Fungal Cytochrome P450s and the P450 Complement (CYPome) of Fusarium graminearum
Abstract
:1. Introduction
2. Fungal CYPs
3. CYPs Related to Secondary Metabolite Biosynthesis
3.1. Aflatoxins and Sterigmatocystin
3.2. Fumonisins
3.3. Host-Selective Toxins
3.4. Dothistromin
3.5. Botridial
3.6. Ochratoxin A
4. Xenobiotic-Metabolizing CYPs
5. CYPs Required for Fungal Development and Virulence
6. CYPs of F. graminearum
6.1. Trichothecenes
6.2. Xenobiotic-Metabolizing CYPs in F. graminearum
6.3. CYPs Required for Fungal Development and Virulence in F. graminearum
7. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Nelson, D.R.; Koymans, L.; Kamataki, T.; Stegeman, J.J.; Feyereisen, R.; Waxman, D.J.; Waterman, M.R.; Gotoh, O.; Coon, M.J.; Estabrook, R.W. P450 superfamily: Update on new sequences, gene mapping, accession numbers and nomenclature. Pharmacogenetics 1996, 6, 1–42. [Google Scholar] [CrossRef] [PubMed]
- Lamb, D.C.; Lei, L.; Warrilow, A.G.; Lepesheva, G.I.; Mullins, J.G.; Waterman, M.R.; Kelly, S.L. The first virally encoded cytochrome P450. J. Virol. 2009, 83, 8266–8269. [Google Scholar] [CrossRef] [PubMed]
- Sanglard, D.; Loper, J.C. Characterization of the alkane-inducible cytochrome P450 (P450alk) gene from the yeast Candida tropicalis: Identification of a new P450 gene family. Gene 1989, 76, 121–136. [Google Scholar] [CrossRef]
- Mansuy, D. The great diversity of reactions catalyzed by cytochromes P450. Comp. Biochem. Physiol. C Pharmacol. Toxicol. Endocrinol. 1998, 121, 5–14. [Google Scholar] [CrossRef]
- Sono, M.; Roach, M.P.; Coulter, E.D.; Dawson, J.H. Heme-containing oxygenases. Chem. Rev. 1996, 96, 2841–2888. [Google Scholar] [CrossRef] [PubMed]
- Urlacher, V.B.; Girhard, M. Cytochrome P450 monooxygenases: An update on perspectives for synthetic application. Trends Biotechnol. 2012, 30, 26–36. [Google Scholar] [CrossRef] [PubMed]
- Nelson, D.R.; Nebert, D.W. Cytochrome P450 (CYP) gene superfamily. In eLS; John Wiley & Sons, Ltd.: Hoboken, NJ, USA, 2001; ISBN 9780470015902. [Google Scholar]
- Guengerich, F.P. Human cytochrome P450 enzymes. In Cytochrome P450; Springer: Cham, Switzerland, 2005; pp. 377–530. ISBN 978-3-319-12108-6. [Google Scholar]
- Schuler, M.A. P450s in plants, insects, and their fungal pathogens. In Cytochrome P450: Structure, Mechanism, and Biochemistry; Ortiz de Montellano, P.R., Ed.; Springer: Cham, Switzerland, 2015; pp. 409–449. ISBN 978-3-319-12108-6. [Google Scholar]
- Jensen, N.B.; Zagrobelny, M.; Hjernø, K.; Olsen, C.E.; Houghton-Larsen, J.; Borch, J.; Møller, B.L.; Bak, S. Convergent evolution in biosynthesis of cyanogenic defence compounds in plants and insects. Nat. Commun. 2011, 2, 273. [Google Scholar] [CrossRef] [PubMed]
- Cui, S.; Wang, L.; Ma, L.; Geng, X. P450-mediated detoxification of botanicals in insects. Phytoparasitica 2016, 44, 585–599. [Google Scholar] [CrossRef]
- Bolwell, G.P.; Bozak, K.; Zimmerlin, A. Plant cytochrome P450. Phytochemistry 1994, 37, 1491–1506. [Google Scholar] [CrossRef]
- Park, J.; Lee, S.; Choi, J.; Ahn, K.; Park, B.; Park, J.; Kang, S.; Lee, Y.H. Fungal cytochrome P450 database. BMC Genom. 2008, 9, 402. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Lee, M.-K.; Jefcoate, C.; Kim, S.-C.; Chen, F.; Yu, J.-H. Fungal cytochrome P450 monooxygenases: Their distribution, structure, functions, family expansion, and evolutionary origin. Genome Biol. Evol. 2014, 6, 1620–1634. [Google Scholar] [CrossRef] [PubMed]
- van den Brink, H.J.M.; van Gorcom, R.F.M.; van den Hondel, C.A.M.J.J.; Punt, P.J. Cytochrome P450 enzyme systems in fungi. Fungal Genet. Biol. 1998, 23, 1–17. [Google Scholar] [CrossRef] [PubMed]
- George, H.L.; Hirschi, K.D.; VanEtten, H.D. Biochemical properties of the products of cytochrome P450 genes (PDA) encoding pisatin demethylase activity in Nectria haematococca. Arch. Microbiol. 1998, 170, 147–154. [Google Scholar] [CrossRef] [PubMed]
- Cerniglia, C.E.; Hebert, R.L.; Szaniszlo, P.J.; Gibson, D.T. Fungal transformation of naphthalene. Arch. Microbiol. 1978, 117, 135–143. [Google Scholar] [CrossRef] [PubMed]
- Sutherland, J.B. Detoxification of polycyclic aromatic hydrocarbons by fungi. J. Ind. Microbiol. 1992, 9, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Bezalel, L.; Hadar, Y.; Cerniglia, C.E. Enzymatic mechanisms involved in phenanthrene degradation by the white rot fungus Pleurotus ostreatus. Appl. Environ. Microbiol. 1997, 63, 2495–2501. [Google Scholar] [PubMed]
- Shin, J.Y.; Bui, D.-C.; Lee, Y.; Nam, H.; Jung, S.; Fang, M.; Kim, J.-C.; Lee, T.; Kim, H.; Choi, G.J.; et al. Functional characterization of cytochrome P450 monooxygenases in the cereal head blight fungus Fusarium graminearum. Environ. Microbiol. 2017, 19, 2053–2067. [Google Scholar] [CrossRef] [PubMed]
- Fan, J.; Urban, M.; Parker, J.E.; Brewer, H.C.; Kelly, S.L.; Hammond-Kosack, K.E.; Fraaije, B.A.; Liu, X.; Cools, H.J. Characterization of the sterol 14α-demethylases of Fusarium graminearum identifies a novel genus-specific CYP51 function. New Phytol. 2013, 198, 821–835. [Google Scholar] [CrossRef] [PubMed]
- Skaggs, B.; Alexander, J.; Pierson, C.; Schweitzer, K.; Chun, K.; Koegel, C.; Barbuch, R.; Bard, M. Cloning and characterization of the Saccharomyces cerevisiae C-22 sterol desaturase gene, encoding a second cytochrome P-450 involved in ergosterol biosynthesis. Gene 1996, 169, 105–109. [Google Scholar] [CrossRef]
- Yun, Y.; Yin, D.; Dawood, D.H.; Liu, X.; Chen, Y.; Ma, Z. Functional characterization of FgERG3 and FgERG5 associated with ergosterol biosynthesis, vegetative differentiation and virulence of Fusarium graminearum. Fungal Genet. Biol. 2014, 68, 60–70. [Google Scholar] [CrossRef] [PubMed]
- Newsome, A.W.; Nelson, D.; Corran, A.; Kelly, S.L.; Kelly, D.E. The cytochrome P450 complement (CYPome) of Mycosphaerella graminicola. Biotechnol. Appl. Biochem. 2013, 60, 52–64. [Google Scholar] [CrossRef] [PubMed]
- Lamb, D.C.; Skaug, T.; Song, H.-L.; Jackson, C.J.; Podust, L.M.; Waterman, M.R.; Kell, D.B.; Kelly, D.E.; Kelly, S.L. The cytochrome P450 complement (CYPome) of Streptomyces coelicolor A3(2). J. Biol. Chem. 2002, 277, 24000–24005. [Google Scholar] [CrossRef] [PubMed]
