How Next-Generation Sequencing Has Aided Our Understanding of the Sequence Composition and Origin of B Chromosomes
Abstract
:1. Recent Discoveries Related to the Origin and Evolution of B Chromosomes
2. The Acquisition of Sequences Enriched in B Chromosome
3. The In Silico-Based Identification of B Chromosome-Enriched Sequences
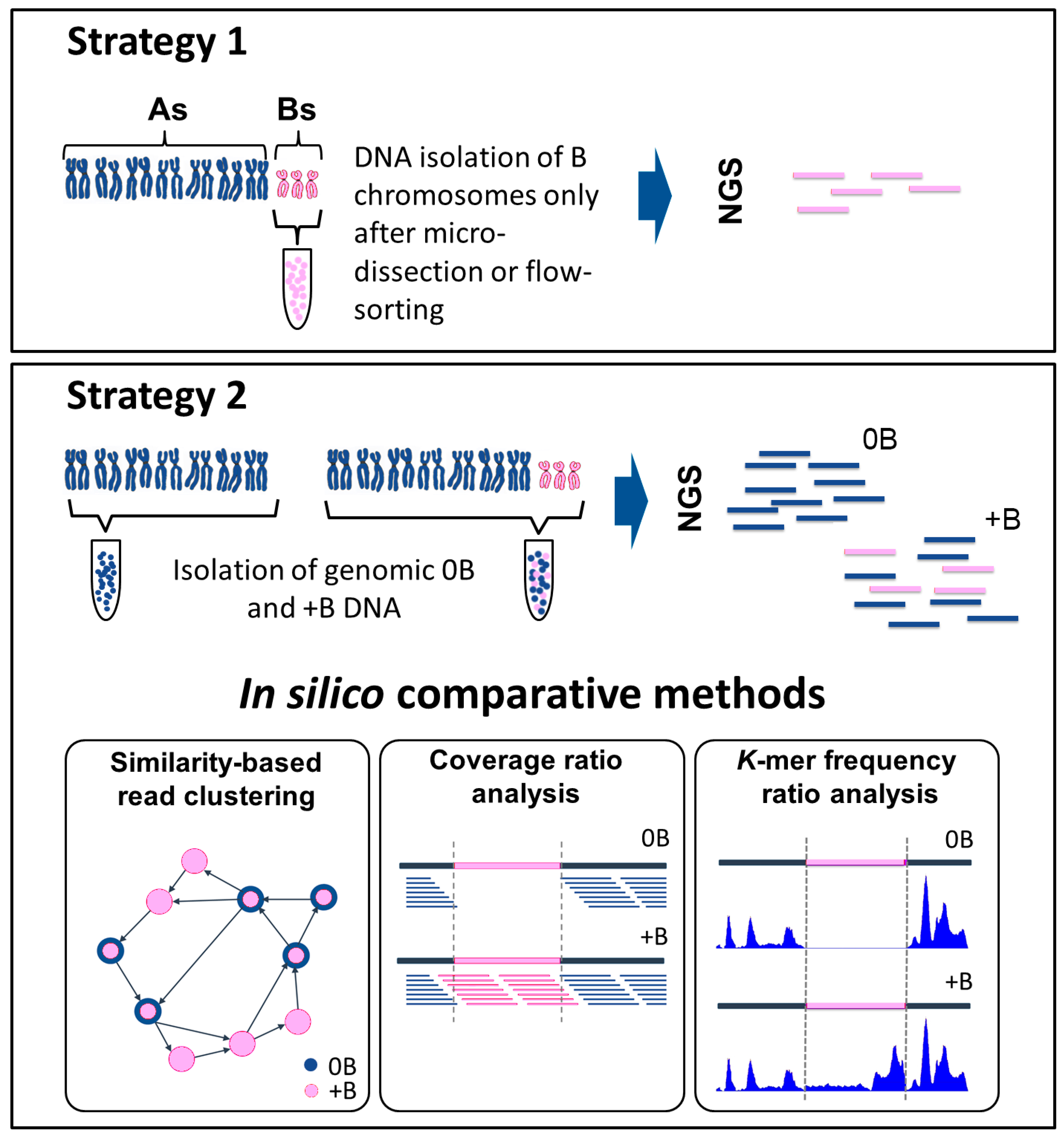
3.1. Strategy 1—The Direct Route: Isolating, then Sequencing Micro-Dissected or Flow-Sorted B Chromosomes
3.2. Strategy 2—The Indirect Route: Comparing Whole Genome Sequence Acquired from Individuals Carrying and Not Carrying B Chromosomes
3.2.1. Similarity-Based Read Clustering
3.2.2. Coverage Ratio Analysis
3.2.3. k-mer Frequency Ratio Analysis
3.3. Benefits and Merits of Indirect and Direct Strategies
3.4. An In Silico Method Used to Identify B Chromosome Sequences
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Jones, R.N. B-chromosome drive. Am. Nat. 1991, 137, 430–442. [Google Scholar] [CrossRef]
- Houben, A. B chromosomes—A matter of chromosome drive. Front. Plant Sci. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Martis, M.M.; Klemme, S.; Banaei-Moghaddam, A.M.; Blattner, F.R.; Macas, J.; Schmutzer, T.; Scholz, U.; Gundlach, H.; Wicker, T.; Simkova, H.; et al. Selfish supernumerary chromosome reveals its origin as a mosaic of host genome and organellar sequences. Proc. Natl. Acad. Sci. USA 2012, 109, 13343–13346. [Google Scholar] [CrossRef] [PubMed]
- Klemme, S.; Banaei-Moghaddam, A.M.; Macas, J.; Wicker, T.; Novák, P.; Houben, A. High-copy sequences reveal distinct evolution of the rye B-chromosome. New Phytol. 2013, 199, 550–558. [Google Scholar] [CrossRef] [PubMed]
- Ruban, A.; Fuchs, J.; Marques, A.; Schubert, V.; Soloviev, A.; Raskina, O.; Badaeva, E.; Houben, A. B chromosomes of Aegilops speltoides are enriched in organelle genome-derived sequences. PLoS ONE 2014, 9, e90214. [Google Scholar] [CrossRef] [PubMed]
- Marques, A.; Klemme, S.; Guerra, M.; Houben, A. Cytomolecular characterization of de novo formed rye B chromosome variants. Mol. Cytogenet. 2012, 5. [Google Scholar] [CrossRef] [PubMed]
- Banaei-Moghaddam, A.M.; Meier, K.; Karimi-Ashtiyani, R.; Houben, A. Formation and expression of pseudogenes on the B chromosome of rye. Plant Cell 2013, 25, 2536–2544. [Google Scholar] [CrossRef] [PubMed]
- Ma, W.; Gabriel, T.S.; Martis, M.M.; Gursinsky, T.; Schubert, V.; Vrána, J.; Doležel, J.; Grundlach, H.; Altschmied, L.; Scholz, U.; et al. Rye B chromosomes encode a functional Argonaute-like protein with in vitro slicer activities similar to its A chromosome paralog. New Phytol. 2016, 213, 916–928. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Liu, X.; Benny, U.; Kistler, H.C.; VanEtten, H.D. Genes determining pathogenicity to pea are clustered on a supernumerary chromosome in the fungal plant pathogen Nectria haematococca. Plant J. 2001, 25, 305–314. [Google Scholar] [CrossRef] [PubMed]
