Yeast Irc22 Is a Novel Dsk2-Interacting Protein that Is Involved in Salt Tolerance
Abstract
:1. Introduction
2. Experimental Section
2.1. Strains and Media
2.2. Plasmids and Mutants
2.3. Pull-Down Assays and Immunoblotting
2.4. Degradation Assay of N-End Rule Substrate
2.5. Growth Sensitivity to Salt Stress, UV Irradiation and Oxidative Stress
2.6. Antibodies
3. Results and Discussion
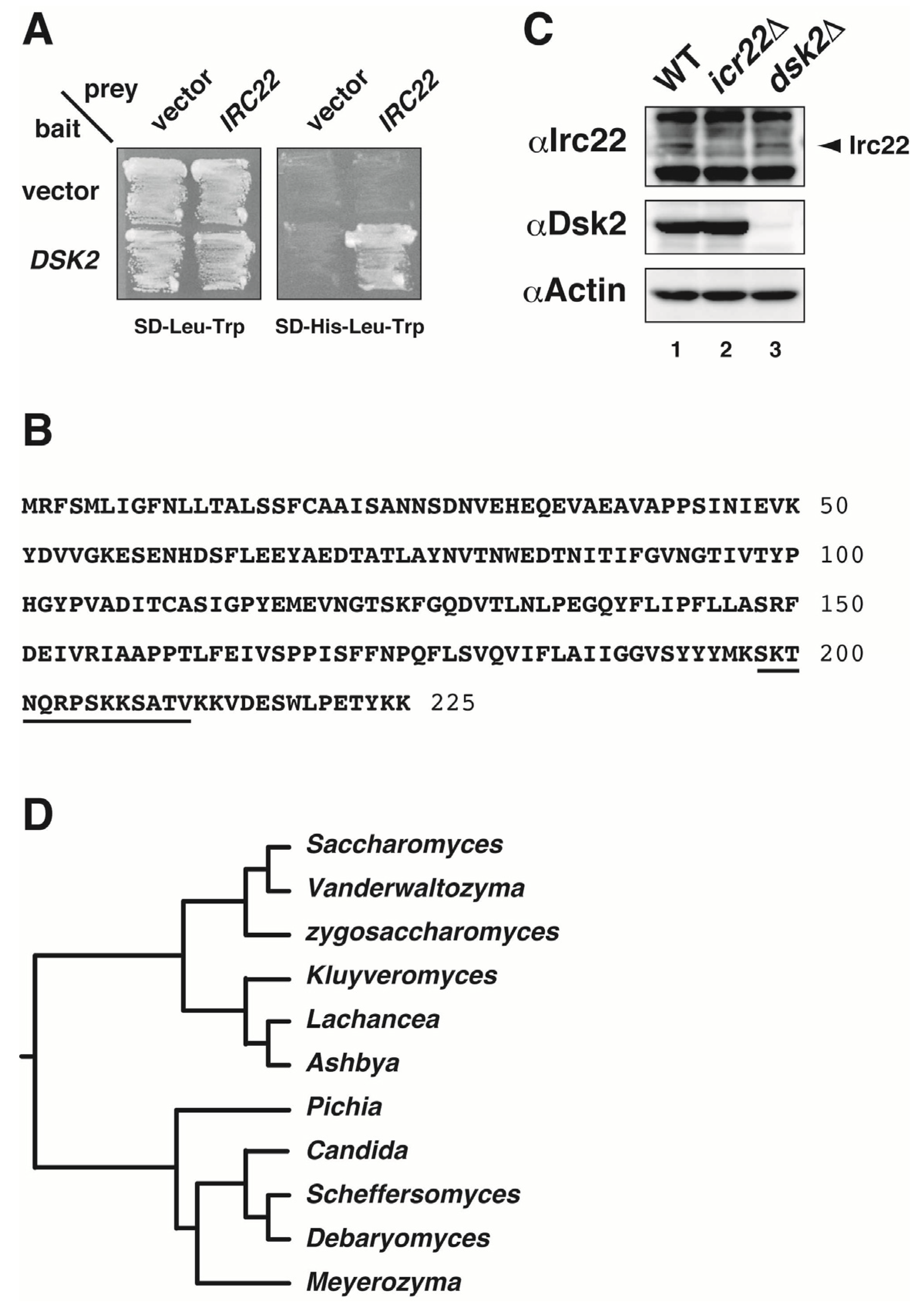
3.1. Identification of Irc22 as a Novel Dsk2-Interacting Protein in Yeast
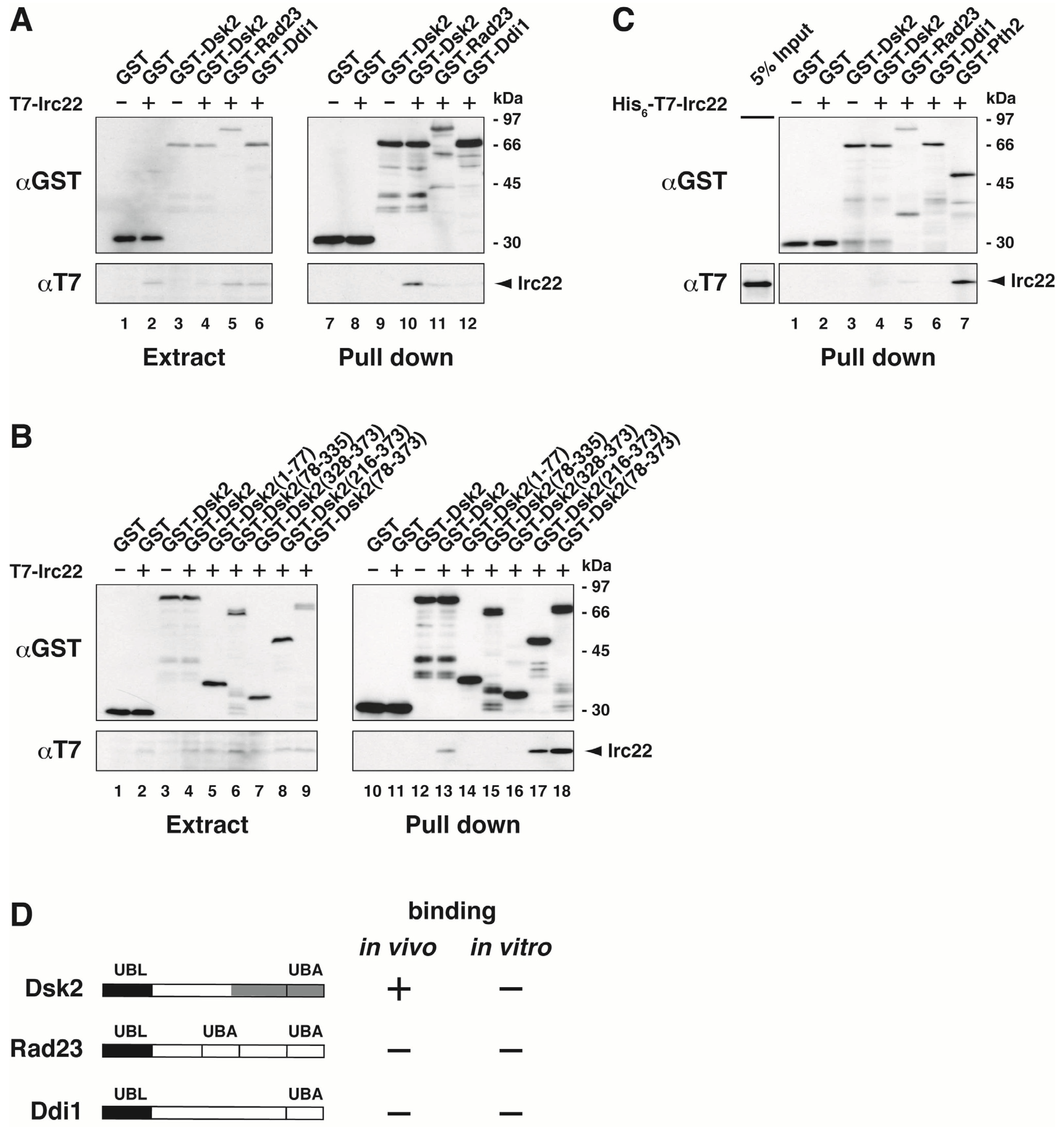
3.2. Irc22 Interacts with Dsk2 But Not with Rad23 and Ddi1
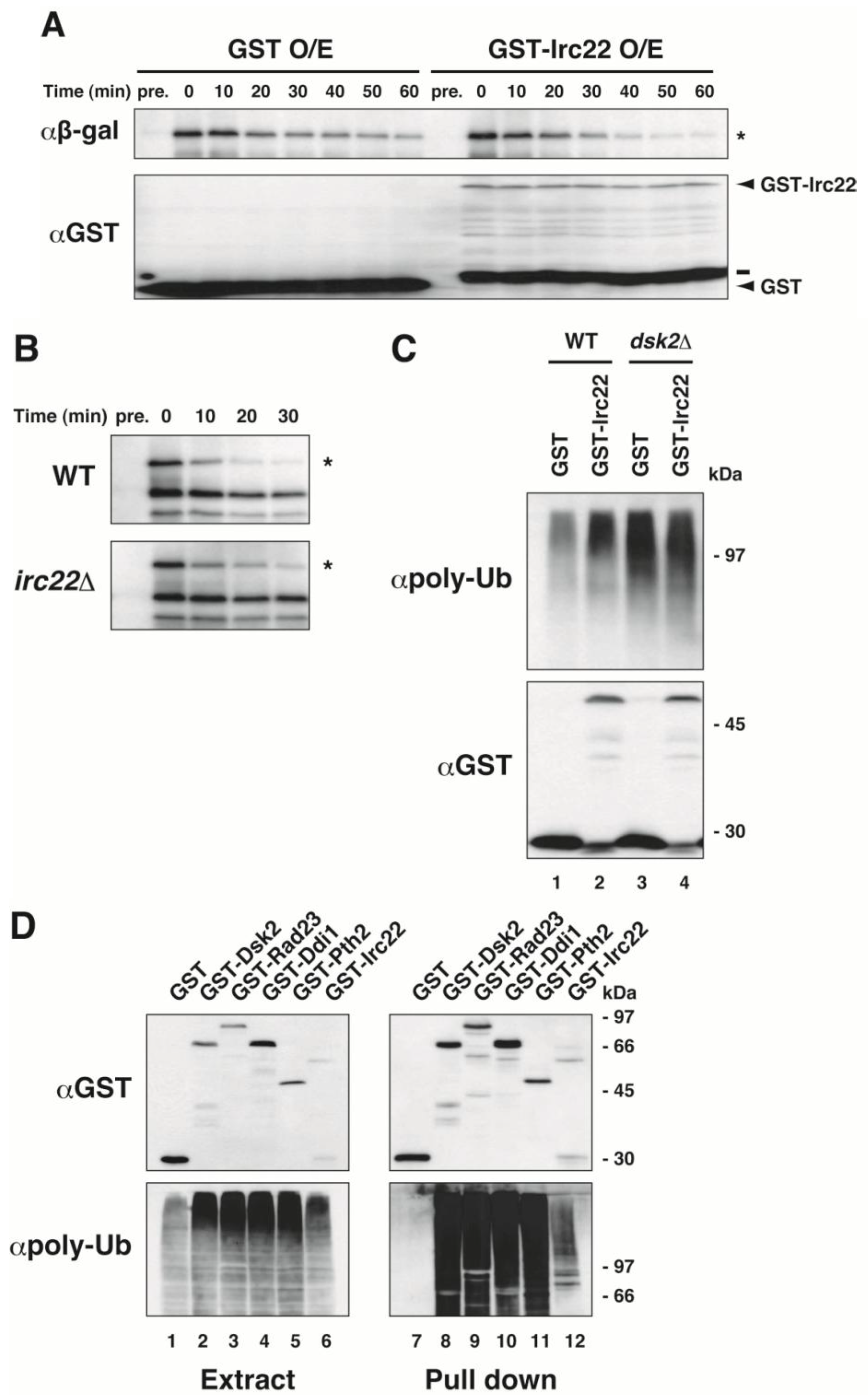
3.3. Irc22 Affects the Ubiquitin-Dependent Degradation Pathway in Yeast
3.4. IRC22 is Involved in Salt Stress Responses in Yeast

3.5. Discussion
3.5.1. Distinct Functions of the Middle Stretch of UBL-UBA Proteins
3.5.2. Irc22 Acts as a Negative Regulator of Dsk2 in Salt Tolerance
3.5.3. Possible Link between the UBL-UBA Protein Dsk2 and Salt Stress Tolerance in Yeast
4. Conclusions
Supplementary Files
Acknowledgments
Author Contributions
Conflicts of Interest
References and Notes
- Finley, D. Recognition and processing of ubiquitin-protein conjugates by the proteasome. Annu. Rev. Biochem. 2009, 78, 477–513. [Google Scholar] [CrossRef]
- Finley, D.; Ulrich, H.D.; Sommer, T.; Kaiser, P. The ubiquitin-proteasome system of Saccharomyces cerevisiae. Genetics 2012, 192, 319–360. [Google Scholar] [CrossRef]
- Elsasser, S.; Finley, D. Delivery of ubiquitinated substrates to protein-unfolding machines. Nat. Cell Biol. 2005, 7, 742–749. [Google Scholar] [CrossRef]
- Kirkin, V.; Dikic, I. Role of ubiquitin- and Ubl-binding proteins in cell signaling. Curr. Opn. Cell Biol. 2007, 19, 199–205. [Google Scholar] [CrossRef]