- Lamb, D.C.; Ikeda, H.; Nelson, D.R.; Ishikawa, J.; Skaug, T.; Jackson, C.; Omura, S.; Waterman, M.R.; Kelly, S.L. Cytochrome P450 complement (CYPome) of the avermectin-producer Streptomyces avermitilis and comparison to that of Streptomyces coelicolor A3(2). Biochem. Biophys. Res. Commun. 2003, 307, 610–619. [Google Scholar] [CrossRef]
- Kelly, D.E.; Kraševec, N.; Mullins, J.; Nelson, D.R. The CYPome (cytochrome P450 complement) of Aspergillus nidulans. Fungal Genet. Biol. 2009, 46, S53–S61. [Google Scholar] [CrossRef] [PubMed]
- Dejong, C.A.; Wilson, J.Y. The cytochrome P450 superfamily Complement (CYPome) in the annelid Capitella teleta. PLoS ONE 2014, 9, e107728. [Google Scholar] [CrossRef] [PubMed]
- Moktali, V.; Park, J.; Fedorova-Abrams, N.D.; Park, B.; Choi, J.; Lee, Y.-H.; Kang, S. Systematic and searchable classification of cytochrome P450 proteins encoded by fungal and oomycete genomes. BMC Genom. 2012, 13, 525. [Google Scholar] [CrossRef] [PubMed]
- Hirosue, S.; Tazaki, M.; Hiratsuka, N.; Yanai, S.; Kabumoto, H.; Shinkyo, R.; Arisawa, A.; Sakaki, T.; Tsunekawa, H.; Johdo, O.; et al. Insight into functional diversity of cytochrome P450 in the white-rot basidiomycete Phanerochaete chrysosporium: Involvement of versatile monooxygenase. Biochem. Biophys. Res. Commun. 2011, 407, 118–123. [Google Scholar] [CrossRef] [PubMed]
- Olicón-Hernández, D.R.; González-López, J.; Aranda, E. Overview on the biochemical potential of filamentous fungi to degrade pharmaceutical compounds. Front. Microbiol. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Warrilow, A.; Ugochukwu, C.; Lamb, D.; Kelly, D.; Kelly, S. Expression and characterization of CYP51, the ancient sterol 14-demethylase activity for cytochromes P450 (CYP), in the white-rot fungus Phanerochaete chrysosporium. Lipids 2008, 43, 1143–1153. [Google Scholar] [CrossRef] [PubMed]
- Aoyama, Y.; Yoshida, Y.; Sonoda, Y.; Sato, Y. Deformylation of 32-oxo-24,25-dihydrolanosterol by the purified cytochrome P-45014DM (lanosterol 14α-demethylase) from yeast evidence confirming the intermediate step of lanosterol 14a-demethylation. J. Biol. Chem. 1989, 264, 18502–18505. [Google Scholar] [PubMed]
- van Nistelrooy, J.G.M.; van den Brink, J.M.; van Kan, J.A.L.; van Gorcom, R.F.M.; de Waard, M.A. Isolation and molecular characterisation of the gene encoding eburicol 14α-demethylase (CYP51) from Penicillium italicum. Mol. Gen. Genet. 1996, 250, 725–733. [Google Scholar] [PubMed]
- Kalb, V.; Loper, J.; Dey, C.; Woods, C.; Sutter, T. Isolation of a cytochrome P-450 structural gene from Saccharomyces cerevisiae. Gene 1986, 45, 237–245. [Google Scholar] [CrossRef]
- Kalb, V.F.; Woods, C.W.; Turi, T.G.; Dey, C.R.; Sutter, T.R.; Loper, J.C. Primary structure of the P450 lanosterol demethylase gene from Saccharomyces cerevisiae. DNA 1987, 6, 529–537. [Google Scholar] [CrossRef] [PubMed]
- Lai, M.; Kirsch, D. Nucleotide sequence of cytochrome P450 L1A1 (lanosterol 14α-demethylase) from Candida albicans. Nucleic Acids Res. 1989, 17, 804. [Google Scholar] [CrossRef] [PubMed]
- Morace, G.; Sanguinetti, M.; Posteraro, B.; Cascio, G.L.; Fadda, G. Identification of various medically important Candida species in clinical specimens by PCR-restriction enzyme analysis. J. Clin. Microbiol. 1997, 35, 667–672. [Google Scholar] [PubMed]
- Burgener-Kairuz, P.; Zuber, J.; Jaunin, P.; Buchman, T.; Bille, J.; Rossier, M. Rapid detection and identification of Candida albicans and Torulopsis (Candida) glabrata in clinical specimens by species-specific nested PCR amplification of a cytochrome P-450 lanosterol-α-demethylase (L1A1) gene fragment. J. Clin. Microbiol. 1994, 32, 1902–1907. [Google Scholar] [PubMed]
- Aoyama, Y.; Yoshida, Y.; Sato, R. Yeast cytochrome P-450 catalyzing lanosterol 14α-demethylation. J. Biol. Chem. 1984, 259, 1661–1666. [Google Scholar] [PubMed]
- Sheehan, D.J.; Hitchcock, C.A.; Sibley, C.M. Current and emerging azole antifungal agents. Clin. Microbiol. Rev. 1999, 12, 40–79. [Google Scholar] [PubMed]
- Chen, C.; Turi, T.G.; Sanglard, D.; Loper, J.C. Isolation of the Candida tropicalis gene for P450 lanosterol demethylase and its expression in Saccharomyces cerevisiae. Biochem. Biophys. Res. Commun. 1987, 146, 1311–1317. [Google Scholar] [CrossRef]
- Ohkuma, M.; Masuda, Y.; Mee Park, S.; Ohtomo, R.; Ohta, A.; Takagi, M. Evidence that the expression of the gene for NADPH-cytochrome P-450 reductase is n-alkane-inducible in Candida maltosa. Biosci. Biotechnol. Biochem. 1995, 59, 1328–1330. [Google Scholar] [CrossRef] [PubMed]
- Schunck, W.; Vogel, F.; Gross, B.; Kärgel, E.; Mauersberger, S.; Köpke, K.; Gengnagel, C.; Müller, H. Comparison of two cytochromes P-450 from Candida maltosa: Primary structures, substrate specificities and effects of their expression in Saccharomyces cerevisiae on the proliferation of the endoplasmic reticulum. Eur. J. Cell Biol. 1991, 55, 336–345. [Google Scholar] [PubMed]
- Seghezzi, W.; Sanglard, D.; Fiechter, A. Characterization of a second alkane-inducible cytochrome P450-encoding gene, CYP52A2, from Candida tropicalis. Gene 1991, 106, 51–60. [Google Scholar] [CrossRef]
- Seghezzi, W.; Meili, C.; Ruffiner, R.; Kuenzi, R.; Sanglard, D.; Fiechter, A. Identification and characterization of additional members of the cytochrome P450 multigene family CYP52 of Candida tropicalis. DNA Cell Biol. 1992, 11, 767–780. [Google Scholar] [CrossRef] [PubMed]
- Lottermoser, K.; Schunck, W.H.; Asperger, O. Cytochromes P450 of the sophorose lipid-producing yeast Candida apicola: Heterogeneity and polymerase chain reaction-mediated cloning of two genes. Yeast 1996, 12, 565–575. [Google Scholar] [CrossRef]
- Eschenfeldt, W.H.; Zhang, Y.; Samaha, H.; Stols, L.; Eirich, L.D.; Wilson, C.R.; Donnelly, M.I. Transformation of fatty acids catalyzed by cytochrome P450 monooxygenase enzymes of Candida tropicalis. Appl. Environ. Microbiol. 2003, 69, 5992–5999. [Google Scholar] [CrossRef] [PubMed]