- Harimoto, Y.; Hatta, R.; Kodama, M.; Yamamoto, M.; Otani, H.; Tsuge, T. Expression profiles of genes encoded by the supernumerary chromosome controlling AM-toxin biosynthesis and pathogenicity in the apple pathotype of Alternaria alternata. Mol. Plant Microbe Interact. 2007, 20, 1463–1476. [Google Scholar] [CrossRef] [PubMed]
- Vanheule, A.; Audenaert, K.; Warris, S.; van de Geest, H.; Schijlen, E.; Höfte, M.; De Saeger, S.; Haesaert, G.; Waalwijk, C.; van der Lee, T. Living apart together: Crosstalk between the core and supernumerary genomes in a fungal plant pathogen. BMC Genom. 2016, 17, 670. [Google Scholar] [CrossRef] [PubMed]
- Coleman, J.J.; Rounsley, S.D.; Rodriguez-Carres, M.; Kuo, A.; Wasmann, C.C.; Grimwood, J.; Schmutz, J.; Taga, M.; White, G.J.; Zhou, S.; et al. The genome of Nectaria haematococca: Contribution of supernumerary chromosomes to gene expansion. PLoS Genet. 2009, 5, e1000618. [Google Scholar] [CrossRef] [PubMed]
- Croll, D.; McDonald, B.A. The accessory genome as a cradle for adaptive evolution in pathogens. PLoS Pathog. 2012, 8, e1002608. [Google Scholar] [CrossRef] [PubMed]
- Navarro-Domínguez, B.; Ruiz-Ruano, F.J.; Cabrero, J.; Corral, J.M.; López-León, M.D.; Sharbel, T.F.; Camacho, J.P.M. Protein-coding genes in B chromosomes of the grasshopper Eyprepocnemis plorans. Sci. Rep. 2017, 7, 45200. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Ruano, F.J.; Cabrero, J.; Lopez-Leon, M.D.; Camacho, J.P. Satellite DNA content illuminates the ancestry of a supernumerary (B) chromosome. Chromosoma 2017, 126, 487–500. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Ruano, F.J.; Cabrero, J.; Lopez-Leon, M.D.; Sanchez, A.; Camacho, J.P.M. Quantitative sequence characterization for repetitive DNA content in the supernumerary chromosome of the migratory locust. Chromosoma 2017. [Google Scholar] [CrossRef] [PubMed]
- Akbari, O.S.; Antoshechkin, I.; Hay, B.A.; Ferree, P.M. Transcriptome profiling of Nasonia vitripennis testis reveals novel transcripts expressed from the selfish B chromosome, paternal sex ratio. G3 2013, 3, 1597–1605. [Google Scholar] [CrossRef] [PubMed]
- Nur, U.; Werren, J.H.; Eickbush, D.G.; Burke, W.D.; Eickbush, T.H. A “selfish” B chromosome that enhances its transmission by eliminating the paternal genome. Science 1988, 240, 512–514. [Google Scholar] [CrossRef] [PubMed]