- Hochstrasser, M. Origin and function of ubiquitin-like proteins. Nature 2009, 458, 422–429. [Google Scholar] [CrossRef]
- Su, V.; Lau, A.F. Ubiquitin-like and ubiquitin-associated domain proteins: significance in proteasomal degradation. Cell. Mol. Life Sci. 2009, 66, 2819–2833. [Google Scholar] [CrossRef]
- Wilkinson, C.R.M.; Seeger, M.; Hartmann-Petersen, R.; Stone, M.; Wallace, M.; Semple, C.; Gordon, C. Proteins containing the UBA domain are able to bind to multi-ubiquitin chains. Nat. Cell Biol. 2001, 3, 939–943. [Google Scholar]
- Funakoshi, M.; Sasaki, T.; Nishimoto, T.; Kobayashi, H. Budding yeast Dsk2p is a polyubiquitin-binding protein that can interact with the proteasome. Proc. Natl. Acad. Sci. USA 2002, 99, 745–750. [Google Scholar] [CrossRef]
- Rao, H.; Sastry, A. Recognition of specific ubiquitin conjugates is important for the proteolytic functions of the ubiquitin-associated domain proteins Dsk2 and Rad23. J. Biol. Chem. 2002, 277, 11691–11695. [Google Scholar] [CrossRef]
- Elsasser, S.; Gali, R.R.; Schwickart, M.; Larsen, C.N.; Leggett, D.S.; Muller, B.; Feng, M.T.; Tubing, F.; Dittmar, G.A.; Finley, D. Proteasome subunit Rpn1 binds ubiquitin-like protein domains. Nat. Cell Biol. 2002, 4, 725–730. [Google Scholar] [CrossRef]
- Elsasser, S.; Chandler-Militello, D.; Muller, B.; Hanna, J.; Finley, D. Rad23 and Rpn10 serve as alternative ubiquitin receptors for the proteasome. J. Biol. Chem. 2004, 279, 26817–26822. [Google Scholar] [CrossRef]
- Husnjak, K.; Elsasser, S.; Zhang, N.; Chen, X.; Randles, L.; Shi, Y.; Hofmann, K.; Walters, K.J.; Finley, D.; Dikic, I. Proteasome subunit Rpn13 is a novel ubiquitin receptor. Nature 2008, 453, 481–488. [Google Scholar] [CrossRef]
