-3p-1 | GRNC_zma_lcl|Zmays_AC211588.3_FGT002 | Maize (N↑) | [116] | |
| zma-miR160a-5p | zma_eTM_miR160a-5p-5 | GRNC_zma_lcl|Zmays_GRMZM5G849473_T01 | Maize (N↑) | [116] | |
| zma_eTM_miR160a-5p-4 | GRNC_zma_lcl|Zmays_GRMZM2G531719_T01 | ||||
| zma_eTM_miR160a-5p-3 | GRNC_zma_lcl|Zmays_GRMZM2G149698_T05 | ||||
| zma_eTM_miR160a-5p-2 | GRNC_zma_lcl|Zmays_GRMZM2G011007_T01 | ||||
| zma_eTM_miR160a-5p-1 | PNRD_zma_GRMZM5G849473_T01 | ||||
| zma-miR160b-5p | zma_eTM_miR160b-5p-5 | GRNC_zma_lcl|Zmays_GRMZM5G849473_T01 | Maize (N↑) | [116] | |
| zma_eTM_miR160b-5p-4 | GRNC_zma_lcl|Zmays_GRMZM2G531719_T01 | ||||
| zma_eTM_miR160b-5p-3 | GRNC_zma_lcl|Zmays_GRMZM2G149698_T05 | ||||
| zma_eTM_miR160b-5p-2 | GRNC_zma_lcl|Zmays_GRMZM2G011007_T01 | ||||
| zma_eTM_miR160b-5p-1 | PNRD_zma_GRMZM5G849473_T01 | ||||
| zma-miR160c-5p | zma_eTM_miR160c-5p-5 | GRNC_zma_lcl|Zmays_GRMZM5G849473_T01 | Maize (N↑) | [116] | |
| zma_eTM_miR160c-5p-4 | GRNC_zma_lcl|Zmays_GRMZM2G531719_T01 | ||||
| zma_eTM_miR160c-5p-3 | GRNC_zma_lcl|Zmays_GRMZM2G149698_T05 | ||||
| zma_eTM_miR160c-5p-2 | GRNC_zma_lcl|Zmays_GRMZM2G011007_T01 | ||||
| zma_eTM_miR160c-5p-1 | PNRD_zma_GRMZM5G849473_T01 | ||||
| zma-miR160d-3p | zma_eTM_miR160d-3p-3 | GRNC_zma_lcl|Zmays_GRMZM2G064666_T02 | Maize (N↑) | [113] | |
| zma_eTM_miR160d-3p-2 | GRNC_zma_lcl|Zmays_GRMZM2G054392_T01 | ||||
| zma_eTM_miR160d-3p-1 | GRNC_zma_lcl|Zmays_GRMZM2G052412_T01 | ||||
| zma-miR160d-5p | zma_eTM_miR160d-5p-5 | GRNC_zma_lcl|Zmays_GRMZM5G849473_T01 | Maize (N↑) | [116] | |
| zma_eTM_miR160d-5p-4 | GRNC_zma_lcl|Zmays_GRMZM2G531719_T01 | ||||
| zma_eTM_miR160d-5p-3 | GRNC_zma_lcl|Zmays_GRMZM2G149698_T05 | ||||
| zma_eTM_miR160d-5p-2 | GRNC_zma_lcl|Zmays_GRMZM2G011007_T01 | ||||
| zma_eTM_miR160d-5p-1 | PNRD_zma_GRMZM5G849473_T01 | ||||
| zma-miR160e | zma_eTM_miR160e-5 | GRNC_zma_lcl|Zmays_GRMZM5G849473_T01 | Maize (N↑) | [116] | |
| zma_eTM_miR160e-4 | GRNC_zma_lcl|Zmays_GRMZM2G531719_T01 | ||||
| zma_eTM_miR160e-3 | GRNC_zma_lcl|Zmays_GRMZM2G149698_T05 | ||||
| zma_eTM_miR160e-2 | GRNC_zma_lcl|Zmays_GRMZM2G011007_T01 | ||||
| zma_eTM_miR160e-1 | PNRD_zma_GRMZM5G849473_T01 | ||||
| zma-miR160g-5p | zma_eTM_miR160g-5p-5 | GRNC_zma_lcl|Zmays_GRMZM5G849473_T01 | Maize (N↑) | [116] | |
| zma_eTM_miR160g-5p-4 | GRNC_zma_lcl|Zmays_GRMZM2G531719_T01 | ||||
| zma_eTM_miR160g-5p-3 | GRNC_zma_lcl|Zmays_GRMZM2G149698_T05 | ||||
| zma_eTM_miR160g-5p-2 | GRNC_zma_lcl|Zmays_GRMZM2G011007_T01 | ||||
| zma_eTM_miR160g-5p-1 | PNRD_zma_GRMZM5G849473_T01 | ||||
| zma-miR164f-5p | zma_eTM_miR164f-5p-1 | GRNC_zma_lcl|Zmays_GRMZM2G008252_T01 | Maize (N↑) | [116] | |
| zma-miR164f-3p | zma_eTM_miR164f-3p-1 | GRNC_zma_lcl|Zmays_GRMZM5G837428_T01 | Maize (N↑) | [113] | |
| zma-miR166j-3p | zma_eTM_miR166j-3p-4 | GRNC_zma_lcl|Zmays_GRMZM2G134604_T01 | Maize (N↓) | [117] | |
