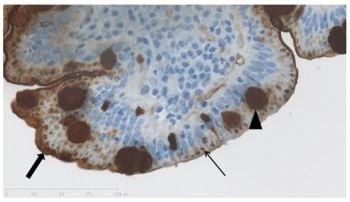
Lectin Staining Shows no Evidence of Involvement of Glycocalyx/Mucous Layer Carbohydrate Structures in Development of Celiac Disease
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Design
| Group A Patient ID | CD + T1D Untreated | CD + T1D GFD | Group B Patient ID | CD | Group C Patient ID | Non-CD |
|---|---|---|---|---|---|---|
| A1 | 3A | 0 | B1 | 2 | C1 | 0 |
| A2 | 3C | 0 | B2 | 3B | C2 | 0 |
| A3 | 2 | 0 | B3 | 3C | C3 | 0 |
| A4 | 2 | 0 | B4 | 3B | C4 | 0 |
| A5 | 3B | 0 | B5 | 3B | C5 | 0 |
| A6 | 3C | 0 | B6 | 3C | C6 | 0 |
| A7 | 3C | 0 | B7 | 3C | C7 | 0 |
| A8 | 3A | 0 | B8 | 3C | C8 | 0 |
| A9 | 3B | 0 | B9 | 3C | C9 | 0 |
| A10 | 3C | 0 | B10 | 3C | C10 | 0 |
2.2. Lectin Histochemistry
2.3. Ethics
2.4. Statistical Methods
3. Results

| Group | A CD + T1D Untreated | A CD + T1D GFD | B CD | C Non-CD | |
|---|---|---|---|---|---|
| Score | |||||
| 3 | 0 | 2 | 0 | 2 | |
| 2 | 5 | 4 | 1 | 0 | |
| 1 | 5 | 4 | 9 | 8 | |
| 0 | 0 | 0 | 0 | 0 | |
| Median score | 1.5 | 2 | 1 | 1 | |

| Group | A CD + T1D Untreated | A CD + T1D GFD | B CD | C Non-CD | |
|---|---|---|---|---|---|
| Score | |||||
| 3 | 5 | 4 | 10 | 5 | |
| 2 | 5 | 6 | 0 | 4 | |
| 1 | 0 | 0 | 0 | 0 | |
| 0 | 0 | 0 | 0 | 1 | |
| Median score | 2.5 | 2 | 3 | 2.5 | |
4. Discussion
5. Conclusions
Acknowledgments
Conflicts of Interest
References
- Abadie, V.; Sollid, L.M.; Barreiro, L.B.; Jabri, B. Integration of genetic and immunological insights into a model of celiac disease pathogenesis. Annu. Rev. Immunol. 2011, 29, 493–525. [Google Scholar] [CrossRef]
- Dubois, P.C.; Trynka, G.; Franke, L.; Hunt, K.A.; Romanos, J.; Curtotti, A.; Zhernakova, A.; Heap, G.A.; Adany, R.; Aromaa, A.; et al. Multiple common variants for celiac disease influencing immune gene expression. Nat. Genet. 2010, 42, 295–302. [Google Scholar] [CrossRef]
- Kverka, M.; Tlaskalova-Hogenova, H. Two faces of microbiota in inflammatory and autoimmune diseases: Triggers and drugs. Acta Pathol. Microbiol. Immunol. Scand. 2013, 121, 403–421. [Google Scholar]
- Le Chatelier, E.; Nielsen, T.; Qin, J.; Prifti, E.; Hildebrand, F.; Falony, G.; Almeida, M.; Arumugam, M.; Batto, J.M.; Kennedy, S.; et al. Richness of human gut microbiome correlates with metabolic markers. Nature 2013, 500, 541–546. [Google Scholar] [CrossRef]
- Robbe, C.; Capon, C.; Coddeville, B.; Michalski, J.C. Structural diversity and specific distribution of o-glycans in normal human mucins along the intestinal tract. Biochem. J. 2004, 384, 307–316. [Google Scholar] [CrossRef]
- Brinck, U.; Korabiowska, M.; Bosbach, R.; Gabius, H.J. Detection of inflammation- and neoplasia-associated alterations in human large intestine using plant/invertebrate lectins, galectin-1 and neoglycoproteins. Acta Anat. 1998, 161, 219–233. [Google Scholar] [CrossRef] [Green Version]
