Oligopeptide M13 Phage Display in Pathogen Research
Abstract
:1. M13 Phage Display
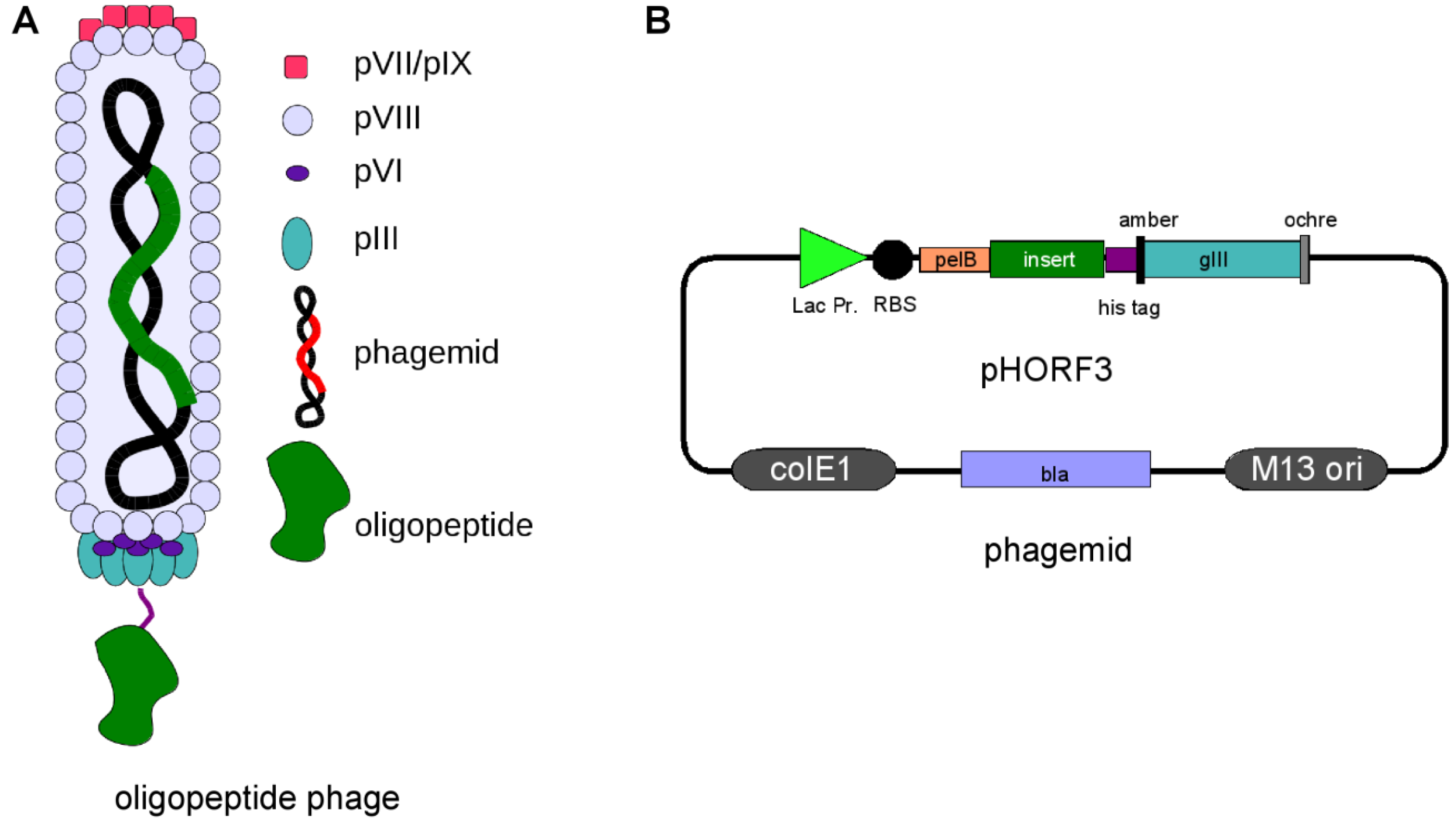
2. Panning
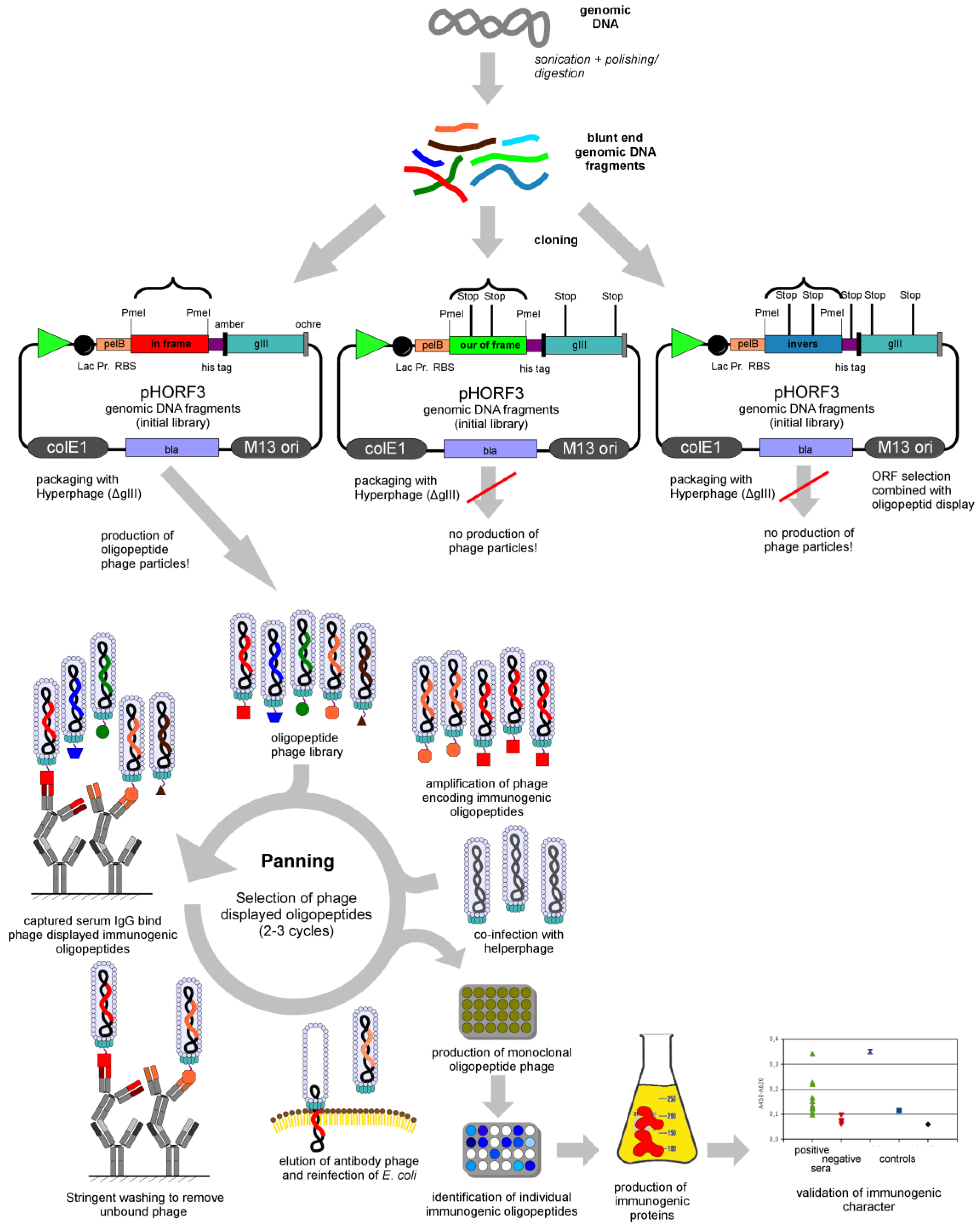
3. Genomic Libraries
4. cDNA Libraries
5. Applications of Oligopeptide Phage Display in Pathogen Research
| Pathogen/Disease | Target Type | Phage System | Library Type | Reference |
|---|---|---|---|---|
| Arabidopsis thaliana (=host) | Pseudomonas cells (=target) | T7 | cDNA | [13] |
| Blue tongue virus (BTV) | two mAbs | M13/ pIII | genomic | [85] |
| Cowdria ruminantium | MAP1 antibodies | M13/ pIII + lambda | genomic | [85] |
| Human cytomegalovirus (HCMV) | serum | lambda | genomic | [19] |
| Human immunodeficiency virus (HIV)/ AIDS | GP140 antibody | M13/ pIII | gp140 gene | [59] |
| Mycobacterium avium (MAP)/ Johne's disease (JD) | serum | lambda | genomic | [86] |
| Mycobacterium tuberculosis/ tuberculosis | serum | M13/ pIII | genomic | [87] |
| Mycoplasma hyopneumoniae/ porcine enzootic pneumonia | serum | M13/ pIII | genomic | [22] |
| Mycoplasma mycoides/ CBPP | serum | M13/ pIII | genomic | [55] |
| Mycoplasma mycoides/ CBPP | serum | M13/ pIII | genomic | [74] |
| Mycoplasma pneumoniae/ pneumonia | serum | lambda | genomic | [20] |
| Plasmodium falciparum/ malaria | erythrocytes | T7 | cDNA | [75] |
| Salmonella Typhimurium/ salmonellosis | serum | M13/ pIII | genomic | [23] |
| Streptococcus pneumoniae/ pneumonia | serum | lambda | genomic | [18] |
| Streptococcus pneumoniae/ pneumonia | serum | lambda | genomic | [14] |
| Taenia solium/ Neurocysticercosis (NCC) | serum | M13/ pVIII | genomic | [76] |
6. Technical Work Flow
7. Summary
Acknowledgments
Conflicts of Interest
References and Notes
- Colwill, K.; Gräslund, S. A roadmap to generate renewable protein binders to the human proteome. Nat. Methods 2011, 8, 551–558. [Google Scholar] [CrossRef]
- Liu, B.A.; Engelmann, B.W.; Nash, P.D. High-throughput analysis of peptide-binding modules. Proteomics 2012, 12, 1527–1546. [Google Scholar] [CrossRef]