- Scheller, U.; Zimmer, T.; Kärgel, E.; Schunck, W.-H. Characterization of the n-alkane and fatty acid hydroxylating cytochrome P450 forms 52A3 and 52A4. Arch. Biochem. Biophys. 1996, 328, 245–254. [Google Scholar] [CrossRef] [PubMed]
- Iida, T.; Sumita, T.; Ohta, A.; Takagi, M. The cytochrome P450ALK multigene family of an n-alkane-assimilating yeast, Yarrowia lipolytica: Cloning and characterization of genes coding for new CYP52 family members. Yeast 2000, 16, 1077–1087. [Google Scholar] [CrossRef]
- Faber, B.W.; van Gorcom, R.F.; Duine, J.A. Purification and characterization of benzoate-para-hydroxylase, a cytochrome P450 (CYP53A1), from Aspergillus niger. Arch. Biochem. Biophys. 2001, 394, 245–254. [Google Scholar] [CrossRef] [PubMed]
- Fraser, J.A.; Davis, M.A.; Hynes, M.J. The genes gmdA, encoding an amidase, and bzuA, encoding a cytochrome P450, are required for benzamide utilization in Aspergillus nidulans. Fungal Genet. Biol. 2002, 35, 135–146. [Google Scholar] [CrossRef] [PubMed]
- Fukuda, H.; Fujii, T.; Sukita, E.; Tazaki, M.; Nagahama, S.; Ogawa, T. Reconstitution of the isobutene-forming reaction catalyzed by cytochrome P450 and P450 reductase from Rhodotorula minuta: Decarboxylation with the formation of isobutene. Biochem. Biophys. Res. Commun. 1994, 201, 516–522. [Google Scholar] [CrossRef] [PubMed]
- van Gorcom, R.F.; Boschloo, J.G.; Kuijvenhoven, A.; Lange, J.; van Vark, A.J.; Bos, C.J.; van Balken, J.A.; Pouwels, P.H.; van den Hondel, C.A. Isolation and molecular characterisation of the benzoate-para-hydroxylase gene (bphA) of Aspergillus niger: A member of a new gene family of the cytochrome P450 superfamily. Mol. Gen. Genet. 1990, 223, 192–197. [Google Scholar] [CrossRef] [PubMed]
- Podobnik, B.; Stojan, J.; Lah, L.; Krasevec, N.; Seliskar, M.; Rizner, T.L.; Rozman, D.; Komel, R. CYP53A15 of Cochliobolus lunatus, a target for natural antifungal compounds. J. Med. Chem. 2008, 51, 3480–3486. [Google Scholar] [CrossRef] [PubMed]
- Matsuzaki, F.; Wariishi, H. Molecular characterization of cytochrome P450 catalyzing hydroxylation of benzoates from the white-rot fungus Phanerochaete chrysosporium. Biochem. Biophys. Res. Commun. 2005, 334, 1184–1190. [Google Scholar] [CrossRef] [PubMed]
- Attar, R.; Grotewold, E.; Taccioli, G.; Aisemberg, G.; Torres, H.; Judewicz, N. A cycloheximide-inducible gene of Neurospora crassa belongs to the cytochrome P-450 superfamily. Nucleic Acids Res. 1989, 17, 7535–7536. [Google Scholar] [CrossRef] [PubMed]
- Kizawa, H.; Tomura, D.; Oda, M.; Fukamizu, A.; Hoshino, T.; Gotoh, O.; Yasui, T.; Shoun, H. Nucleotide sequence of the unique nitrate/nitrite-inducible cytochrome P-450 cDNA from Fusarium oxysporum. J. Biol. Chem. 1991, 266, 10632–10637. [Google Scholar] [PubMed]
- Kudo, T.; Tomura, D.; Liu, D.; Dai, X.; Shoun, H. Two isozymes of P450nor of Cylindrocarpon tonkinense: Molecular cloning of the cDNAs and genes, expressions in the yeast, and the putative NAD(P)H-binding site. Biochimie 1996, 78, 792–799. [Google Scholar] [CrossRef]
- Usuda, K.; Toritsuka, N.; Matsuo, Y.; Kim, D.-H.; Shoun, H. Denitrification by the fungus Cylindrocarpon tonkinense: Anaerobic cell growth and two isozyme forms of cytochrome P-450nor. Appl. Environ. Microbiol. 1995, 61, 883–889. [Google Scholar] [PubMed]
- Kaya, M.; Matsumura, K.; Higashida, K.; Hata, Y.; Kawato, A.; Abe, Y.; Akita, O.; Takaya, N.; Shoun, H. Cloning and enhanced expression of the cytochrome P450nor gene (nicA; CYP55A5) encoding nitric oxide reductase from Aspergillus oryzae. Biosci. Biotechnol. Biochem. 2004, 68, 2040–2049. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Takaya, N.; Kitazume, T.; Kondo, T.; Shoun, H. Purification and cDNA cloning of nitric oxide reductase cytochrome P450nor (CYP55A4) from Trichosporon cutaneum. Eur. J. Biochem. 2001, 268, 3198–3204. [Google Scholar] [CrossRef] [PubMed]
- Shiro, Y.; Fujii, M.; Iizuka, T.; Adachi, S.-I.; Tsukamoto, K.; Nakahara, K.; Shoun, H. Spectroscopic and kinetic studies on reaction of cytochrome P450nor with nitric oxide implication for its nitric oxide reduction mechanism. J. Biol. Chem. 1995, 270, 1617–1623. [Google Scholar] [CrossRef] [PubMed]
- Briza, P.; Breitenbach, M.; Ellinger, A.; Segall, J. Isolation of two developmentally regulated genes involved in spore wall maturation in Saccharomyces cerevisiae. Genes Dev. 1990, 4, 1775–1789. [Google Scholar] [CrossRef] [PubMed]
- Briza, P.; Eckerstorfer, M.; Breitenbach, M. The sporulation-specific enzymes encoded by the DIT1 and DIT2 genes catalyze a two-step reaction leading to a soluble LL-dityrosine-containing precursor of the yeast spore wall. Proc. Natl. Acad. Sci. USA 1994, 91, 4524–4528. [Google Scholar] [CrossRef] [PubMed]
- Melo, N.; Moran, G.; Warrilow, A.; Dudley, E.; Smith, S.; Sullivan, D.; Lamb, D.; Kelly, D.; Coleman, D.; Kelly, S. CYP56 (Dit2p) in Candida albicans: Characterization and investigation of its role in growth and antifungal drug susceptibility. Antimicrob. Agents Chemother. 2008, 52, 3718–3724. [Google Scholar] [CrossRef] [PubMed]
- Weltring, K.-M.; Turgeon, B.G.; Yoder, O.; VanEtten, H.D. Isolation of a phytoalexin-detoxification gene from the plant pathogenic fungus Nectria haematococca by detecting its expression in Aspergillus nidulans. Gene 1988, 68, 335–344. [Google Scholar] [CrossRef]
- Maloney, A.P.; VanEtten, H.D. A gene from the fungal plant pathogen Nectria haematococca that encodes the phytoalexin-detoxifying enzyme pisatin demethylase defines a new cytochrome P450 family. Mol. Gen. Genet. 1994, 243, 506–514. [Google Scholar] [CrossRef] [PubMed]
- Funnell, D.L.; VanEtten, H.D. Pisatin demethylase genes are on dispensable chromosomes while genes for pathogenicity on carrot and ripe tomato are on other chromosomes in Nectria haematococca. Mol. Plant Microbe Interact. 2002, 15, 840–846. [Google Scholar] [CrossRef] [PubMed]
- Hohn, T.M.; Desjardins, A.E.; McCormick, S.P. The Tri4 gene of Fusarium sporotrichioides encodes a cytochrome P450 monooxygenase involved in trichothecene biosynthesis. Mol. Gen. Genet. 1995, 248, 95–102. [Google Scholar] [CrossRef] [PubMed]
