Association of Differentiation-Related Gene-1 (DRG1) with Breast Cancer Survival and in Vitro Impact of DRG1 Suppression
Abstract
:1. Introduction
2. Experimental Section
2.1. Cell Lines and Culture Conditions
2.2. Patient Samples and Preparation
| Clinical data | Sample numbers | |
|---|---|---|
| Grade | ||
| 1 | 20 | |
| 2 | 38 | |
| 3 | 54 | |
| TNM | ||
| 1 | 60 | |
| 2 | 40 | |
| 3 | 7 | |
| 4 | 4 | |
| NPI | ||
| 1 | 57 | |
| 2 | 38 | |
| 3 | 15 | |
| Outcome | ||
| Disease free | 80 | |
| With metastasis | 7 | |
| With local recurrence | 5 | |
| Died of breast cancer | 14 | |
2.3. DRG1 Knockdown in Human Breast Cancer Cell Lines
| Primer | Forward Primers | Reverse primers |
|---|---|---|
| DRG1 Conventional | GGACGATTTCACAAAAACAT | CATCTTCATACTGCAAAGCA |
| DRG1 Quantitative | TGCTACAGCTGATGACCTC | ACTGAACCTGACCTGACCGTA CACCAATTCCTCAATGGAGAT |
| GAPDH Conventional | GGCTGCTTTTAACTCTGGTA | GACTGTGGTCATGAGTCCTT |
| GAPDH Quantitative | CTGAGTACGTCGTGGAGTC | ACTGAACCTGACCGTACACAGAGA TGATGACCCTTTTG |
| DRG1 Ribozyme1 | CTGCAGCAGTGTTGACTTCCCCACACTGATGAGTCCGTGAGGA | ACTAGTGATGCTCGAATTGGATTTGTTGGTTTTCCATTTCGTCCTCACGGACT |
| DRG1 Ribozyme2 | CTGCAGGACATCCAGAACAATCAACTGATGAGTCCGTGAGGA | ACTAGTGGCCCGAACCTGTAACTTGATTTCGTCCTCACGGACT |
2.4. RNA Extraction and Reverse Transcription-Polymerase Chain Reaction
2.5. Quantitative-Polymerase Chain Reaction (Q-PCR)
2.6. SDS-PAGE and Western Blotting
2.7. In Vitro Growth Assay
2.8. In Vitro Matrigel Adhesion Assay
2.9. Electric Cell-Substrate Impedance Sensing Analysis of Breast Cancer Cell Lines Migration
2.10. In Vitro Matrigel Invasion Assay
2.11. Statistical Analysis
3. Results
3.1. Correlation of DRG1 Transcript Expression with Grade, TNM and NPI Status
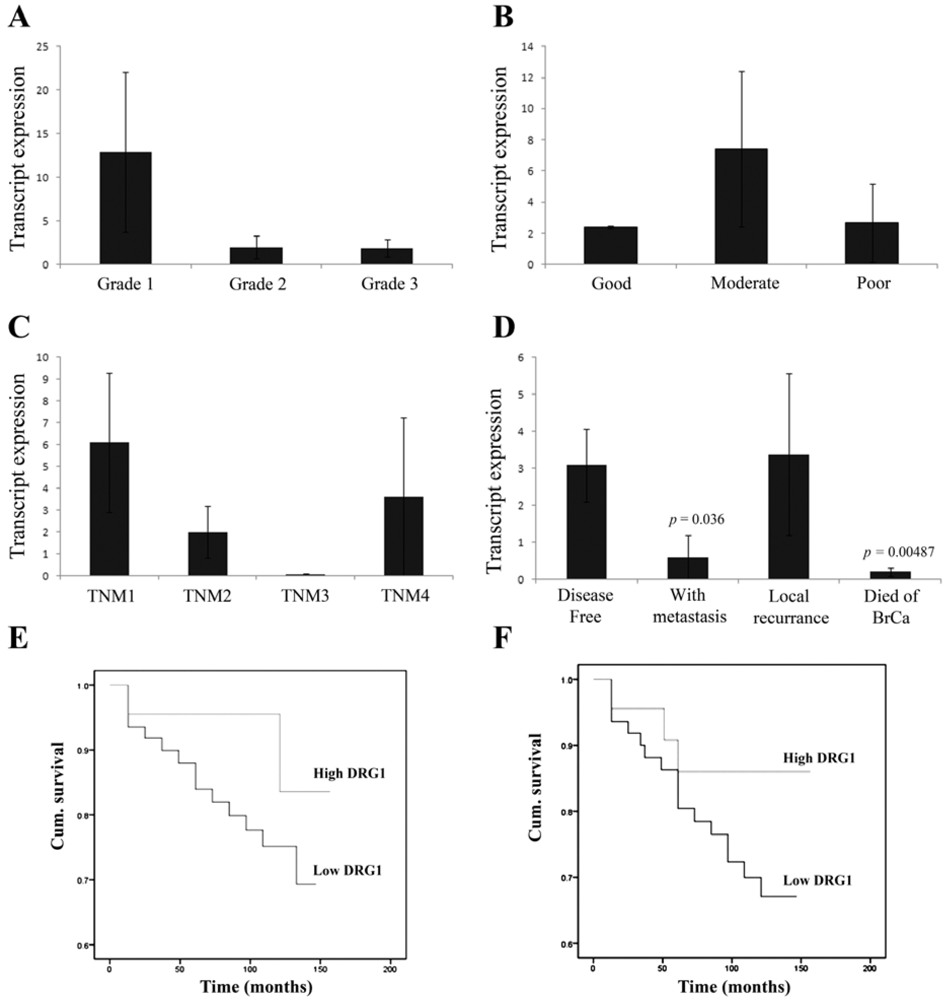
3.2. Reduced DRG1 Expression Is Associated with Poorer Patient Prognosis
3.3. Knockdown of DRG1 in Breast Cancer Cell Lines
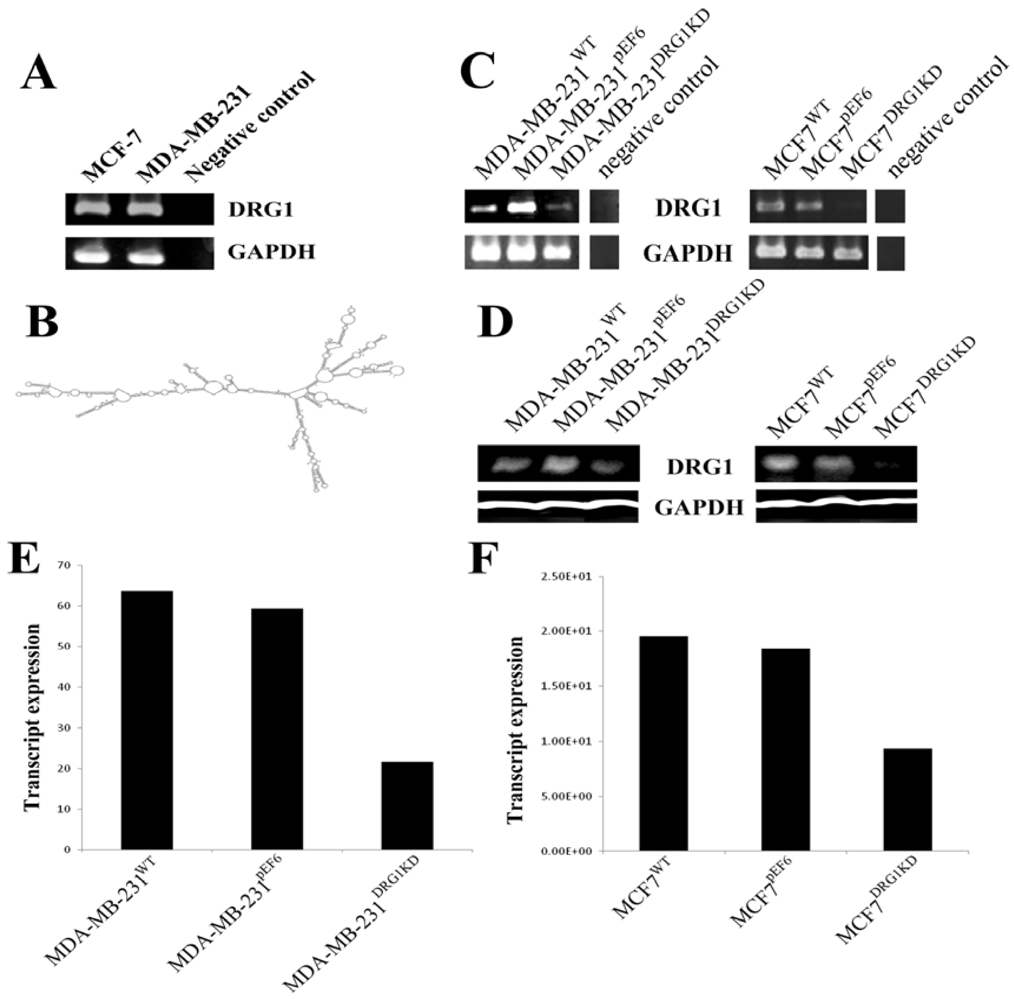