- Fatimababy, A.S.; Lin, Y.L.; Usharani, R.; Radjacommare, R.; Wang, H.T.; Tsai, H.L.; Lee, Y.; Fu, H. Cross-species divergence of the major recognition pathways of ubiquitylated substrates for ubiquitin/26S proteasome-mediated proteolysis. FEBS J. 2010, 277, 796–816. [Google Scholar] [CrossRef]
- Gomez, T.A.; Kolawa, N.; Gee, M.; Sweredoski, M.J.; Deshaies, R.J. Identification of a functional docking site in the Rpn1 LRR domain for the UBL-UBA domain protein Ddi1. BMC Biol. 2011, 9, 33. [Google Scholar] [CrossRef]
- Rosenzweig, R.; Bronner, V.; Zhang, D.; Fushman, D.; Glickman, M.H. Rpn1 and Rpn2 coordinate ubiquitin processing factors at proteasome. J. Biol. Chem. 2012, 287, 14659–14671. [Google Scholar] [CrossRef]
- Ivantsiv, Y.; Kaplun, L.; Tzirkin-Goldin, R.; Shabek, N.; Raveh, D. Unique role for the Ubl-UbA protein Ddi1 in turnover of SCFUfo1 complexes. Mol. Cell. Biol. 2006, 26, 1579–1588. [Google Scholar] [CrossRef]
- Ghaboosi, N.; Deshaies, R.J. A conditional yeast E1 mutant blocks the ubiquitin-proteasome pathway and reveals a role for ubiquitin conjugates in targeting Rad23 to the proteasome. Mol. Biol. Cell 2007, 18, 1953–1963. [Google Scholar] [CrossRef]
- Kaur, N.; Zhao, Q.; Xie, Q.; Hu, J. Arabidopsis RING peroxins are E3 ubiquitin ligases that interact with two homologous ubiquitin receptor proteins. J. Integr. Plant Biol. 2013, 55, 108–120. [Google Scholar] [CrossRef]
- Kim, I.; Mi, K.; Rao, H. Multiple interactions of rad23 suggest a mechanism for ubiquitylated substrate delivery important in proteolysis. Mol. Biol. Cell 2004, 15, 3357–3365. [Google Scholar] [CrossRef]