- Cheng, J.; Kalliomaki, M.; Heilig, H.G.; Palva, A.; Lahteenoja, H.; de Vos, W.M.; Salojarvi, J.; Satokari, R. Duodenal microbiota composition and mucosal homeostasis in pediatric celiac disease. BMC Gastroenterol. 2013, 13, 113. [Google Scholar] [CrossRef]
- Forsberg, G.; Fahlgren, A.; Horstedt, P.; Hammarstrom, S.; Hernell, O.; Hammarstrom, M.L. Presence of bacteria and innate immunity of intestinal epithelium in childhood celiac disease. Am. J. Gastroenterol. 2004, 99, 894–904. [Google Scholar] [CrossRef]
- Volta, U.; Tovoli, F.; Caio, G. Clinical and immunological features of celiac disease in patients with type 1 diabetes mellitus. Expert Rev. Gastroenterol. Hepatol. 2011, 5, 479–487. [Google Scholar] [CrossRef]
- Camarca, M.E.; Mozzillo, E.; Nugnes, R.; Zito, E.; Falco, M.; Fattorusso, V.; Mobilia, S.; Buono, P.; Valerio, G.; Troncone, R.; et al. Celiac disease in type 1 diabetes mellitus. Ital. J. Pediatr. 2012, 38, 10. [Google Scholar] [CrossRef]
- Antonioli, D.A. Celiac disease: A progress report. Mod. Pathol. 2003, 16, 342–346. [Google Scholar] [CrossRef]
- Roth, Z.; Yehezkel, G.; Khalaila, I. Identification and quantification of protein glycosylation. Int. J. Carbohydr. Chem. 2012, 2012. [Google Scholar] [CrossRef]
- GraphPad Prism, version 4.03 for Windows; GraphPad Software: San Diego, CA, USA, 2005.
- Herold, K.C.; Vignali, D.A.; Cooke, A.; Bluestone, J.A. Type 1 diabetes: Translating mechanistic observations into effective clinical outcomes. Nat. Rev. Immunol. 2013, 13, 243–256. [Google Scholar] [CrossRef]
- Vecchi, M.; Torgano, G.; de Franchis, R.; Tronconi, S.; Agape, D.; Ronchi, G. Evidence of altered structural and secretory glycoconjugates in the jejunal mucosa of patients with gluten sensitive enteropathy and subtotal villous atrophy. Gut 1989, 30, 804–810. [Google Scholar] [CrossRef]
- Barresi, G.; Tuccari, G.; Tedeschi, A.; Magazzu, G. Lectin binding sites in duodeno-jejunal mucosae from coeliac children. Histochemistry 1988, 88, 105–112. [Google Scholar] [CrossRef]
- Gronlund, M.M.; Arvilommi, H.; Kero, P.; Lehtonen, O.P.; Isolauri, E. Importance of intestinal colonisation in the maturation of humoral immunity in early infancy: A prospective follow up study of healthy infants aged 0–6 months. Arch. Dis. Childh. Fetal Neonatal Ed. 2000, 83, F186–F192. [Google Scholar] [CrossRef]
- Lanning, D.; Sethupathi, P.; Rhee, K.J.; Zhai, S.K.; Knight, K.L. Intestinal microflora and diversification of the rabbit antibody repertoire. J. Immunol. 2000, 165, 2012–2019. [Google Scholar]
- Rhee, K.J.; Sethupathi, P.; Driks, A.; Lanning, D.K.; Knight, K.L. Role of commensal bacteria in development of gut-associated lymphoid tissues and preimmune antibody repertoire. J. Immunol. 2004, 172, 1118–1124. [Google Scholar]