- Kimura, M.; Tokai, T.; Takahashi-Ando, N.; Ohsato, S.; Fujimura, M. Molecular and genetic studies of Fusarium trichothecene biosynthesis: Pathways, genes, and evolution. Biosci. Biotechnol. Biochem. 2007, 71, 2105–2123. [Google Scholar] [CrossRef] [PubMed]
- Ehrlich, K.C.; Chang, P.-K.; Yu, J.; Cotty, P.J. Aflatoxin biosynthesis cluster gene cypA is required for G aflatoxin formation. Appl. Environ. Microbiol. 2004, 70, 6518–6524. [Google Scholar] [CrossRef] [PubMed]
- Bhatnagar, D.; Ehrlich, K.; Cleveland, T. Molecular genetic analysis and regulation of aflatoxin biosynthesis. Appl. Microbiol. Biotechnol. 2003, 61, 83–93. [Google Scholar] [CrossRef] [PubMed]
- Wen, Y.; Hatabayashi, H.; Arai, H.; Kitamoto, H.K.; Yabe, K. Function of the cypX and moxY genes in aflatoxin biosynthesis in Aspergillus parasiticus. Appl. Environ. Microbiol. 2005, 71, 3192–3198. [Google Scholar] [CrossRef] [PubMed]
- Keller, N.P.; Kantz, N.J.; Adams, T.H. Aspergillus nidulans verA is required for production of the mycotoxin sterigmatocystin. Appl. Environ. Microbiol. 1994, 60, 1444–1450. [Google Scholar] [PubMed]
- Brown, D.; Yu, J.; Kelkar, H.; Fernandes, M.; Nesbitt, T.; Keller, N.; Adams, T.; Leonard, T. Twenty-five coregulated transcripts define a sterigmatocystin gene cluster in Aspergillus nidulans. Proc. Natl. Acad. Sci. USA 1996, 93, 1418–1422. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Chang, P.-K.; Cary, J.W.; Wright, M.; Bhatnagar, D.; Cleveland, T.E.; Payne, G.A.; Linz, J.E. Comparative mapping of aflatoxin pathway gene clusters in Aspergillus parasiticus and Aspergillus flavus. Appl. Environ. Microbiol. 1995, 61, 2365–2371. [Google Scholar] [PubMed]
- Lamb, D.C.; Maspahy, S.; Kelly, D.E.; Manning, N.J.; Geber, A.; Bennett, J.E.; Kelly, S.L. Purification, reconstitution, and inhibition of cytochrome P-450 sterol Δ22-desaturase from the pathogenic fungus Candida glabrata. Antimicrob. Agents Chemother. 1999, 43, 1725–1728. [Google Scholar] [PubMed]
- Doddapaneni, H.; Subramanian, V.; Yadav, J.S. Physiological regulation, xenobiotic induction, and heterologous expression of P450 monooxygenase Gene pc-3 (CYP63A3), a new member of the CYP63 gene cluster in the white-rot fungus Phanerochaete chrysosporium. Curr. Microbiol. 2005, 50, 292–298. [Google Scholar] [CrossRef] [PubMed]
- Prieto, R.; Woloshuk, C. ord1, an oxidoreductase gene responsible for conversion of O-methylsterigmatocystin to aflatoxin in Aspergillus flavus. Appl. Environ. Microbiol. 1997, 63, 1661–1666. [Google Scholar] [PubMed]
- De Groot, P.W.; Schaap, P.J.; Van Griensven, L.J.; Visser, J. Isolation of developmentally regulated genes from the edible mushroom Agaricus bisporus. Microbiology 1997, 143, 1993–2001. [Google Scholar] [CrossRef] [PubMed]
- Rojas, M.C.; Hedden, P.; Gaskin, P.; Tudzynski, B. The P450–1 gene of Gibberella fujikuroi encodes a multifunctional enzyme in gibberellin biosynthesis. Proc. Natl. Acad. Sci. USA 2001, 98, 5838–5843. [Google Scholar] [CrossRef] [PubMed]
- Tudzynski, B.; Rojas, M.A.C.; Gaskin, P.; Hedden, P. The gibberellin 20-oxidase of Gibberella fujikuroi is a multifunctional monooxygenase. J. Biol. Chem. 2002, 277, 21246–21253. [Google Scholar] [CrossRef] [PubMed]
- Mingot, J.M.; Peñalva, M.A.; Fernández-Cañón, J.M. Disruption of phacA, an Aspergillus nidulans gene encoding a novel cytochrome P450 monooxygenase catalyzing phenylacetate 2-hydroxylation, results in penicillin overproduction. J. Biol. Chem. 1999, 274, 14545–14550. [Google Scholar] [CrossRef] [PubMed]
- Ferrer-Sevillano, F.; Fernández-Cañón, J.M. Novel phacB-encoded cytochrome P450 monooxygenase from Aspergillus nidulans with 3-hydroxyphenylacetate 6-hydroxylase and 3,4-dihydroxyphenylacetate 6-hydroxylase activities. Eukaryot. Cell 2007, 6, 514–520. [Google Scholar] [CrossRef] [PubMed]
- Nakayama, N.; Takemae, A.; Shoun, H. Cytochrome P450foxy, a catalytically self-sufficient fatty acid hydroxylase of the fungus Fusarium oxysporum. J. Biochem. 1996, 119, 435–440. [Google Scholar] [CrossRef] [PubMed]
- Kitazume, T.; Tanaka, A.; Takaya, N.; Nakamura, A.; Matsuyama, S.; Suzuki, T.; Shoun, H. Kinetic analysis of hydroxylation of saturated fatty acids by recombinant P450foxy produced by an Escherichia coli expression system. Eur. J. Biochem. 2002, 269, 2075–2082. [Google Scholar] [CrossRef] [PubMed]
- Butchko, R.A.; Plattner, R.D.; Proctor, R.H. Deletion analysis of FUM genes involved in tricarballylic ester formation during fumonisin biosynthesis. J. Agric. Food Chem. 2006, 54, 9398–9404. [Google Scholar] [CrossRef] [PubMed]
- Bojja, R.S.; Cerny, R.L.; Proctor, R.H.; Du, L. Determining the biosynthetic sequence in the early steps of the fumonisin pathway by use of three gene-disruption mutants of Fusarium verticillioides. J. Agric. Food Chem. 2004, 52, 2855–2860. [Google Scholar] [CrossRef] [PubMed]
- The UniProt Consortium. UniProt: The universal protein knowledgebase. Nucl. Acids Res. 2017, 45, D158–D169. [Google Scholar]
- Gao, D.; Du, L.; Yang, J.; Wu, W.-M.; Liang, H. A critical review of the application of white rot fungus to environmental pollution control. Crit. Rev. Biotechnol. 2010, 30, 70–77. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Viraraghavan, T. Fungal decolorization of dye wastewaters: A review. Bioresour. Technol. 2001, 79, 251–262. [Google Scholar] [CrossRef]
- Evelin, H.; Kapoor, R.; Giri, B. Arbuscular mycorrhizal fungi in alleviation of salt stress: A review. Ann. Bot. 2009, 104, 1263–1280. [Google Scholar] [CrossRef] [PubMed]
- Casadevall, A. Determinants of virulence in the pathogenic fungi. Fungal Biol. Rev. 2007, 21, 130–132. [Google Scholar] [CrossRef] [PubMed]
- Desjardins, A.E.; Proctor, R.H. Molecular biology of Fusarium mycotoxins. Int. J. Food Microbiol. 2007, 119, 47–50. [Google Scholar] [CrossRef] [PubMed]
- Moss, M.O. Mycotoxin review—1. Aspergillus and Penicillium. Mycologist 2002, 16, 116–119. [Google Scholar]