- Medicherla, B.; Kostova, Z.; Schaefer, A.; Wolf, D.H. A genomic screen identifies Dsk2p and Rad23p as essential components of ER-associated degradation. EMBO Rep. 2004, 5, 692–697. [Google Scholar] [CrossRef]
- Ishii, T.; Funakoshi, M.; Kobayashi, H. Yeast Pth2 is a UBL domain-binding protein that participates in the ubiquitin-proteasome pathway. EMBO J. 2006, 25, 5492–5503. [Google Scholar] [CrossRef]
- Hanzelmann, P.; Stingele, J.; Hofmann, K.; Schindelin, H.; Rassi, S. The yeast E4 ubiquitin ligase Ufd2 interacts with the ubiquitin-like domains of Rad23 and Dsk2 via a novel and distinct ubiquitin-like binding domain. J. Biol. Chem. 2010, 285, 20390–20398. [Google Scholar]
- Matiuhin, Y.; Kirkpatrick, D.S.; Ziv, I.; Kim, W.; Dakshinamurthy, A.; Kleifeld, O.; Gygi, S.P.; Reis, N.; Glickman, M.H. Extraproteasomal Rpn10 restricts access of the polyuibiquitin-binding protein Dsk2 to proteasome. Mol. Cell 2008, 32, 415–425. [Google Scholar] [CrossRef]
- Zhang, D.; Chen, T.; Ziv, I.; Rosenzweig, R.; Matiuhin, Y.; Bronner, V.; Glickman, M.H.; Fushman, D. Together, Rpn10 and Dsk2 can serve as a polyubiquitin chain-length sensor. Mol. Cell 2009, 36, 1018–1033. [Google Scholar] [CrossRef]
- Heinen, C.; Acs, K.; Hoogstraten, D.; Dantuma, N.P. C-terminal UBA domains protect ubiquitin receptors by preventing initiation of protein degradation. Nat. Commun. 2011, 2, 191. [Google Scholar] [CrossRef]
- Biggins, S.; Ivanovska, I.; Rose, M.D. Yeast ubiquitin-like genes are involved in duplication of the microtubule organizing center. J. Cell Biol. 1996, 133, 1331–1346. [Google Scholar] [CrossRef]
- Saeki, Y.; Saitoh, A.; Toh-e, A.; Yokosawa, H. Ubiquitin-like proteins and Rpn10 play cooperative roles in ubiquitin-dependent proteolysis. Biochem. Biophys. Res. Commun. 2002, 293, 986–992. [Google Scholar] [CrossRef]