- Aldrich, J.C.; Ferree, P.M. Genome silencing and elimination: Insights from a “selfish” B chromosome. Front. Genet. 2017, 8, 50. [Google Scholar] [CrossRef] [PubMed]
- Aldrich, J.C.; Leibholz, A.; Cheema, M.S.; Ausiό, J.; Ferree, P.M. A ‘selfish’ B chromosome induces genome elimination by disrupting the histone code in the jewel wasp Nasonia vitripennis. Sci. Rep. 2017, 7, 42551. [Google Scholar] [CrossRef] [PubMed]
- Bauerly, E.; Hughes, S.E.; Vietti, D.R.; Miller, D.E.; McDowell, W.; Hawley, R.S. Discovery of supernumerary B chromosomes in Drosophila melanogaster. Genetics 2014, 196, 1007–1016. [Google Scholar] [CrossRef] [PubMed]
- Valente, G.T.; Conte, M.A.; Fantinatti, B.E.A.; Cabral-de-Mello, D.C.; Carvalho, R.F.; Vicari, M.R.; Kocher, T.D.; Martins, C. Origin and evolution of B chromosomes in the cichlid fish Astatotilapia latifasciata based on integrated genomic analyses. Mol. Biol. Evol. 2014, 31, 2061–2072. [Google Scholar] [CrossRef] [PubMed]
- Clark, F.E.; Conte, M.A.; Ferreira-Bravo, I.A.; Poletto, A.B.; Martins, C.; Kocher, T.D. Dynamic sequence evolution of a sex-associated B chromosome in Lake Malawi cichlid fish. J. Hered. 2017, 108, 53–62. [Google Scholar] [CrossRef] [PubMed]
- Camacho, J.P.M.; Schmid, M.; Cabrero, J. B chromosomes and sex in animals. Sex. Dev. 2011, 5, 155–166. [Google Scholar] [CrossRef] [PubMed]
- Becker, S.E.; Thomas, R.; Trifonov, V.A.; Wayne, R.K.; Graphodatsky, A.S.; Breen, M. Anchoring the dog to its relatives reveals new evolutionary breakpoints across 11 species of the Canidae and provides new clues for the role of B chromosomes. Chromosom. Res. 2011, 19, 685–708. [Google Scholar] [CrossRef] [PubMed]
- Graphodatsky, A.S.; Kukekova, A.V.; Yudkin, D.V.; Trifonov, V.A.; Vorobieva, N.V.; Beklemisheva, V.R.; Perelman, P.L.; Graphodatskaya, D.A.; Trut, L.N.; Yang, F.; et al. The proto-oncogene C-KIT maps to canid B-chromosomes. Chromosom. Res. 2005, 13, 113–122. [Google Scholar] [CrossRef] [PubMed]
- Yudkin, D.V.; Trifonov, V.A.; Kukekova, A.V.; Vorobieva, N.V.; Rubtsova, N.V.; Yang, F.; Acland, G.M.; Ferguson-Smith, M.A.; Graphodatsky, A.S. Mapping of KIT adjacent sequences on canid autosomes and B chromosomes. Cytogenet. Genome Res. 2007, 116, 100–103. [Google Scholar] [CrossRef] [PubMed]
- Makunin, A.I.; Kichigin, I.G.; Larkin, D.M.; O’Brien, P.C.; Ferguson-Smith, M.A.; Yang, F.; Proskuryakova, A.A.; Vorobieva, N.V.; Chernyaeva, E.N.; O’Brien, S.J.; et al. Contrasting origin of B chromosomes in two cervids (Siberian roe deer and grey brocket deer) unravelled by chromosome-specific DNA sequencing. BMC Genom. 2016, 17, 618. [Google Scholar] [CrossRef] [PubMed]