- Rentero, I.; Heinis, C. Screening of large molecule diversities by phage display. Chimia (Aarau) 2011, 65, 843–845. [Google Scholar] [CrossRef]
- Schirrmann, T.; Meyer, T.; Schütte, M.; Frenzel, A.; Hust, M. Phage display for the generation of antibodies for proteome research, diagnostics and therapy. Molecules 2011, 16, 412–426. [Google Scholar] [CrossRef]
- Skerra, A. Alternative non-antibody scaffolds for molecular recognition. Curr Opin Biotechnol 2007, 18, 295–304. [Google Scholar] [CrossRef]
- Ullman, C.G.; Frigotto, L.; Cooley, R.N. In vitro methods for peptide display and their applications. Brief Funct. Genomics 2011, 10, 125–134. [Google Scholar] [CrossRef]
- Smith, G.P. Filamentous fusion phage: novel expression vectors that display cloned antigens on the virion surface. Science 1985, 228, 1315–1317. [Google Scholar]
- Hoogenboom, H.R. Selecting and screening recombinant antibody libraries. Nat Biotechnol 2005, 23, 1105–1116. [Google Scholar] [CrossRef]
- Paschke, M. Phage display systems and their applications. Appl. Microbiol. Biotechnol 2006, 70, 2–11. [Google Scholar] [CrossRef]
- Schirrmann, T.; Hust, M. Construction of human antibody gene libraries and selection of antibodies by phage display. Methods Mol. Biol 2010, 651, 177–209. [Google Scholar] [CrossRef]
- Danner, S.; Belasco, J.G. T7 phage display: a novel genetic selection system for cloning RNA-binding proteins from cDNA libraries. Proc. Natl. Acad. Sci. U.S.A. 2001, 98, 12954–12959. [Google Scholar] [CrossRef]
- Houshmand, H.; Fröman, G.; Magnusson, G. Use of bacteriophage T7 displayed peptides for determination of monoclonal antibody specificity and biosensor analysis of the binding reaction. Anal. Biochem. 1999, 268, 363–370. [Google Scholar] [CrossRef]
- Rioja, C.; Van Wees, S.C.; Charlton, K.A.; Pieterse, C.M.J.; Lorenzo, O.; García-Sánchez, S. Wide screening of phage-displayed libraries identifies immune targets in planta. PLoS ONE 2013, 8, e54654. [Google Scholar]