- Diaz-Martinez, L.A.; Kang, Y.; Walters, K.J.; Clarke, D.J. Yeast UBL-UBA proteins have partially redundant functions in cell cycle control. Cell Division 2006, 1, 28. [Google Scholar] [CrossRef]
- Bertolaet, B.L.; Clarke, D.J.; Wolff, M.; Watson, M.H.; Henze, M.; Divita, G.; Reed, S.I. UBA domains mediate protein-protein interactions between two DNA damage-inducible proteins. J. Mol. Biol. 2001, 313, 955–963. [Google Scholar] [CrossRef]
- Ryu, K.S.; Lee, K.J.; Bae, S.H.; Kim, B.K.; Kim, K.A.; Choi, B.S. Binding surface mapping of intra- and interdomain interactions among hHR23B, ubiquitin, and polyubiquitin binding site 2 of S5a. J. Biol. Chem. 2003, 278, 36621–36627. [Google Scholar]
- Sasaki, T.; Funakoshi, M.; Endicott, J.A.; Kobayashi, H. Budding yeast Dsk2 protein forms a homodimer via its C-terminal UBA domain. Biochem. Biophys. Res. Commun. 2005, 336, 530–535. [Google Scholar] [CrossRef]
- Kang, Y.; Vossler, R.A.; Diaz-Martinez, L.A.; Winter, N.S.; Clarke, D.J.; Walters, K.J. UBL/UBA ubiquitin receptor proteins bind a common tetraubiquitin chain. J. Mol. Biol. 2006, 356, 1027–1035. [Google Scholar] [CrossRef]
- Lowe, E.D.; Hassan, N.; Trempe, J.F.; Fonso, L.; Noble, M.E.; Endicott, J.A.; Johnson, L.N.; Brown, N.R. Structures of the Dsk2 UBL and UBA domains and their complex. Acta Cryst. 2006, D62, 177–188. [Google Scholar]
- Kang, Y.; Zhang, N.; Koepp, D.M.; Walters, K.J. Ubiquitin receptor proteins hHR23a and hPLIC2 interact. J. Mol. Biol. 2007, 365, 1093–1101. [Google Scholar] [CrossRef]