- Ventura, K.; O’Brien, P.C.M.; do Nascimento Moreira, C.; Yonenaga-Yassuda, Y.; Ferguson-Smith, M.A. On the origin and evolution of the extant system of B chromosomes in Oryzomyini radiation (Rodentia, Sigmodontinae). PLoS ONE 2015, 10, e0136663. [Google Scholar] [CrossRef] [PubMed]
- Rubtsov, N.B.; Karamysheva, T.V.; Andreenkova, O.V.; Bochkaerev, M.N.; Kartavtseva, I.V.; Roslik, G.V.; Borissov, Y.M. Comparative analysis of micro and macro B chromosomes in the Korean field mouse Apodemus peninsulae (Rodentia, Murinae) performed by chromosome microdissection and FISH. Cytogenet. Genome Res. 2004, 106, 289–294. [Google Scholar] [CrossRef] [PubMed]
- Sharbel, T.F.; Green, D.M.; Houben, A. B-chromosome origin in the endemic New Zealand frog Leiopelma hochstetteri through sex chromosome devolution. Genome 1998, 41, 14–22. [Google Scholar] [CrossRef] [PubMed]
- Sandery, M.J.; Forster, J.W.; Macadam, S.R.; Blunden, R.; Jones, R.N.; Brown, D.M. Isolation of a sequence common to A- and B-chromosomes of rye (Secale cereale) by microcloning. Plant Mol. Biol. Rep. 1991, 9, 21–30. [Google Scholar] [CrossRef]
- Houben, A. Chromosome microdissection and utilization of microisolated DNA. In Plant Cytogenetics. Plant Genetics and Genomics: Crops and Models; Bass, H., Birchler, J., Eds.; Springer: New York, NY, USA, 2011; Volume 4, pp. 257–270. [Google Scholar]
- Kosyakova, N.; Liehr, T.; Al-Rikabi, H.A.B. FISH-microdissection. In Fluorescence In Situ Hybridization (FISH). Application Guide, 2nd ed.; Liehr, T., Ed.; Springer: Berlin/Heidelberg, Germany, 2017; pp. 81–100. [Google Scholar]
- Zhang, Y.X.; Deng, C.L.; Hu, Z.M. The chromosome microdissection and microcloning technique. Methods Mol. Biol. 2016, 1429, 151–160. [Google Scholar] [CrossRef] [PubMed]
- Zhou, R.N.; Hu, Z.M. The development of chromosome microdissection and microcloning technique and its applications in genomic research. Curr. Genom. 2007, 8, 67–72. [Google Scholar] [CrossRef]
- Houben, A.; Kynast, R.G.; Heim, U.; Hermann, H.; Jones, R.N.; Forster, J.W. Molecular cytogenetic characterisation of the terminal heterochromatic segment of the B-chromosome of rye (Secale cereale). Chromosoma 1996, 105, 97–103. [Google Scholar] [CrossRef] [PubMed]
- Houben, A.; Leach, C.R.; Verlin, D.; Rofe, R.; Timmis, J.N. A repetitive DNA sequence common to the different B chromosomes of the genus Brachycome. Chromosoma 1997, 106, 513–519. [Google Scholar] [CrossRef] [PubMed]
- Dolezel, J.; Kubalakova, M.; Paux, E.; Bartos, J.; Feuillet, C. Chromosome-based genomics in the cereals. Chromosom. Res. 2007, 15, 51–66. [Google Scholar] [CrossRef] [PubMed]