- Maragos, C.; Busman, M. Rapid and advanced tools for mycotoxin analysis: A review. Food Addit. Contam. Part A Chem. Anal. Control Expo. Risk Assess. 2010, 27, 688–700. [Google Scholar] [CrossRef] [PubMed]
- Hannemann, F.; Bichet, A.; Ewen, K.M.; Bernhardt, R. Cytochrome P450 systems-biological variations of electron transport chains. Biochim. Biophys. Acta 2007, 1770, 330–344. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Chang, P.-K.; Ehrlich, K.C.; Cary, J.W.; Bhatnagar, D.; Cleveland, T.E.; Payne, G.A.; Linz, J.E.; Woloshuk, C.P.; Bennett, J.W. Clustered pathway genes in aflatoxin biosynthesis. Appl. Environ. Microbiol. 2004, 70, 1253–1262. [Google Scholar] [CrossRef] [PubMed]
- Takaoka, S.; Kurata, M.; Harimoto, Y.; Hatta, R.; Yamamoto, M.; Akimitsu, K.; Tsuge, T. Complex regulation of secondary metabolism controlling pathogenicity in the phytopathogenic fungus Alternaria alternata. New Phytol. 2014, 202, 1297–1309. [Google Scholar] [CrossRef] [PubMed]
- Ruswandi, S.; Kitani, K.; Akimitsu, K.; Tsuge, T.; Shiraishi, T.; Yamamoto, M. Structural analysis of cosmid clone pcAFT-2 carrying AFT10-1 encoding an acyl-CoA dehydrogenase involved in AF-toxin production in the strawberry pathotype of Alternaria alternata. J. Gen. Plant Pathol. 2005, 71, 107–116. [Google Scholar] [CrossRef]
- Deighton, N.; Muckenschnabel, I.; Colmenares, A.J.; Collado, I.G.; Williamson, B. Botrydial is produced in plant tissues infected by Botrytis cinerea. Phytochemistry 2001, 57, 689–692. [Google Scholar] [CrossRef]
- Siewers, V.; Viaud, M.; Jimenez-Teja, D.; Collado, I.G.; Gronover, C.S.; Pradier, J.-M.; Tudzynsk, B.; Tudzynski, P. Functional analysis of the cytochrome P450 monooxygenase gene bcbot1 of Botrytis cinerea indicates that botrydial is a strain-specific virulence factor. Mol. Plant Microbe Interact. 2005, 18, 602–612. [Google Scholar] [CrossRef] [PubMed]
- Collado, I.G.; Sánchez, A.J.M.; Hanson, J.R. Fungal terpene metabolites: Biosynthetic relationships and the control of the phytopathogenic fungus Botrytis cinerea. Nat. Prod. Rep. 2007, 24, 674–686. [Google Scholar] [CrossRef] [PubMed]
- Wight, W.D.; Kim, K.-H.; Lawrence, C.B.; Walton, J.D. Biosynthesis and role in virulence of the histone deacetylase inhibitor depudecin from Alternaria brassicicola. Mol. Plant Microbe Interact. 2009, 22, 1258–1267. [Google Scholar] [CrossRef] [PubMed]
- Chettri, P.; Ehrlich, K.C.; Cary, J.W.; Collemare, J.; Cox, M.P.; Griffiths, S.A.; Olson, M.A.; de Wit, P.J.; Bradshaw, R.E. Dothistromin genes at multiple separate loci are regulated by AflR. Fungal Genet. Biol. 2013, 51, 12–20. [Google Scholar] [CrossRef] [PubMed]
- Wallwey, C.; Li, S.-M. Ergot alkaloids: Structure diversity, biosynthetic gene clusters and functional proof of biosynthetic genes. Nat. Prod. Rep. 2011, 28, 496–510. [Google Scholar] [CrossRef] [PubMed]
- Coyle, C.M.; Panaccione, D.G. An ergot alkaloid biosynthesis gene and clustered hypothetical genes from Aspergillus fumigatus. Appl. Environ. Microbiol. 2005, 71, 3112–3118. [Google Scholar] [CrossRef] [PubMed]
- Seo, J.-A.; Proctor, R.H.; Plattner, R.D. Characterization of four clustered and coregulated genes associated with fumonisin biosynthesis in Fusarium verticillioides. Fungal Genet. Biol. 2001, 34, 155–165. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.-Q.; Ahn, J.-H.; Walton, J.D. A putative branched-chain-amino-acid transaminase gene required for HC-toxin biosynthesis and pathogenicity in Cochliobolus carbonum. Microbiology 1999, 145, 3539–3546. [Google Scholar] [CrossRef] [PubMed]
- Walton, J.D. HC-toxin. Phytochemistry 2006, 67, 1406–1413. [Google Scholar] [CrossRef] [PubMed]
- Paterson, R.R.M.; Lima, N. Molecular Biology of Food and Water Borne Mycotoxigenic and Mycotic Fungi; CRC Press: Boca Raton, FL, USA, 2015; ISBN 1466559888. [Google Scholar]
- McMillan, L.; Carr, R.; Young, C.; Astin, J.; Lowe, R.; Parker, E.; Jameson, G.; Finch, S.; Miles, C.; McManus, O. Molecular analysis of two cytochrome P450 monooxygenase genes required for paxilline biosynthesis in Penicillium paxilli, and effects of paxilline intermediates on mammalian maxi-K ion channels. Mol. Genet. Genom. 2003, 270, 9–23. [Google Scholar] [CrossRef] [PubMed]
- Saikia, S.; Parker, E.J.; Koulman, A.; Scott, B. Defining paxilline biosynthesis in Penicillium paxilli. J. Biol. Chem. 2007, 282, 16829–16837. [Google Scholar] [CrossRef] [PubMed]
- Hidalgo, P.I.; Ullán, R.V.; Albillos, S.M.; Montero, O.; Fernández-Bodega, M.Á.; García-Estrada, C.; Fernández-Aguado, M.; Martín, J.-F. Molecular characterization of the PR-toxin gene cluster in Penicillium roqueforti and Penicillium chrysogenum: Cross talk of secondary metabolite pathways. Fungal Genet. Biol. 2014, 62, 11–24. [Google Scholar] [CrossRef] [PubMed]
- Alexander, N.J.; Hohn, T.M.; McCormick, S.P. The TRI11 gene of Fusarium sporotrichioides encodes a cytochrome P-450 monooxygenase required for C-15 hydroxylation in trichothecene biosynthesis. Appl. Environ. Microbiol. 1998, 64, 221–225. [Google Scholar] [PubMed]
- Kim, S.; Thiessen, P.A.; Bolton, E.E.; Chen, J.; Fu, G.; Gindulyte, A.; Han, L.; He, J.; He, S.; Shoemaker, B.A. PubChem substance and compound databases. Nucleic Acids Res. 2015, 44, D1202–D1213. [Google Scholar] [CrossRef] [PubMed]
- Brown, M.; Brown-Jenco, C.; Payne, G. Genetic and molecular analysis of aflatoxin biosynthesis. Fungal Genet. Biol. 1999, 26, 81–98. [Google Scholar] [CrossRef] [PubMed]
- Fratamico, P.M.; Bhunia, A.K.; Smith, J.L. Foodborne Pathogens: Microbiology and Molecular Biology; Horizon Scientific Press: Poole, UK, 2005; ISBN 190445500X. [Google Scholar]
- Bennett, J. Mycotoxins, mycotoxicoses, mycotoxicology and mycopathologia. Mycopathologia 1987, 100, 3–5. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Chang, P.-K.; Cary, J.W.; Bhatnagar, D.; Cleveland, T.E. avnA, a gene encoding a cytochrome P-450 monooxygenase, is involved in the conversion of averantin to averufin in aflatoxin biosynthesis in Aspergillus parasiticus. Appl. Environ. Microbiol. 1997, 63, 1349–1356. [Google Scholar] [PubMed]