3.4. Effect of DRG1 Knockdown on the Growth of Breast Cancer Cells
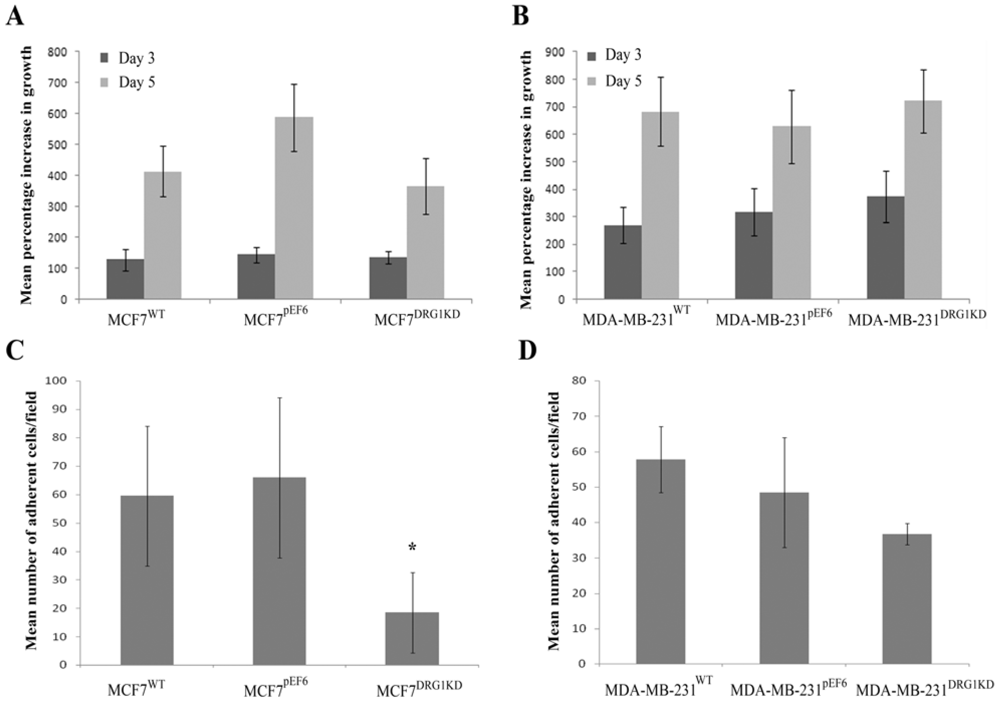
3.5. Effect of DRG1 Knockdown on Adhesion of Breast Cancer Cells
3.6. Effect of DRG1 Knockdown on Invasion of Breast Cancer Cells
3.7. Effect of DRG1 Knockdown on Migrational Rates of Breast Cancer Cells
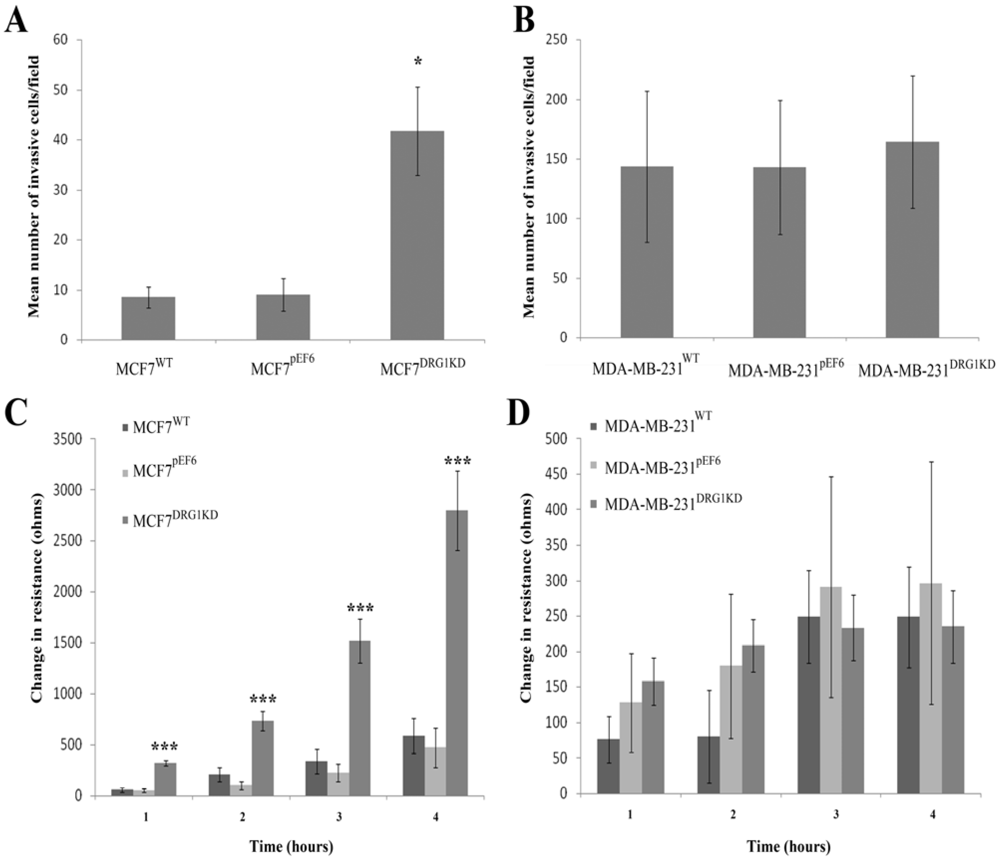
4. Discussion
5. Conclusions
Acknowledgements
References
- Sobel, M.E. Metastasis suppressor genes. J. Natl. Cancer Inst. 1990, 82, 267–276. [Google Scholar] [CrossRef]
- Freije, J.M.; MacDonald, N.J.; Steeg, P.S. Nm23 and tumour metastasis: Basic and translational advances. Biochem. Soc. Symp. 1998, 63, 261–271. [Google Scholar]
- van Belzen, N.; Dinjens, W.N.; Diesveld, M.P.; Groen, N.A.; van der Made, A.C.; Nozawa, Y.; Vlietstra, R.; Trapman, J.; Bosman, F.T. A novel gene which is up-regulated during colon epithelial cell differentiation and down-regulated in colorectal neoplasms. Lab. Invest. 1997, 77, 85–92. [Google Scholar]
- Guan, R.J.; Ford, H.L.; Fu, Y.; Li, Y.; Shaw, L.M.; Pardee, A.B. Drg-1 as a differentiation-related, putative metastatic suppressor gene in human colon cancer. Cancer Res. 2000, 60, 749–755. [Google Scholar]
- Kokame, K.; Kato, H.; Miyata, T. Homocysteine-respondent genes in vascular endothelial cells identified by differential display analysis. GRP78/BiP and novel genes. J. Biol. Chem. 1996, 271, 29659–29665. [Google Scholar]
- Lin, T.M.; Chang, C. Cloning and characterization of TDD5, an androgen target gene that is differentially repressed by testosterone and dihydrotestosterone. Proc. Natl. Acad. Sci. USA 1997, 94, 4988–4993. [Google Scholar]
- Zhou, D.; Salnikow, K.; Costa, M. Cap43, a novel gene specifically induced by Ni2+ compounds. Cancer Res. 1998, 58, 2182–2189. [Google Scholar]