- Telenius, H.; Carter, N.P.; Bebb, C.E.; Nordenskjold, M.; Ponder, B.A.J.; Tunnacliffe, A. Degenerate oligonucleotide-primed PCR—General amplification of target DNA by a single degenerate primer. Genomics 1992, 13, 718–725. [Google Scholar] [CrossRef]
- Dean, F.B.; Nelson, J.R.; Giesler, T.L.; Lasken, R.S. Rapid amplification of plasmid and phage DNA using Phi29 DNA polymerase and multiply-primed rolling circle amplification. Genome Res. 2001, 11, 1095–1099. [Google Scholar] [CrossRef] [PubMed]
- Šimková, H.; Svensson, J.T.; Condamine, P.; Hřibová, E.; Suchánková, P.; Bhat, P.R.; Bartoš, J.; Šafář, J.; Close, T.J.; Doležel, J. Coupling amplified DNA from flow-sorted chromosomes to high-density SNP mapping in barley. BMC Genom. 2008, 9, 294. [Google Scholar] [CrossRef] [PubMed]
- Long, H.; Qi, Z.X.; Sun, X.M.; Chen, C.B.; Li, X.L.; Song, W.Q.; Chen, R.Y. Characters of DNA constitution in the rye B chromosome. J. Integr. Plant Biol. 2008, 50, 183–189. [Google Scholar] [CrossRef] [PubMed]
- Jamilena, M.; Garrido-Ramos, M.; Rejon, M.R.; Rejon, C.R.; Parker, J.S. Characterisation of repeated sequences from microdissected B chromosomes of Crepis capillaris. Chromosoma 1995, 104, 113–120. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.M.; Lin, B.Y. Cloning and characterization of maize B chromosome sequences derived from microdissection. Genetics 2003, 164, 299–310. [Google Scholar] [PubMed]
- Amorim, I.C.; Milani, D.; Cabral-de-Mello, D.C.; Rocha, M.F.; Moura, R.C. Possible origin of B chromosome in Dichotomius sericeus (Coleoptera). Genome 2016, 59, 575–580. [Google Scholar] [CrossRef] [PubMed]
- Teruel, M.; Cabrero, J.; Montiel, E.E.; Acosta, M.J.; Sanchez, A.; Camacho, J.P. Microdissection and chromosome painting of X and B chromosomes in Locusta migratoria. Chromosom. Res. 2009, 17, 11–18. [Google Scholar] [CrossRef] [PubMed]
- Bugrov, A.G.; Karamysheva, T.V.; Perepelov, E.A.; Elisaphenko, E.A.; Rubtsov, D.N.; Warchalowska-Sliwa, E.; Tatsuta, H.; Rubtsov, N.B. DNA content of the B chromosomes in grasshopper Podisma kanoi Storozh. (Orthoptera, Acrididae). Chromosom. Res. 2007, 15, 315–325. [Google Scholar] [CrossRef] [PubMed]
- Bugrov, A.G.; Karamysheva, T.V.; Pyatkova, M.S.; Rubtsov, D.N.; Andreenkova, O.V.; Warchalowska-Sliwa, E.; Rubtsov, N.B. B chromosomes of the Podisma sapporensis Shir. (Orthoptera, Acrididae) analysed by chromosome microdissection and FISH. Folia Biol. 2003, 51, 1–11. [Google Scholar]
- Menezes-de-Carvalho, N.Z.; Palacios-Gimenez, O.M.; Milani, D.; Cabral-de-Mello, D.C. High similarity of U2 snDNA sequence between A and B chromosomes in the grasshopper Abracris flavolineata. Mol. Genet. Genom. 2015, 290, 1787–1792. [Google Scholar] [CrossRef] [PubMed]