| zma_eTM_miR166j-3p-3 | zma_eTM_miR166j-3p-3 | ||||
| zma_eTM_miR166j-3p-2 | zma_eTM_miR166j-3p-2 | ||||
| zma_eTM_miR166j-3p-1 | zma_TCONS_00089106 | ||||
| zma-miR166k-3p | zma_eTM_miR166k-3p-4 | GRNC_zma_lcl|Zmays_GRMZM2G134604_T01 | Maize (N↓) | [117] | |
| zma_eTM_miR166k-3p-3 | GRNC_zma_lcl|Zmays_GRMZM2G110279_T02 | ||||
| zma_eTM_miR166k-3p-2 | GRNC_zma_lcl|Zmays_GRMZM2G110279_T01 | ||||
| zma_eTM_miR166k-3p-1 | zma_TCONS_00089106 | ||||
| zma-miR166n-3p | zma_eTM_miR166n-3p-4 | GRNC_zma_lcl|Zmays_GRMZM2G134604_T01 | Maize (N↓) | [117] | |
| zma_eTM_miR166n-3p-3 | GRNC_zma_lcl|Zmays_GRMZM2G110279_T02 | ||||
| zma_eTM_miR166n-3p-2 | GRNC_zma_lcl|Zmays_GRMZM2G110279_T01 | ||||
| zma_eTM_miR166n-3p-1 | zma_TCONS_00089106 | ||||
| zma-miR167g-3p | zma_eTM_miR167g-3p-1 | zma_TCONS_00081049 | Maize (N↓) | [116] | |
| zma-miR167g-5p | zma_eTM_miR167g-5p-2 | GRNC_zma_lcl|Zmays_GRMZM2G174168_T02 | Maize (N↓) | [116] | |
| zma_eTM_miR167g-5p-1 | GRNC_zma_lcl|Zmays_GRMZM2G174168_T01 | ||||
| zma-miR167h-3p | zma_eTM_miR167h-3p-7 | GRNC_zma_lcl|Zmays_GRMZM2G326635_T01 | Maize (N↓) | [116] | |
| zma_eTM_miR167h-3p-6 | GRNC_zma_lcl|Zmays_GRMZM2G175272_T03 | ||||
| zma_eTM_miR167h-3p-5 | GRNC_zma_lcl|Zmays_GRMZM2G159741_T04 | ||||
| zma_eTM_miR167h-3p-4 | GRNC_zma_lcl|Zmays_GRMZM2G158766_T04 | ||||
| zma_eTM_miR167h-3p-3 | GRNC_zma_lcl|Zmays_GRMZM2G125239_T04 | ||||
| zma_eTM_miR167h-3p-2 | PNRD_zma_TCONS_00034773 | ||||
| zma_eTM_miR167h-3p-1 | zma_TCONS_00012947 | ||||
| zma-miR167h-5p | zma_eTM_miR167h-5p-2 | GRNC_zma_lcl|Zmays_GRMZM2G174168_T02 | Maize (N↓) | [116] | |
| zma_eTM_miR167h-5p-1 | GRNC_zma_lcl|Zmays_GRMZM2G174168_T01 | ||||
| gma-miR169f | gma_eTM_miR169f-3 | GRNC_gma_lcl|Gmax_Glyma.19G136600.8 | Soybean (N↑) | [118] | |
| gma_eTM_miR169f-2 | GRNC_gma_lcl|Gmax_Glyma.19G136600.3 | ||||
| gma_eTM_miR169f-1 | GRNC_gma_lcl|Gmax_Glyma.19G136600.2 | ||||
| gma-miR169g | gma_eTM_miR169g-3 | GRNC_gma_lcl|Gmax_Glyma.19G136600.8 | Soybean (N↑) | [118] | |
| gma_eTM_miR169g-2 | GRNC_gma_lcl|Gmax_Glyma.19G136600.3 | ||||
| gma_eTM_miR169g-1 | GRNC_gma_lcl|Gmax_Glyma.19G136600.2 | ||||
| ath-miR169a-3p | ath_eTM_miR169a-3p-2 | GRNC_ath_lcl|Athaliana_AT1G44940.2 | A. thaliana(P↓) | [113] | |
| ath_eTM_miR169a-3p-1 | GRNC_ath_lcl|Athaliana_AT1G44940.1 | ||||
| zma-miR319a-5p | zma_eTM_miR319a-5p-4 | GRNC_zma_lcl|Zmays_GRMZM2G438722_T03 | Maize (N↑) | [116] | |
| zma_eTM_miR319a-5p-3 | zma_TCONS_00089764 | ||||
| zma_eTM_miR319a-5p-2 | zma_TCONS_00089763 | ||||
| zma_eTM_miR319a-5p-1 | zma_TCONS_00024738 | ||||
| zma-miR395d-5p | zma_eTM_miR395d-5p-1 | zma_TCONS_00091080 | Maize (N↓) | [116] | |
| zma-miR395g-5p | zma_eTM_miR395g-5p-1 | zma_TCONS_00091080 | |||
| gma-miR398b | gma_eTM_miR398b-1 | GRNC_gma_lcl|Gmax_Glyma.12G204100.1 | Soybean (P↓) | [114] | |