- Hooper, L.V.; Wong, M.H.; Thelin, A.; Hansson, L.; Falk, P.G.; Gordon, J.I. Molecular analysis of commensal host-microbial relationships in the intestine. Science 2001, 291, 881–884. [Google Scholar] [CrossRef]
- Garcia-Lafuente, A.; Antolin, M.; Guarner, F.; Crespo, E.; Malagelada, J.R. Modulation of colonic barrier function by the composition of the commensal flora in the rat. Gut 2001, 48, 503–507. [Google Scholar] [CrossRef]
- Bai, A.P.; Ouyang, Q.; Zhang, W.; Wang, C.H.; Li, S.F. Probiotics inhibit TNF-α-induced interleukin-8 secretion of HT29 cells. World J. Gastroenterol. 2004, 10, 455–457. [Google Scholar]
- Cox, M.J.; Cookson, W.O.; Moffatt, M.F. Sequencing the human microbiome in health and disease. Hum. Mol. Genet. 2013. [Google Scholar] [CrossRef]
- Benson, A.K.; Kelly, S.A.; Legge, R.; Ma, F.; Low, S.J.; Kim, J.; Zhang, M.; Oh, P.L.; Nehrenberg, D.; Hua, K.; et al. Individuality in gut microbiota composition is a complex polygenic trait shaped by multiple environmental and host genetic factors. Proc. Natl. Acad. Sci. USA 2010, 107, 18933–18938. [Google Scholar] [CrossRef]
- McKnite, A.M.; Perez-Munoz, M.E.; Lu, L.; Williams, E.G.; Brewer, S.; Andreux, P.A.; Bastiaansen, J.W.; Wang, X.; Kachman, S.D.; Auwerx, J.; et al. Murine gut microbiota is defined by host genetics and modulates variation of metabolic traits. PLoS One 2012, 7, e39191. [Google Scholar]
- Lee, S.M.; Donaldson, G.P.; Mikulski, Z.; Boyajian, S.; Ley, K.; Mazmanian, S.K. Bacterial colonization factors control specificity and stability of the gut microbiota. Nature 2013, 501, 426–429. [Google Scholar] [CrossRef]
- Makivuokko, H.; Lahtinen, S.J.; Wacklin, P.; Tuovinen, E.; Tenkanen, H.; Nikkila, J.; Bjorklund, M.; Aranko, K.; Ouwehand, A.C.; Matto, J. Association between the abo blood group and the human intestinal microbiota composition. BMC Microbiol. 2012, 12, 94. [Google Scholar] [CrossRef]
- Wacklin, P.; Makivuokko, H.; Alakulppi, N.; Nikkila, J.; Tenkanen, H.; Rabina, J.; Partanen, J.; Aranko, K.; Matto, J. Secretor genotype (FUT2 gene) is strongly associated with the composition of bifidobacteria in the human intestine. PLoS One 2011, 6, e20113. [Google Scholar] [CrossRef]
- Parmar, A.S.; Alakulppi, N.; Paavola-Sakki, P.; Kurppa, K.; Halme, L.; Farkkila, M.; Turunen, U.; Lappalainen, M.; Kontula, K.; Kaukinen, K.; et al. Association study of FUT2 (rs601338) with celiac disease and inflammatory bowel disease in the finnish population. Tissue Antigens 2012, 80, 488–493. [Google Scholar] [CrossRef]
- Sanz, Y.; de Pama, G.; Laparra, M. Unraveling the ties between celiac disease and intestinal microbiota. Int. Rev. Immunol. 2011, 30, 207–218. [Google Scholar] [CrossRef]
- Plot, L.; Amital, H. Infectious associations of celiac disease. Autoimmun. Rev. 2009, 8, 316–319. [Google Scholar] [CrossRef]
- Collado, M.C.; Donat, E.; Ribes-Koninckx, C.; Calabuig, M.; Sanz, Y. Specific duodenal and faecal bacterial groups associated with paediatric coeliac disease. J. Clin. Pathol. 2009, 62, 264–269. [Google Scholar] [CrossRef]