- Dantuma, N.P.; Heinen, C.; Hoogstraten, D. The ubiquitin receptor Rad23: at the crossroads of nucleotide excision repair and proteasomal degradation. DNA repair 2009, 8, 449–460. [Google Scholar] [CrossRef]
- Li, Y.; Yan, J.; Kim, I.; Liu, C.; Huo, K.; Rao, H. Rad4 regulates protein turnover at a postubiquitylation step. Mol. Biol. Cell 2010, 21, 177–185. [Google Scholar] [CrossRef]
- Sirkis, R.; Gerst, J.E.; Fass, D. Ddi1, a eukaryotic protein with the retroviral protease fold. J. Mol. Biol. 2006, 364, 376–387. [Google Scholar] [CrossRef]
- Gabriely, G.; Kama, R.; Gelin-Licht, R.; Gerst, J.E. Different domains of the UBL-UBA ubiquitin receptor, Ddi1/Vsm1, are involved in its multiple cellular roles. Mol. Biol. Cell 2008, 19, 3625–3637. [Google Scholar] [CrossRef]
- Kaye, F.J.; Modi, S.; Ivanovska, I.; Koonin, E.V.; Thress, K.; Kubo, A.; Kornbluth, S.; Rose, M.D. A family of ubiquitin-like proteins binds the ATPase domain of Hsp70-like Stech. FEBS Lett. 2000, 467, 348–352. [Google Scholar] [CrossRef]
- Woznik, M.; Ronder, C.; Lemon, K.; Rima, B.; Mankertz, A.; Finsterbusch, T. Mumps virus small hydrophobic protein targets ataxin-1 ubiquitin-like interacting protein (ubiquilin 4). J. Gen. Virol. 2010, 91, 2773–2781. [Google Scholar] [CrossRef]
- Sun, X.; Meng, X.; Xu, Z.; Song, R. Expression of the 26S proteasome subunit RPN10 is upregulated by salt stress in Dunaliella viridis. J. Plant Physiol. 2010, 167, 1003–1008. [Google Scholar] [CrossRef]
- Cui, F.; Liu, L.; Zhao, Q.; Zhang, Z.; Li, Q.; Lin, B.; Wu, Y.; Tang, S.; Xie, Q. Arabidopsis ubiquitin conjugate UBC32 is an ERAD component that functions in Brassinosteroid-mediated salt stress tolerance. Plant Cell 2012, 24, 233–244. [Google Scholar] [CrossRef]