| ath-miR399f | ath_eTM_miR399f-5 | GRNC_ath_lcl|Athaliana_AT5G03545.1 | A. thaliana (N↑↓) | A. thaliana (P↑) | [113] |
| ath_eTM_miR399f-4 | GRNC_ath_lcl|Athaliana_AT3G09922.1 | ||||
| ath_eTM_miR399f-3 | PNRD_ath_At4-2 | ||||
| ath-miR399e | ath_eTM_miR399f-2 | PNRD_ath_At4 | |||
| ath_eTM_miR399f-1 | PNRD_ath_AtIPS1 | ||||
| ath_eTM_miR399e-1 | GRNC_ath_lcl|Athaliana_AT1G53708.1 | A. thaliana (N↑↓) | A. thaliana (P↑) | [113] | |
| ath-miR399d | ath_eTM_miR399d-5 | GRNC_ath_lcl|Athaliana_AT5G03545.1 | A. thaliana (N↑↓) | A. thaliana (P↑) | [113] |
| ath_eTM_miR399d-4 | GRNC_ath_lcl|Athaliana_AT3G09922.1 | ||||
| ath_eTM_miR399d-3 | PNRD_ath_At4-2 | ||||
| ath-miR399b | ath_eTM_miR399d-2 | PNRD_ath_At4 | |||
| ath_eTM_miR399d-1 | PNRD_ath_AtIPS1 | ||||
| ath_eTM_miR399b-6 | GRNC_ath_lcl|Athaliana_AT5G03545.1 | A. thaliana (N↑↓) | A. thaliana (P↑) | [113] | |
| ath_eTM_miR399b-5 | GRNC_ath_lcl|Athaliana_AT3G09922.1 | ||||
| ath_eTM_miR399b-4 | GRNC_ath_lcl|Athaliana_AT1G53708.1 | ||||
| ath-miR399a | ath_eTM_miR399b-3 | PNRD_ath_At4-2 | |||
| ath_eTM_miR399b-2 | PNRD_ath_At4 | ||||
| ath_eTM_miR399b-1 | PNRD_ath_AtIPS1 | ||||
| ath_eTM_miR399a-6 | GRNC_ath_lcl|Athaliana_AT5G03545.1 | A. thaliana (N↑↓) | A. thaliana (P↑) | [113] | |
| ath_eTM_miR399a-5 | GRNC_ath_lcl|Athaliana_AT3G09922.1 | ||||
| ath_eTM_miR399a-4 | GRNC_ath_lcl|Athaliana_AT1G53708.1 | ||||
| ath_eTM_miR399a-3 | PNRD_ath_At4-2 | ||||
| ath_eTM_miR399a-2 | PNRD_ath_At4 | ||||
| ath_eTM_miR399a-1 | PNRD_ath_AtIPS1 | ||||
| tae-miR408 | tae_eTM_miR408-2 | GRNC_tae_lcl|Taestivum_Traes_5BL_EED36D3B9.35 | Wheat (P↓) | [115] | |
| tae_eTM_miR408-1 | tae_TCONS_00103503 | ||||
| osa-miR444a-3p.2 | osa_eTM_miR444a-3p.2-1 | GRNC_osa_lcl|Osativa_LOC_Os12g19080.1 | Rice((N↑) | [119] | |
| ath-miR827 | ath_eTM_miR827-2 | GRNC_ath_lcl|Athaliana_AT3G02832.1 gene = AT3G02832 | A. thaliana (P↑) | [112] | |
| ath_eTM_miR827-1 | PNRD_ath_NONATHT001723 | ||||
| ath-miR828 | ath_eTM_miR828-1 | PNRD_ath_NONATHT000094 | A. thaliana (P↑) | [112] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Borah, P.; Das, A.; Milner, M.J.; Ali, A.; Bentley, A.R.; Pandey, R. Long Non-Coding RNAs as Endogenous Target Mimics and Exploration of Their Role in Low Nutrient Stress Tolerance in Plants. Genes 2018, 9, 459. https://doi.org/10.3390/genes9090459
Borah P, Das A, Milner MJ, Ali A, Bentley AR, Pandey R. Long Non-Coding RNAs as Endogenous Target Mimics and Exploration of Their Role in Low Nutrient Stress Tolerance in Plants. Genes. 2018; 9(9):459. https://doi.org/10.3390/genes9090459
Chicago/Turabian StyleBorah, Priyanka, Antara Das, Matthew J. Milner, Arif Ali, Alison R. Bentley, and Renu Pandey. 2018. "Long Non-Coding RNAs as Endogenous Target Mimics and Exploration of Their Role in Low Nutrient Stress Tolerance in Plants" Genes 9, no. 9: 459. https://doi.org/10.3390/genes9090459