- Beghetto, E.; Gargano, N. Antigen discovery using whole-genome phage display libraries. Methods Mol. Biol. 2013, 1061, 79–95. [Google Scholar] [CrossRef]
- Huse, W.D.; Sastry, L.; Iverson, S.A.; Kang, A.S.; Alting-Mees, M.; Burton, D.R.; Benkovic, S.J.; Lerner, R.A. Generation of a large combinatorial library of the immunoglobulin repertoire in phage lambda. Science 1989, 246, 1275–1281. [Google Scholar]
- Kang, A.S.; Jones, T.M.; Burton, D.R. Antibody redesign by chain shuffling from random combinatorial immunoglobulin libraries. Proc. Natl. Acad. Sci. U.S.A. 1991, 88, 11120–11123. [Google Scholar] [CrossRef]
- Mullinax, R.L.; Gross, E.A.; Amberg, J.R.; Hay, B.N.; Hogrefe, H.H.; Kubitz, M.M.; Greener, A.; Alting-Mees, M.; Ardourel, D.; Short, J.M. Identification of human antibody fragment clones specific for tetanus toxoid in a bacteriophage lambda immunoexpression library. Proc. Natl. Acad. Sci. U.S.A. 1990, 87, 8095–8099. [Google Scholar] [CrossRef]
- Beghetto, E.; Gargano, N.; Ricci, S.; Garufi, G.; Peppoloni, S.; Montagnani, F.; Oggioni, M.; Pozzi, G.; Felici, F. Discovery of novel Streptococcus pneumoniae antigens by screening a whole-genome lambda-display library. FEMS Microbiol Lett 2006, 262, 14–21. [Google Scholar] [CrossRef]
- Beghetto, E.; Paolis, F.D.; Spadoni, A.; Del Porto, P.; Buffolano, W.; Gargano, N. Molecular dissection of the human B cell response against cytomegalovirus infection by lambda display. J. Virol. Methods 2008, 151, 7–14. [Google Scholar] [CrossRef]
- Beghetto, E.; De Paolis, F.; Montagnani, F.; Cellesi, C.; Gargano, N. Discovery of new Mycoplasma pneumoniae antigens by use of a whole-genome lambda display library. Microbes Infect 2009, 11, 66–73. [Google Scholar] [CrossRef]