- Piquemal, D.; Joulia, D.; Balaguer, P.; Basset, A.; Marti, J.; Commes, T. Differential expression of the RTP/Drg1/Ndr1 gene product in proliferating and growth arrested cells. Biochim. Biophys. Acta 1450, 364–373. [Google Scholar]
- Okuda, T.; Kondoh, H. Identification of new genes Ndr2 and Ndr3 which are related to Ndr1/RTP/Drg1 but show distinct tissue specificity and response to N-myc. Biochem. Biophys. Res. Commun. 1999, 266, 208–215. [Google Scholar] [CrossRef]
- Ulrix, W.; Swinnen, J.V.; Heyns, W.; Verhoeven, G. The differentiation-related gene 1, Drg1, is markedly upregulated by androgens in LNCaP prostatic adenocarcinoma cells. FEBS Lett. 1999, 455, 23–26. [Google Scholar] [CrossRef]
- Park, H.; Adams, M.A.; Lachat, P.; Bosman, F.; Pang, S.C.; Graham, C.H. Hypoxia induces the expression of a 43-kDa protein (PROXY-1) in normal and malignant cells. Biochem. Biophys. Res. Commun. 2000, 276, 321–328. [Google Scholar] [CrossRef]
- Unoki, M.; Nakamura, Y. Growth-suppressive effects of BPOZ and EGR2, two genes involved in the PTEN signaling pathway. Oncogene 2001, 20, 4457–4465. [Google Scholar] [CrossRef]
- Le, N.T.; Richardson, D.R. Iron chelators with high antiproliferative activity up-regulate the expression of a growth inhibitory and metastasis suppressor gene: A link between iron metabolism and proliferation. Blood 2004, 104, 2967–2975. [Google Scholar] [CrossRef]
- Kurdistani, S.K.; Arizti, P.; Reimer, C.L.; Sugrue, M.M.; Aaronson, S.A.; Lee, S.W. Inhibition of tumor cell growth by RTP/rit42 and its responsiveness to p53 and DNA damage. Cancer Res. 1998, 58, 4439–4444. [Google Scholar]
- Masuda, K.; Ono, M.; Okamoto, M.; Morikawa, W.; Otsubo, M.; Migita, T.; Tsuneyoshi, M.; Okuda, H.; Shuin, T.; Naito, S.; et al. Downregulation of Cap43 gene by von Hippel-Lindau tumor suppressor protein in human renal cancer cells. Int. J. Cancer 2003, 105, 803–810. [Google Scholar] [CrossRef]