- Gruber, S.; Diniz, D.; Sobrinho-Scudeler, P.; Foresti, F.; Haddad, C.; Kasahara, S. Possible interspecific origin of the B chromosome of Hypsiboas albopunctatus (Spix, 1824) (Anura, Hylidae), revealed by microdissection, chromosome painting, and reverse hybridisation. Comp. Cytogenet. 2014, 8, 17–29. [Google Scholar] [CrossRef] [PubMed]
- Scudeler, P.E.; Diniz, D.; Wasko, A.P.; Oliveira, C.; Foresti, F. Whole chromosome painting of B chromosomes of the red-eye tetra Moenkhausia sanctaefilomenae (Teleostei, Characidae). Comp. Cytogenet. 2015, 9, 661–669. [Google Scholar] [CrossRef] [PubMed]
- Vicari, M.R.; de Mello Pistune, H.F.; Castro, J.P.; de Almeida, M.C.; Bertollo, L.A.; Moreira-Filho, O.; Camacho, J.P.; Artoni, R.F. New insights on the origin of B chromosomes in Astyanax scabripinnis obtained by chromosome painting and FISH. Genetica 2011, 139, 1073–1081. [Google Scholar] [CrossRef] [PubMed]
- De A. Silva, D.M.; Daniel, S.N.; Camacho, J.P.; Utsunomia, R.; Ruiz-Ruano, F.J.; Penitente, M.; Pansonato-Alves, J.C.; Hashimoto, D.T.; Oliveira, C.; Porto-Foresti, F.; et al. Origin of B chromosomes in the genus Astyanax (Characiformes, Characidae) and the limits of chromosome painting. Mol. Genet. Genom. 2016, 291, 1407–1418. [Google Scholar] [CrossRef] [PubMed]
- Voltolin, T.A.; Laudicina, A.; Senhorini, J.A.; Bortolozzi, J.; Oliveira, C.; Foresti, F.; Porto-Foresti, F. Origin and molecular organization of supernumerary chromosomes of Prochilodus lineatus (Characiformes, Prochilodontidae) obtained by DNA probes. Genetica 2010, 138, 1133–1139. [Google Scholar] [CrossRef] [PubMed]
- Brinkman, J.N.; Sessions, S.K.; Houben, A.; Green, D.M. Structure and evolution of supernumerary chromosomes in the Pacific giant salamander, Dicamptodon tenebrosus. Chromosom. Res. 2000, 8, 477–485. [Google Scholar] [CrossRef]
- Karamysheva, T.V.; Andreenkova, O.V.; Bochkaerev, M.N.; Borissov, Y.M.; Bogdanchikova, N.; Borodin, P.M.; Rubtsov, N.B. B chromosomes of Korean field mouse Apodemus peninsulae (Rodentia, Murinae) analysed by microdissection and FISH. Cytogenet. Genome Res. 2002, 96, 154–160. [Google Scholar] [CrossRef] [PubMed]
- Thind, A.K.; Wicker, T.; Simkova, H.; Fossati, D.; Moullet, O.; Brabant, C.; Vrana, J.; Dolezel, J. Rapid cloning of genes in hexaploid wheat using cultivar-specific long-range chromosome assembly. Nat. Biotechnol. 2017, 35, 793–796. [Google Scholar] [CrossRef] [PubMed]
- Putnam, N.H.; O’Connell, B.L.; Stites, J.C.; Rice, B.J.; Blanchette, M.; Calef, R.; Troll, C.J.; Fields, A.; Hartley, P.D.; Sugnet, C.W.; et al. Chromosome-scale shotgun assembly using an in vitro method for long-range linkage. Genome Res. 2016, 26, 342–350. [Google Scholar] [CrossRef] [PubMed]