- Faix, P.H.; Burg, M.A.; Gonzales, M.; Ravey, E.P.; Baird, A.; Larocca, D. Phage display of cDNA libraries: enrichment of cDNA expression using open reading frame selection. Biotechniques 2004, 36, 1018–1022, 1024, 1026–1029. [Google Scholar]
- Kügler, J.; Nieswandt, S.; Gerlach, G.F.; Meens, J.; Schirrmann, T.; Hust, M. Identification of immunogenic polypeptides from a Mycoplasma hyopneumoniae genome library by phage display. Appl. Microbiol. Biotechnol 2008, 80, 447–458. [Google Scholar] [CrossRef]
- Meyer, T.; Schirrmann, T.; Frenzel, A.; Miethe, S.; Stratmann-Selke, J.; Gerlach, G.F.; Strutzberg-Minder, K.; Dübel, S.; Hust, M. Identification of immunogenic proteins and generation of antibodies against Salmonella typhimurium using phage display. BMC Biotechnol. 2012, 12, 29. [Google Scholar]
- Russel, M.; Linderoth, N.A.; Sali, A. Filamentous phage assembly: variation on a protein export theme. Gene 1997, 192, 23–32. [Google Scholar] [CrossRef]
- Thie, H.; Schirrmann, T.; Paschke, M.; Dübel, S.; Hust, M. SRP and Sec pathway leader peptides for antibody phage display and antibody fragment production in E. coli. N Biotechnol 2008, 25, 49–54. [Google Scholar]
- Dunn, I.S. Assembly of functional bacteriophage lambda virions incorporating C-terminal peptide or protein fusions with the major tail protein. J. Mol. Biol. 1995, 248, 497–506. [Google Scholar] [CrossRef]
- Maruyama, I.N.; Maruyama, H.I.; Brenner, S. Lambda foo: a lambda phage vector for the expression of foreign proteins. Proc. Natl. Acad. Sci. U.S.A. 1994, 91, 8273–8277. [Google Scholar] [CrossRef]
- Crameri, R.; Kodzius, R. The powerful combination of phage surface display of cDNA libraries and high throughput screening. Comb. Chem. High Throughput Screen. 2001, 4, 145–155. [Google Scholar] [CrossRef]
- Paschke, M.; Höhne, W. A twin-arginine translocation (Tat)-mediated phage display system. Gene 2005, 350, 79–88. [Google Scholar] [CrossRef]
- Bratkovic, T. Progress in phage display: evolution of the technique and its application. Cell. Mol. Life Sci. 2010, 67, 749–767. [Google Scholar] [CrossRef]