- Bandyopadhyay, S.; Pai, S.K.; Gross, S.C.; Hirota, S.; Hosobe, S.; Miura, K.; Saito, K.; Commes, T.; Hayashi, S.; Watabe, M.; et al. The Drg-1 gene suppresses tumor metastasis in prostate cancer. Cancer Res. 2003, 63, 1731–1736. [Google Scholar]
- Shah, M.A.; Kemeny, N.; Hummer, A.; Drobnjak, M.; Motwani, M.; Cordon-Cardo, C.; Gonen, M.; Schwartz, G.K. Drg1 expression in 131 colorectal liver metastases: Correlation with clinical variables and patient outcomes. Clin. Cancer Res. 2005, 11, 3296–3302. [Google Scholar] [CrossRef]
- Bandyopadhyay, S.; Pai, S.K.; Hirota, S.; Hosobe, S.; Takano, Y.; Saito, K.; Piquemal, D.; Commes, T.; Watabe, M.; Gross, S.C.; et al. Role of the putative tumor metastasis suppressor gene Drg-1 in breast cancer progression. Oncogene 2004, 23, 5675–5681. [Google Scholar]
- Kim, K.T.; Ongusaha, P.P.; Hong, Y.K.; Kurdistani, S.K.; Nakamura, M.; Lu, K.P.; Lee, S.W. Function of Drg1/Rit42 in p53-dependent mitotic spindle checkpoint. J. Biol. Chem. 2004, 279, 38597–38602. [Google Scholar]
- Cangul, H. Hypoxia upregulates the expression of the NDRG1 gene leading to its overexpression in various human cancers. BMC Genet. 2004, 5, 27. [Google Scholar] [CrossRef] [Green Version]
- Kovacevic, Z.; Richardson, D.R. The metastasis suppressor, Ndrg-1: A new ally in the fight against cancer. Carcinogenesis 2006, 27, 2355–2366. [Google Scholar] [CrossRef]
- Caruso, R.P.; Levinson, B.; Melamed, J.; Wieczorek, R.; Taneja, S.; Polsky, D.; Chang, C.; Zeleniuch-Jacquotte, A.; Salnikow, K.; Yee, H.; et al. Altered N-myc downstream-regulated gene 1 protein expression in African-American compared with caucasian prostate cancer patients. Clin. Cancer Res. 2004, 10, 222–227. [Google Scholar]
- Jiang, W.G.; Davies, G.; Martin, T.A.; Parr, C.; Watkins, G.; Mason, M.D.; Mokbel, K.; Mansel, R.E. Targeting matrilysin and its impact on tumor growth in vivo: The potential implications in breast cancer therapy. Clin. Cancer Res. 2005, 11, 6012–6019. [Google Scholar]