- Dolezel, J.; Vrana, J.; Capal, P.; Kubalakova, M.; Buresova, V.; Simkova, H. Advances in plant chromosome genomics. Biotechnol. Adv. 2014, 32, 122–136. [Google Scholar] [CrossRef] [PubMed]
- Novák, P.; Neumann, P.; Macas, J. Graph-based clustering and characterization of repetitive sequences in next-generation sequencing data. BMC Bioinform. 2010, 11. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Kumke, K.; Macas, J.; Fuchs, J.; Altschmied, L.; Kour, J.; Dhar, M.K.; Houben, A. Plantago lagopus B chromosome is enriched in 5S rDNA-derived satellite DNA. Cytogenet. Genome Res. 2016, 148, 68–73. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009, 25, 1754–1760. [Google Scholar] [CrossRef] [PubMed]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed]
- Kurtz, S.; Narechania, A.; Stein, J.C.; Ware, D. A new method to compute k-mer frequencies and its application to annotate large repetitive plant genomes. BMC Genom. 2008, 9, 517. [Google Scholar] [CrossRef] [PubMed]
- Marçais, G.; Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 2011, 27, 764–770. [Google Scholar] [CrossRef] [PubMed]
- Schmutzer, T.; Ma, L.; Pousarebani, N.; Bull, F.; Stein, N.; Houben, A.; Scholz, U. Kmasker—A tool for in silico prediction of single-copy FISH probes for the large-genome species Hordeum vulgare. Cytogenet. Genome Res. 2014, 142, 66–78. [Google Scholar] [CrossRef] [PubMed]
- Aliyeva-Schnorr, L.; Beier, S.; Karafiátová, M.; Schmutzer, T.; Scholz, U.; Doležel, J.; Stein, N.; Houben, A. Cytogenetic mapping with centromeric bacterial artificial chromosomes contigs shows that this recombination-poor region comprises more than half of barley chromosome 3H. Plant J. 2015, 84, 385–394. [Google Scholar] [CrossRef] [PubMed]
- Vu, G.T.H.; Schmutzer, T.; Bull, F.; Cao, H.X.; Fuchs, J.; Tran, T.D.; Jovtchev, G.; Pistrick, K.; Stein, N.; Pecinka, A.; et al. Comparative genome analysis reveals divergent genome size evolution in a carnivorous plant genus. Plant Genome 2015, 8. [Google Scholar] [CrossRef]
- Mayer, K.F.; Martis, M.; Hedley, P.E.; Simkova, H.; Liu, H.; Morris, J.A.; Steuernagel, B.; Taudien, S.; Roessner, S.; Gundlach, H.; et al. Unlocking the barley genome by chromosomal and comparative genomics. Plant Cell 2011, 23, 1249–1263. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Houben, A.; Banaei-Moghaddam, A.M.; Klemme, S.; Timmis, J.N. Evolution and biology of supernumerary B chromosomes. Cell. Mol. Life Sci. 2014, 71, 467–478. [Google Scholar] [CrossRef] [PubMed]