- Di Cagno, R.; de Angelis, M.; de Pasquale, I.; Ndagijimana, M.; Vernocchi, P.; Ricciuti, P.; Gagliardi, F.; Laghi, L.; Crecchio, C.; Guerzoni, M.E.; et al. Duodenal and faecal microbiota of celiac children: Molecular, phenotype and metabolome characterization. BMC Microbiol. 2011, 11, 219. [Google Scholar] [CrossRef]
- Nistal, E.; Caminero, A.; Herran, A.R.; Arias, L.; Vivas, S.; de Morales, J.M.; Calleja, S.; de Miera, L.E.; Arroyo, P.; Casqueiro, J. Differences of small intestinal bacteria populations in adults and children with/without celiac disease: Effect of age, gluten diet, and disease. Inflamm. Bowel Dis. 2012, 18, 649–656. [Google Scholar] [CrossRef]
- Cinova, J.; de Palma, G.; Stepankova, R.; Kofronova, O.; Kverka, M.; Sanz, Y.; Tuckova, L. Role of intestinal bacteria in gliadin-induced changes in intestinal mucosa: Study in germ-free rats. PLoS One 2011, 6, e16169. [Google Scholar]
- D’Arienzo, R.; Stefanile, R.; Maurano, F.; Mazzarella, G.; Ricca, E.; Troncone, R.; Auricchio, S.; Rossi, M. Immunomodulatory effects of Lactobacillus casei administration in a mouse model of gliadin-sensitive enteropathy. Scand. J. Immunol. 2011, 74, 335–341. [Google Scholar] [CrossRef]
- Larsson, J.M.; Karlsson, H.; Sjovall, H.; Hansson, G.C. A complex, but uniform o-glycosylation of the human MUC2 mucin from colonic biopsies analyzed by nanoLC/MSn. Glycobiology 2009, 19, 756–766. [Google Scholar] [CrossRef]
- Larsson, J.M.; Karlsson, H.; Crespo, J.G.; Johansson, M.E.; Eklund, L.; Sjovall, H.; Hansson, G.C. Altered o-glycosylation profile of MUC2 mucin occurs in active ulcerative colitis and is associated with increased inflammation. Inflamm. Bowel Dis. 2011, 17, 2299–2307. [Google Scholar] [CrossRef]
- Brockhausen, I. Sulphotransferases acting on mucin-type oligosaccharides. Biochem. Soc. Transact. 2003, 31, 318–325. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Toft-Hansen, H.; Nielsen, C.; Biagini, M.; Husby, S.; Lillevang, S.T. Lectin Staining Shows no Evidence of Involvement of Glycocalyx/Mucous Layer Carbohydrate Structures in Development of Celiac Disease. Nutrients 2013, 5, 4540-4552. https://doi.org/10.3390/nu5114540
Toft-Hansen H, Nielsen C, Biagini M, Husby S, Lillevang ST. Lectin Staining Shows no Evidence of Involvement of Glycocalyx/Mucous Layer Carbohydrate Structures in Development of Celiac Disease. Nutrients. 2013; 5(11):4540-4552. https://doi.org/10.3390/nu5114540
Chicago/Turabian StyleToft-Hansen, Henrik, Christian Nielsen, Matteo Biagini, Steffen Husby, and Søren T. Lillevang. 2013. "Lectin Staining Shows no Evidence of Involvement of Glycocalyx/Mucous Layer Carbohydrate Structures in Development of Celiac Disease" Nutrients 5, no. 11: 4540-4552. https://doi.org/10.3390/nu5114540