- Jiang, W.G.; Grimshaw, D.; Lane, J.; Martin, T.A.; Abounader, R.; Laterra, J.; Mansel, R.E. A hammerhead ribozyme suppresses expression of hepatocyte growth factor/scatter factor receptor c-MET and reduces migration and invasiveness of breast cancer cells. Clin. Cancer Res. 2001, 7, 2555–2562. [Google Scholar]
- Sanders, A.J.; Parr, C.; Mason, M.D.; Jiang, W.G. Suppression of hepatocyte growth factor activator inhibitor-1 leads to a more aggressive phenotype of prostate cancer cells in vitro. Int. J. Mol. Med. 2007, 20, 613–619. [Google Scholar]
- Jiang, W.G.; Martin, T.A.; Lewis-Russell, J.M.; Douglas-Jones, A.; Ye, L.; Mansel, R.E. Eplin-alpha expression in human breast cancer, the impact on cellular migration and clinical outcome. Mol. Cancer 2008, 7, 71. [Google Scholar]
- Keese, C.R.; Wegener, J.; Walker, S.R.; Giaever, I. Electrical wound-healing assay for cells in vitro. Proc. Natl. Acad. Sci. USA 2004, 101, 1554–1559. [Google Scholar]
- Parr, C.; Davies, G.; Nakamura, T.; Matsumoto, K.; Mason, M.D.; Jiang, W.G. The HGF/SF-induced phosphorylation of paxillin, matrix adhesion, and invasion of prostate cancer cells were suppressed by NK4, an HGF/SF variant. Biochem. Biophys. Res. Commun. 2001, 285, 1330–1337. [Google Scholar] [CrossRef]
- Yamada, S.D.; Hickson, J.A.; Hrobowski, Y.; vander Griend, D.J.; Benson, D.; Montag, A.; Karrison, T.; Huo, D.; Rutgers, J.; Adams, S.; et al. Mitogen-activated protein kinase kinase 4 (MKK4) acts as a metastasis suppressor gene in human ovarian carcinoma. Cancer Res. 2002, 62, 6717–6723. [Google Scholar]
- Strzelczyk, B.; Szulc, A.; Rzepko, R.; Kitowska, A.; Skokowski, J.; Szutowicz, A.; Pawelczyk, T. Identification of high-risk stage II colorectal tumors by combined analysis of the NDRG1 gene expression and the depth of tumor invasion. Ann. Surg. Oncol. 2009, 16, 1287–1294. [Google Scholar]