- Banaei-Moghaddam, A.M.; Martis, M.M.; Macas, J.; Gundlach, H.; Himmelbach, A.; Altschmied, L.; Mayer, K.F.; Houben, A. Genes on B chromosomes: Old questions revisited with new tools. Biochim. Biophys. Acta 2015, 1849, 64–70. [Google Scholar] [CrossRef] [PubMed]
- Valente, G.T.; Nakajima, R.T.; Fantinatti, B.E.; Marques, D.F.; Almeida, R.O.; Simoes, R.P.; Martins, C. B chromosomes: From cytogenetics to systems biology. Chromosoma 2017, 126, 73–81. [Google Scholar] [CrossRef] [PubMed]
- Makunin, A.; Dementyeva, P.; Graphodatsky, A.; Volobouev, V.; Kukekova, A.; Trifonov, V. Genes on B chromosomes of vertebrates. Mol. Cytogenet. 2014, 7, 99. [Google Scholar] [CrossRef] [PubMed]
- Leibowitz, M.L.; Zhang, C.Z.; Pellman, D. Chromothripsis: A new mechanism for rapid karyotype evolution. Annu. Rev. Genet. 2015, 49, 183–211. [Google Scholar] [CrossRef] [PubMed]
- Liehr, T.; Mrasek, K.; Kosyakova, N.; Ogilvie, C.M.; Vermeesch, J.; Trifonov, V.; Rubtsov, N. Small supernumerary marker chromosomes (sSMC) in humans; are there B chromosomes hidden among them. Mol. Cytogenet. 2008, 1, 12. [Google Scholar] [CrossRef] [PubMed]
| Species | Reference |
|---|---|
| Rye (Secale cereale) | [32,37,43] |
| Hawks beard (Crepis capillaris) | [44] |
| Brachycome dichromosomatica | [38] |
| Maize (Zea mays) | [45] |
| Dung beetle (Dichotomius sericeus) | [46] |
| Locust (Locusta migratoria) | [16,47] |
| Grasshopper (Podisma kanoi) | [48] |
| Grasshopper (Podisma sapporensis) | [49] |
| Grasshopper (Abracris flavolineata) | [50] |
| Hylid frog (Hypsiboas albopunctatus) | [51] |
| Fish (Moenkhausia sanctaefilomenae) | [52] |
| Fish (Astyanax scabripinnis) | [53] |
| Cichlid fish (Astatotilapia latifasciata) | [22] |
| Fishes genus Astyanax: A. paranae, A. fasciatus, A. bockmanni | [54] |
| Fish (Prochilodus lineatus) | [55] |
| Pacific giant salamander (Dicamptodon tenebrosus) | [56] |
| Korean field mouse (Apodemus peninsulae) | [57] |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ruban, A.; Schmutzer, T.; Scholz, U.; Houben, A. How Next-Generation Sequencing Has Aided Our Understanding of the Sequence Composition and Origin of B Chromosomes. Genes 2017, 8, 294. https://doi.org/10.3390/genes8110294
Ruban A, Schmutzer T, Scholz U, Houben A. How Next-Generation Sequencing Has Aided Our Understanding of the Sequence Composition and Origin of B Chromosomes. Genes. 2017; 8(11):294. https://doi.org/10.3390/genes8110294
Chicago/Turabian StyleRuban, Alevtina, Thomas Schmutzer, Uwe Scholz, and Andreas Houben. 2017. "How Next-Generation Sequencing Has Aided Our Understanding of the Sequence Composition and Origin of B Chromosomes" Genes 8, no. 11: 294. https://doi.org/10.3390/genes8110294