- Kim, J.H.; Kim, W.T. The Arabidopsis RING E3 ubiquitin ligase AtAIRP3/LOG2 participates in positive regulation of high-salt and drought stress responses. Plant Physiol. 2013, 162, 1733–1749. [Google Scholar] [CrossRef]
- Liu, L.; Cui, F.; Li, Q.; Yin, B.; Zhang, H.; Lin, B.; Wu, Y.; Xia, R.; Tang, S.; Xie, Q. The endoplasmic reticulum-associated degradation is necessary for plant salt tolerance. Cell Res. 2011, 21, 957–969. [Google Scholar] [CrossRef]
- Sikorsky, R.S.; Hieter, P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 1989, 122, 19–27. [Google Scholar]
- Adams, A.; Gottschling, D.E.; Kaiser, C.A.; Stearns, T. Gene replacemen. In Methods in Yeast Genetics; Adams, A., Ed.; CSHL Press: Cold Spring Harbor, New York, NY, USA, 1997; pp. 59–70. [Google Scholar]
- Bachmair, A.; Finley, D.; Varshavsky, A. In vivo half-life of a protein is a function of its amino-terminal residue. Science 1986, 234, 179–186. [Google Scholar]
- Varshavsky, A. The N-end rule: Functions, mysteries, uses. Proc. Natl. Acad. Sci. USA 1996, 93, 12142–12149. [Google Scholar] [CrossRef]
- Runder, A.D.; Hardwick, K.G.; Murray, A. Cdc28 activates exit from mitosis in budding yeast. J. Cell Biol. 2000, 149, 1361–1376. [Google Scholar] [CrossRef]
- Saccharomyces Genome Database Home Page. Available online: http://www.yeastgenome.org/ (accessed on 15 January 2014).
- Babu, M.; Vlasblom, J.; Pu, S.; Guo, X.; Graham, C.; Bean, B.D.; Burston, H.E.; Vizeacoumar, F.J.; Snider, J.; Phanse, S.; et al. Interaction landscape of membrane-protein complexes in Saccharomyces cerevisiae. Nature 2012, 489, 585–589. [Google Scholar] [CrossRef]
- OrthoDB: Database of Orthologous Groups Home Page. Available online: http://cegg.unige.ch/orthodb7 (accessed on 15 January 2014).