- Kalaydjieva, L.; Hallmayer, J.; Chandler, D.; Savov, A.; Nikolova, A.; Angelicheva, D.; King, R.H.; Ishpekova, B.; Honeyman, K.; Calafell, F.; et al. Gene mapping in Gypsies identifies a novel demyelinating neuropathy on chromosome 8q24. Nat. Genet. 1996, 14, 214–217. [Google Scholar]
- Maruyama, Y.; Ono, M.; Kawahara, A.; Yokoyama, T.; Basaki, Y.; Kage, M.; Aoyagi, S.; Kinoshita, H.; Kuwano, M. Tumor growth suppression in pancreatic cancer by a putative metastasis suppressor gene Cap43/NDRG1/Drg-1 through modulation of angiogenesis. Cancer Res. 2006, 66, 6233–6242. [Google Scholar]
- Levenson, A.S.; Jordan, V.C. MCF-7: The first hormone-responsive breast cancer cell line. Cancer Res. 1997, 57, 3071–3078. [Google Scholar]
- Yoshida, B.A.; Sokoloff, M.M.; Welch, D.R.; Rinker-Schaeffer, C.W. Metastasis-suppressor genes: A review and perspective on an emerging field. J. Natl. Cancer Inst. 2000, 92, 1717–1730. [Google Scholar] [CrossRef]
- Liu, Y.N.; Lee, W.W.; Wang, C.Y.; Chao, T.H.; Chen, Y.; Chen, J.H. Regulatory mechanisms controlling human E-cadherin gene expression. Oncogene 2005, 24, 8277–8290. [Google Scholar]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Baig, R.M.; Sanders, A.J.; Kayani, M.A.; Jiang, W.G. Association of Differentiation-Related Gene-1 (DRG1) with Breast Cancer Survival and in Vitro Impact of DRG1 Suppression. Cancers 2012, 4, 658-672. https://doi.org/10.3390/cancers4030658
Baig RM, Sanders AJ, Kayani MA, Jiang WG. Association of Differentiation-Related Gene-1 (DRG1) with Breast Cancer Survival and in Vitro Impact of DRG1 Suppression. Cancers. 2012; 4(3):658-672. https://doi.org/10.3390/cancers4030658
Chicago/Turabian StyleBaig, Ruqia Mehmood, Andrew J. Sanders, Mahmood Akhtar Kayani, and Wen G. Jiang. 2012. "Association of Differentiation-Related Gene-1 (DRG1) with Breast Cancer Survival and in Vitro Impact of DRG1 Suppression" Cancers 4, no. 3: 658-672. https://doi.org/10.3390/cancers4030658