- Yabe, K.; Ando, Y.; Hamasaki, T. Biosynthetic relationship among aflatoxins B1, B2, G1, and G2. Appl. Environ. Microbiol. 1988, 54, 2101–2106. [Google Scholar] [PubMed]
- Yu, J.; Chang, P.-K.; Ehrlich, K.C.; Cary, J.W.; Montalbano, B.; Dyer, J.M.; Bhatnagar, D.; Cleveland, T.E. Characterization of the critical amino acids of an Aspergillus parasiticus cytochrome P-450 monooxygenase encoded by ordA that is involved in the biosynthesis of aflatoxins B1, G1, B2, and G2. Appl. Environ. Microbiol. 1998, 64, 4834–4841. [Google Scholar] [PubMed]
- Proctor, R.H.; Desjardins, A.E.; Plattner, R.D.; Hohn, T.M. A polyketide synthase gene required for biosynthesis of fumonisin mycotoxins in Gibberella fujikuroi mating population A. Fungal Genet. Biol. 1999, 27, 100–112. [Google Scholar] [CrossRef] [PubMed]
- Howard, P.C.; Eppley, R.M.; Stack, M.E.; Warbritton, A.; Voss, K.A.; Lorentzen, R.J.; Kovach, R.; Bucci, T.J. Carcinogenicity of fumonisin B1 in a two-year bioassay with Fischer 344 rats and B6C3F1 mice. JSM Mycotoxins 2000, 1999, 45–54. [Google Scholar] [CrossRef]
- Caldas, E.D.; Sadilkova, K.; Ward, B.L.; Jones, A.D.; Winter, C.K.; Gilchrist, D.G. Biosynthetic studies of fumonisin B1 and AAL toxins. J. Agric. Food Chem. 1998, 46, 4734–4743. [Google Scholar] [CrossRef]
- Walton, J.D. Host-selective toxins: Agents of compatibility. Plant Cell 1996, 8, 1723–1733. [Google Scholar] [CrossRef] [PubMed]
- Friesen, T.L.; Faris, J.D.; Solomon, P.S.; Oliver, R.P. Host-specific toxins: Effectors of necrotrophic pathogenicity. Cell. Microbiol. 2008, 10, 1421–1428. [Google Scholar] [CrossRef] [PubMed]
- Akimitsu, K.; Tsuge, T.; Kodama, M.; Yamamoto, M.; Otani, H. Alternaria host-selective toxins: Determinant factors of plant disease. J. Gen. Plant Pathol. 2014, 80, 109–122. [Google Scholar] [CrossRef]
- Brosch, G.; Ransom, R.; Lechner, T.; Walton, J.D.; Loidl, P. Inhibition of maize histone deacetylases by HC toxin, the host-selective toxin of Cochliobolus carbonum. Plant Cell 1995, 7, 1941–1950. [Google Scholar] [CrossRef] [PubMed]
- Youngman, R.J.; Elstner, E. Photodynamic and reductive mechanisms of oxygen activation by the fungal phytotoxins, cercosporin and dothistromin. In Oxygen Radicals in Chemistry and Biology, Proceedings/Third International Conference, Neuherberg, Germany, 10–15 July 1983; Bors, W., Saran, M., Tait, D., Eds.; W. de Gruyter: Berlin, Germany, 1984. [Google Scholar]
- Bradshaw, R. Dothistroma (red-band) needle blight of pines and the dothistromin toxin: A review. For. Pathol. 2004, 34, 163–185. [Google Scholar] [CrossRef]
- Kabir, M.; Ganley, R.; Bradshaw, R. Dothistromin toxin is a virulence factor in dothistroma needle blight of pines. Plant Pathol. 2015, 64, 225–234. [Google Scholar] [CrossRef]
- Govrin, E.M.; Levine, A. The hypersensitive response facilitates plant infection by the necrotrophic pathogen Botrytis cinerea. Curr. Biol. 2000, 10, 751–757. [Google Scholar] [CrossRef]
- Collado, I.; Aleu, J.; Hernández-Galán, R.; Durán-Patrón, R. Botrytis species an intriguing source of metabolites with a wide range of biological activities. Structure, chemistry and bioactivity of metabolites isolated from Botrytis species. Curr. Org. Chem. 2000, 4, 1261–1286. [Google Scholar] [CrossRef]
- Varga, J.; Rigó, K.; Téren, J.; Mesterházy, Á. Recent advances in ochratoxin research I. Production, detection and occurrence of ochratoxins. Cereal Res. Commun. 2001, 29, 85–92. [Google Scholar]
- International Agency for Research on Cancer (IARC). IARC Monographs on the evaluation of carcinogenic risks to humans: Some naturally. IARC Monogr. Eval. Carcinog. Risks Hum. 1993, 56, 489–521. [Google Scholar]
- Gil-Serna, J.; García-Díaz, M.; González-Jaén, M.T.; Vázquez, C.; Patiño, B. Description of an orthologous cluster of ochratoxin A biosynthetic genes in Aspergillus and Penicillium species: A comparative analysis. Int. J. Food Microbiol. 2018, 268, 35–43. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Yu, F.; Schnabel, G.; Wu, J.; Wang, Z.; Ma, Z. Paralogous cyp51 genes in Fusarium graminearum mediate differential sensitivity to sterol demethylation inhibitors. Fungal Genet. Biol. 2011, 48, 113–123. [Google Scholar] [CrossRef] [PubMed]
- Anzenbacher, P.; Anzenbacherova, E. Cytochromes P450 and metabolism of xenobiotics. Cell. Mol. Life Sci. 2001, 58, 737–747. [Google Scholar] [CrossRef] [PubMed]
- Furge, L.L.; Guengerich, F.P. Cytochrome P450 enzymes in drug metabolism and chemical toxicology: An introduction. Biochem. Mol. Biol. Educ. 2006, 34, 66–74. [Google Scholar] [CrossRef] [PubMed]
- Danielson, P.B.; MacIntyre, R.J.; Fogleman, J.C. Molecular cloning of a family of xenobiotic-inducible drosophilid cytochrome P450s: Evidence for involvement in host-plant allelochemical resistance. Proc. Natl. Acad. Sci. USA 1997, 94, 10797–10802. [Google Scholar] [CrossRef] [PubMed]
- Ding, M.; Gao, Q.; Mo, J.; Cheng, J.A. Construction and validation of an insecticide resistance-associated DNA microarray. J. Pestic. Sci. 2007, 32, 32–41. [Google Scholar] [CrossRef]
- Syed, K.; Yadav, J.S. P450 monooxygenases (P450ome) of the model white rot fungus Phanerochaete chrysosporium. Crit. Rev. Microbiol. 2012, 38, 339–363. [Google Scholar] [CrossRef] [PubMed]
- Ichinose, H. Cytochrome P450 of wood-rotting basidiomycetes and biotechnological applications. Biotechnol. Appl. Biochem. 2013, 60, 71–81. [Google Scholar] [CrossRef] [PubMed]
- Matthews, D.E.; Van Etten, H.D. Detoxification of the phytoalexin pisatin by a fungal cytochrome P-450. Arch. Biochem. Biophys. 1983, 224, 494–505. [Google Scholar] [CrossRef]
- Coleman, J.J.; Wasmann, C.C.; Usami, T.; White, G.J.; Temporini, E.D.; McCluskey, K.; VanEtten, H.D. Characterization of the gene encoding pisatin demethylase (FoPDA1) in Fusarium oxysporum. Mol. Plant Microbe Interact. 2011, 24, 1482–1491. [Google Scholar] [CrossRef] [PubMed]
- Parks, L.W.; Smith, S.J.; Crowley, J.H. Biochemical and physiological effects of sterol alterations in yeast—A review. Lipids 1995, 30, 227–230. [Google Scholar] [CrossRef] [PubMed]