- Masutani, C.; Araki, M.; Sugasawa, K.; van der Spek, P.J.; Yamada, A.; Uchida, A.; Maekawa, T.; Bootsma, D.; Hoeijmakers, J.H.; Hanaoka, F. Identification and characterization of XPC-binding domain of hHR23B. Mol. Cell. Biol. 1997, 17, 6915–6923. [Google Scholar]
- Raasi, S.; Varadan, R.; Fushman, D.; Pickart, C.M. Diverse polyubiquitin interaction properties of ubiquitin-associated domains. Nat. Struct. Mol. Biol. 2005, 12, 708–714. [Google Scholar] [CrossRef]
- Kulathu, Y.; Komander, D. Atypical ubiquitylation - the unexplored world of polyubiquitin beyond Lys48 and Lys63 linkages. Nat. Rev. Mol. Cell Biol. 2012, 13, 508–523. [Google Scholar] [CrossRef]
- Nunez, L.R.; Jesch, S.A.; Gasper, M.L.; Almaguer, C.; Villa-Garcia, M.; Ruiz-Noriega, M.; Patton-Vogt, J.; Henry, S.A. Cell wall integrity MAPK pathway is essential for lipid homeostasis. J. Biol. Chem. 2008, 283, 34204–34217. [Google Scholar] [CrossRef]
- Fernandez-Murray, J.P.; Gaspard, G.J.; Jesch, S.A.; McMaster, C.R. NTE1-encoded phophatidylcholine phospholipase B regulates transcription of phospholipid biosynthetic genes. J. Biol. Chem. 2009, 284, 36034–36046. [Google Scholar] [CrossRef]
- Zhu, J.Q.; Zhang, J.T.; Tang, R.J.; Lv, Q.D.; Wang, Q.Q.; Yang, L.; Zhang, H.X. Molecular characterization of ThIPK2, an inositol polyphosphate kinase gene homolog from Thellungiella halophia, and its heterologous expression to improve abiotic stress tolerance in Brassica napus. Physiol. Plant. 2009, 136, 407–425. [Google Scholar] [CrossRef]
- Golani, Y.; Kaye, Y.; Gilhar, O.; Ercetin, M.; Gillaspy, G.; Levine, A. Inositol polyphosphate phosphatidylinositol 5-phosphatase9 (At5PTase9) controls plant salt tolerance by regulating endocytosis. Mol. Plant 2013, 6, 1781–1794. [Google Scholar] [CrossRef]
- Villa-Garcia, M.J.; Choi, M.S.; Hinz, F.I.; Gasper, M.L.; Jesch, S.A.; Henry, S.A. Genome-wide screen for inositol auxotrophy in Saccharomyces cerevisiae implicates lipid metabolism in stress response signaling. Mol. Genet. Genomics 2011, 285, 125–149. [Google Scholar] [CrossRef]
- Karbowski, M.; Youle, R.J. Regulating mitochondorial outer membrane proteins by ubiquitination and protein degradation. Curr. Opin. Cell Biol. 2011, 23, 476–482. [Google Scholar] [CrossRef]
- Jones, C.B.; Ott, E.M.; Keener, J.M.; Curtiss, M.; Sandrin, V.; Babst, M. Regulation of membrane proteasomal degradation by starvation-response pathways. Traffic 2012, 13, 468–482. [Google Scholar] [CrossRef]
- Michaillat, L.; Mayer, A. Identification of genes affecting vacuole membrane fragmentation in Saccharomyces cerevisiae. PLoS One 2013, 8, e54160. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Ishii, T.; Funakoshi, M.; Kobayashi, H.; Sekiguchi, T. Yeast Irc22 Is a Novel Dsk2-Interacting Protein that Is Involved in Salt Tolerance. Cells 2014, 3, 180-198. https://doi.org/10.3390/cells3020180
Ishii T, Funakoshi M, Kobayashi H, Sekiguchi T. Yeast Irc22 Is a Novel Dsk2-Interacting Protein that Is Involved in Salt Tolerance. Cells. 2014; 3(2):180-198. https://doi.org/10.3390/cells3020180
Chicago/Turabian StyleIshii, Takashi, Minoru Funakoshi, Hideki Kobayashi, and Takeshi Sekiguchi. 2014. "Yeast Irc22 Is a Novel Dsk2-Interacting Protein that Is Involved in Salt Tolerance" Cells 3, no. 2: 180-198. https://doi.org/10.3390/cells3020180