- Lepesheva, G.I.; Waterman, M.R. Sterol 14α-demethylase cytochrome P450 (CYP51), a P450 in all biological kingdoms. Biochim. Biophys. Acta 2007, 1770, 467–477. [Google Scholar] [CrossRef] [PubMed]
- Mellado, E.; Garcia-Effron, G.; Buitrago, M.; Alcazar-Fuoli, L.; Cuenca-Estrella, M.; Rodriguez-Tudela, J. Targeted gene disruption of the 14-α sterol demethylase (cyp51A) in Aspergillus fumigatus and its role in azole drug susceptibility. Antimicrob. Agents Chemother. 2005, 49, 2536–2538. [Google Scholar] [CrossRef] [PubMed]
- Sanglard, D.; Ischer, F.; Koymans, L.; Bille, J. Amino acid substitutions in the cytochrome P-450 lanosterol 14α-demethylase (CYP51A1) from azole-resistant Candida albicans clinical isolates contribute to resistance to azole antifungal agents. Antimicrob. Agents Chemother. 1998, 42, 241–253. [Google Scholar] [PubMed]
- Sheng, C.; Miao, Z.; Ji, H.; Yao, J.; Wang, W.; Che, X.; Dong, G.; Lü, J.; Guo, W.; Zhang, W. Three-dimensional model of lanosterol 14α-demethylase from Cryptococcus neoformans: Active-site characterization and insights into azole binding. Antimicrob. Agents Chemother. 2009, 53, 3487–3495. [Google Scholar] [CrossRef] [PubMed]
- Van Bogaert, I.N.A.; Groeneboer, S.; Saerens, K.; Soetaert, W. The role of cytochrome P450 monooxygenases in microbial fatty acid metabolism. FEBS J. 2011, 278, 206–221. [Google Scholar] [CrossRef] [PubMed]
- Shoun, H.; Fushinobu, S.; Jiang, L.; Kim, S.-W.; Wakagi, T. Fungal denitrification and nitric oxide reductase cytochrome P450nor. Philos. Trans. R. Soc. B 2012, 367, 1186–1194. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.-W.; Jia, L.-J.; Zhang, Y.; Jiang, G.; Li, X.; Zhang, D.; Tang, W.-H. In planta stage-specific fungal gene profiling elucidates the molecular strategies of Fusarium graminearum growing inside wheat coleoptiles. Plant Cell 2012, 24, 5159–5176. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Li, Y.; Gao, J.; Suo, Y.; Fu, K.; Li, Y.; Chen, J. Genome sequence and virulence variation-related transcriptome profiles of Curvularia lunata, an important maize pathogenic fungus. BMC Genom. 2014, 15, 627. [Google Scholar] [CrossRef] [PubMed]
- Karlsson, M.; Elfstrand, M.; Stenlid, J.; Olson, Å. A fungal cytochrome P450 is expressed during the interaction between the fungal pathogen Heterobasidion annosum sensu lato and conifer trees. DNA Seq. 2008, 19, 115–120. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.-D.; Wang, X.-Y.; Chen, J.-Y.; Kong, Z.-Q.; Gui, Y.-J.; Li, N.-Y.; Bao, Y.-M.; Dai, X.-F. Identification and characterization of a pathogenicity-related gene VdCYP1 from Verticillium dahliae. Sci. Rep. 2016, 6, 27979. [Google Scholar] [CrossRef] [PubMed]
- Črešnar, B.; Petrič, Š. Cytochrome P450 enzymes in the fungal kingdom. Biochim. Biophys. Acta 2011, 1814, 29–35. [Google Scholar] [CrossRef] [PubMed]
- Leslie, J.F.; Summerell, B.A. The Fusarium Laboratory Manual, 1st ed.; Blackwell Pub.: Ames, Iowa, 2006; pp. 12, 388. ISBN 9780813819198. [Google Scholar]
- Son, H.; Seo, Y.-S.; Min, K.; Park, A.R.; Lee, J.; Jin, J.-M.; Lin, Y.; Cao, P.; Hong, S.-Y.; Kim, E.-K.; et al. A phenome-based functional analysis of transcription factors in the cereal head blight fungus, Fusarium graminearum. PLoS Pathog. 2011, 7, e1002310. [Google Scholar] [CrossRef] [PubMed]
- Brown, D.W.; McCormick, S.P.; Alexander, N.J.; Proctor, R.H.; Desjardins, A.E. A genetic and biochemical approach to study trichothecene diversity in Fusarium sporotrichioides and Fusarium graminearum. Fungal Genet. Biol. 2001, 32, 121–133. [Google Scholar] [CrossRef] [PubMed]
- Harris, L.J.; Alexander, N.J.; Saparno, A.; Blackwell, B.; McCormick, S.P.; Desjardins, A.E.; Robert, L.S.; Tinker, N.; Hattori, J.; Piché, C.; et al. A novel gene cluster in Fusarium graminearum contains a gene that contributes to butenolide synthesis. Fungal Genet. Biol. 2007, 44, 293–306. [Google Scholar] [CrossRef] [PubMed]
- Bahadoor, A.; Schneiderman, D.; Gemmill, L.; Bosnich, W.; Blackwell, B.; Melanson, J.E.; McRae, G.; Harris, L.J. Hydroxylation of longiborneol by a Clm2-encoded CYP450 monooxygenase to produce culmorin in Fusarium graminearum. J. Nat. Prod. 2016, 79, 81–88. [Google Scholar] [CrossRef] [PubMed]
- Grovey, J. The trichothecenes and their biosynthesis. In Progress in the Chemistry of Organic Natural Products; Springer: New York, NY, USA, 2007; pp. 63–130. ISBN 978-3-211-49389-2. [Google Scholar]
- Ueno, Y. Toxicological features of T-2 toxin and related trichothecenes. Fundam. Appl. Toxicol. 1984, 4, S124–S132. [Google Scholar] [CrossRef]
- Pestka, J.J.; Smolinski, A.T. Deoxynivalenol: Toxicology and potential effects on humans. J. Toxicol. Environ. Health B Crit. Rev. 2005, 8, 39–69. [Google Scholar] [CrossRef] [PubMed]
- Minervini, F.; Fornelli, F.; Flynn, K. Toxicity and apoptosis induced by the mycotoxins nivalenol, deoxynivalenol and fumonisin B1 in a human erythroleukemia cell line. Toxicol. In Vitro 2004, 18, 21–28. [Google Scholar] [CrossRef]
- Brown, D.W.; McCormick, S.P.; Alexander, N.J.; Proctor, R.H.; Desjardins, A.E. Inactivation of a cytochrome P-450 is a determinant of trichothecene diversity in Fusarium species. Fungal Genet. Biol. 2002, 36, 224–233. [Google Scholar] [CrossRef]
- Wuchiyama, J.; Kimura, M.; Yamaguchi, I. A trichothecene efflux pump encoded by Tril02 in the biosynthetic gene cluster of Fusarium graminearum. J. Antibiot. 2000, 53, 196–200. [Google Scholar] [CrossRef] [PubMed]
- Tokai, T.; Koshino, H.; Kawasaki, T.; Igawa, T.; Suzuki, Y.; Sato, M.; Fujimura, M.; Eizuka, T.; Watanabe, H.; Kitahara, T. Screening of putative oxygenase genes in the Fusarium graminearum genome sequence database for their role in trichothecene biosynthesis. FEMS Microbiol. Lett. 2005, 251, 193–201. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Zhang, S.; Hou, R.; Zhao, Z.; Zheng, Q.; Xu, Q.; Zheng, D.; Wang, G.; Liu, H.; Gao, X.; et al. Functional analysis of the kinome of the wheat scab fungus Fusarium graminearum. PLoS Pathog. 2011, 7, e1002460. [Google Scholar] [CrossRef] [PubMed]
- Fischer, G.J.; Keller, N.P. Production of cross-kingdom oxylipins by pathogenic fungi: An update on their role in development and pathogenicity. J. Microbiol. 2016, 54, 254–264. [Google Scholar] [CrossRef] [PubMed]
- Winkel-Shirley, B. Flavonoid biosynthesis. A colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol. 2001, 126, 485–493. [Google Scholar] [CrossRef] [PubMed]
- Paxton, J. A new working definition of the term “phytoalexin”. Plant Dis. 1980, 64, 734. [Google Scholar]
- Harris, L.J.; Balcerzak, M.; Johnston, A.; Schneiderman, D.; Ouellet, T. Host-preferential Fusarium graminearum gene expression during infection of wheat, barley, and maize. Fungal Biol. 2016, 120, 111–123. [Google Scholar] [CrossRef] [PubMed]
- Maertens, J. History of the development of azole derivatives. Clinic. Microbiol. Infect. 2004, 10, 1–10. [Google Scholar] [CrossRef]
| Clade 1 | Family | Class 2 | Organism | Function | Reference |
|---|---|---|---|---|---|
| 1 | CYP51 | E, group I, IV | S. cerevisiae, C. albicans, C. kefyr, C. glabrata, C. guilliermondii, C. parapsilosis, C. tropicalis, C. krusei, Ustilago maydis, Schizosaccharomyces pombe, Kluyveromyces marxianus, Penicillium italicum, Fusarium graminearum | Demethylation of eburicol/lanosterol at 14α position | [21,23,32,33,34,35,36,37,38,39,40,41,42] |
| 2 | CYP52 | E, group II | C. maltose, C. tropicalis, C. apicola | n-alkane and fatty acid assimilation | [3,42,43,44,45,46,47,48,49,50] |
| 2 | CYP53 | E, group I | Aspergillus niger, Beauveria bassiana, Cochliobolus lunatus, P. chrysosporium, Rhombophryne minuta | Degradation or detoxification benzoate and its derivatives | [51,52,53,54,55,56] |
| 2 | CYP54 | E, group I | Neurospora crassa | Cycloheximide inducible, but function is unknown | [57] |
| 3 | CYP55 | E, group I | F. oxysporum, Cylindrocarpon tonkinensis, A. oryzae, Trichosporon cutaneum | Denitrification process | [58,59,60,61,62,63] |
| 4 | CYP56 | E, group IV | S. cerevisiae, C. albicans | Formation of dityrosine | [64,65,66] |
| 6 | CYP57 | E, group I | Nectria haematococca | Pisatin detoxification | [67,68,69] |
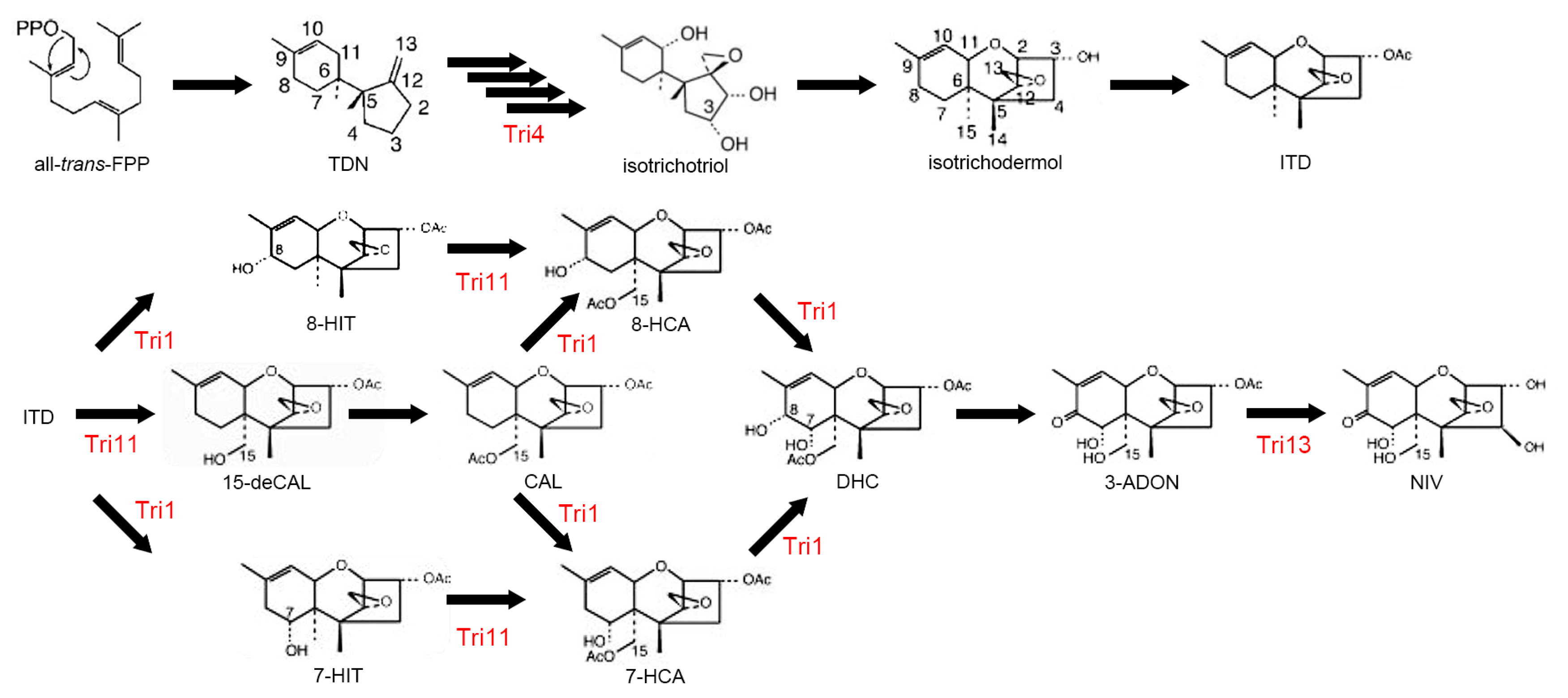
| 6 | CYP58 | E, group I | F. sporotrichioides, F. graminearum | Trichothecene biosynthesis (TRI4) | [70,71] |
| 7 | CYP58 | B | A. flavus, A. parasiticus | Aflatoxin biosynthesis | [72,73,74] |
| 8 | CYP59 | E, group I | A. nidulans | Sterigmatocystin biosynthesis (stcS/verA) | [75,76] |
| 8 | CYP60 | E, group I | A. parasiticus, A. nidulans | o-methylsterigmatocystin to aflatoxin (ord1), sterigmatocystin biosynthesis (stcF and stcL) | [76,77] |
| 8 | CYP61 | E, group I | S. cerevisiae, C. glabrata | Sterol D22-desaturase in ergosterol biosynthesis (erg5) | [22,78] |
| 8 | CYP62 | E, group I | A. nidulans | Sterigmatocystin biosynthesis (stcB) | [76] |
| 8 | CYP63 | E, group I | P. chrysosporium | Unknown function | [79] |
| 8 | CYP64 | E, group I | A. flavus | Conversion of o-methylsterigmatocystin to aflatoxin (ord1) | [80] |
| 8 | CYP65 | E, group I | F. sporotrichioides | Trichothecene biosynthesis (TRI11) | [71] |
| 9 | CYP66 | E, group IV | Agaricus bisporus | Developmental regulation of mushroom | [81] |
| 10 | CYP68, CYP69, CYP503 | E, group I | F. fujikuroi | Gibberellin biosynthesis | [82,83] |
| 10 | CYP504 | E, group I | A. nidulans | Catalyzing phenylacetate 2-hydroxylation | [84,85,86] |
| 14 | CYP505 | E, group IV | F. oxysporum | ω-1 to ω-3 carbon hydroxylation of fatty acids | [86,87] |
| 15 | CYP505 | E, group IV | F. verticillioides | Fumonisin biosynthesis | [88,89] |
| 15 | CYP526 | E, group IV | F. sporotrichioides | Trichothecene biosynthesis | [71] |
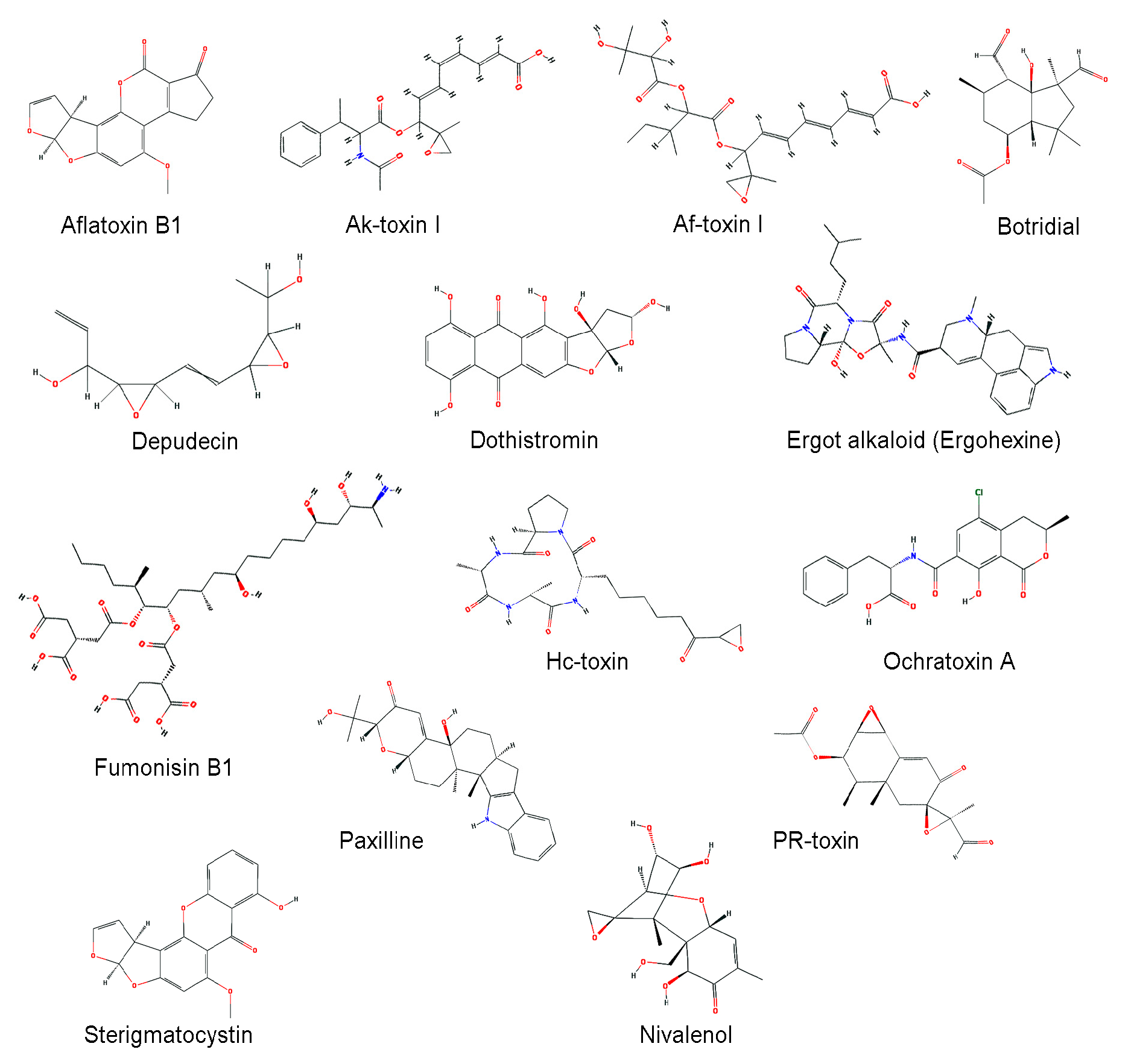
| Mycotoxin | Organism | Characteristics | Reference |
|---|---|---|---|
| Aflatoxin | A. flavus, A. parasiticus, etc. | Carcinogenic compounds posing a potential risk to livestock and human health | [99] |
| Ak-toxin | Alternaria alternata | Host-selective toxin, virulence factor to infect Japanese pear | [100] |
| Af-toxin | A. alternata | Host-selective toxin, virulence factor to infect strawberry | [101] |
| Botridial | Botrytis cinerea | Induction of chlorosis and cell collapse in plant | [102,103,104] |
| Depudecin | A. brassicicola | An inhibitor of histone deacetylase (HDAC) | [105] |
| Dothistromin | Dothistromaseptosporum | A broad-spectrum toxin that generates oxygen radicals by reductive oxygen activation | [106] |
| Ergot alkaloid | Claviceps, Penicillium, and Aspergillus spp. | A complex family of indole derivatives with diverse structures and biological activities | [107,108] |
| Fumonisin | F. verticillioides | Induction of several animal diseases, including leukoencephalomalacia, pulmonary edema, and cancer | [109] |
| Hc-toxin | Cochliobolus carbonum | An inhibitor of histone deacetylases (HDACs) in many organisms, including plants, insects, and mammals | [110,111] |
| Ochratoxin | Aspergillus, and Penicillium spp. | Possible carcinogenic | [112] |
| Paxilline | P. paxilli | A potassium channel blocker | [113,114] |
| PR-toxin | P. roqueforti | Liver toxicity and abortions in cows | [115] |
| Sterigmatocystin | A. nidulans, A. versicolor | A toxic metabolite structurally closely related to the aflatoxins | [75,76] |
| Trichothecene | F. sporotrichioides, F. graminearum | Inhibition of protein synthesis and highly cytotoxic to many eukaryotes | [71,95,116] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shin, J.; Kim, J.-E.; Lee, Y.-W.; Son, H. Fungal Cytochrome P450s and the P450 Complement (CYPome) of Fusarium graminearum. Toxins 2018, 10, 112. https://doi.org/10.3390/toxins10030112
Shin J, Kim J-E, Lee Y-W, Son H. Fungal Cytochrome P450s and the P450 Complement (CYPome) of Fusarium graminearum. Toxins. 2018; 10(3):112. https://doi.org/10.3390/toxins10030112
Chicago/Turabian StyleShin, Jiyoung, Jung-Eun Kim, Yin-Won Lee, and Hokyoung Son. 2018. "Fungal Cytochrome P450s and the P450 Complement (CYPome) of Fusarium graminearum" Toxins 10, no. 3: 112. https://doi.org/10.3390/toxins10030112