- Sainath Rao, S.; Mohan, K.V.K.; Atreya, C.D. A peptide derived from phage display library exhibits antibacterial activity against E. coli and Pseudomonas aeruginosa. PLoS ONE 2013, 8, e56081. [Google Scholar] [CrossRef]
- Deng, Q.; Zhuang, M.; Kong, Y.-Y.; Xie, Y.-H.; Wang, Y. Screening for PreS specific binding ligands with a phage displayed peptides library. World J. Gastroenterol. 2005, 11, 4018–4023. [Google Scholar]
- Fagerlund, A.; Myrset, A.H.; Kulseth, M.A. Construction and characterization of a 9-mer phage display pVIII-library with regulated peptide density. Appl. Microbiol. Biotechnol. 2008, 80, 925–936. [Google Scholar] [CrossRef]
- Felici, F.; Castagnoli, L.; Musacchio, A.; Jappelli, R.; Cesareni, G. Selection of antibody ligands from a large library of oligopeptides expressed on a multivalent exposition vector. J. Mol. Biol. 1991, 222, 301–310. [Google Scholar] [CrossRef]
- Kang, A.S.; Barbas, C.F.; Janda, K.D.; Benkovic, S.J.; Lerner, R.A. Linkage of recognition and replication functions by assembling combinatorial antibody Fab libraries along phage surfaces. Proc. Natl. Acad. Sci. U.S.A. 1991, 88, 4363–4366. [Google Scholar]
- Løset, G.Å.; Roos, N.; Bogen, B.; Sandlie, I. Expanding the versatility of phage display II: improved affinity selection of folded domains on protein VII and IX of the filamentous phage. LoS ONE 2011, 6, e17433. [Google Scholar] [CrossRef]
- Hufton, S.E.; Moerkerk, P.T.; Meulemans, E.V.; de Bruïne, A.; Arends, J.W.; Hoogenboom, H.R. Phage display of cDNA repertoires: the pVI display system and its applications for the selection of immunogenic ligands. J. Immunol. Methods 1999, 231, 39–51. [Google Scholar] [CrossRef]
- Jespers, L.S.; Messens, J.H.; De Keyser, A.; Eeckhout, D.; Van den Brande, I.; Gansemans, Y.G.; Lauwereys, M.J.; Vlasuk, G.P.; Stanssens, P.E. Surface expression and ligand-based selection of cDNAs fused to filamentous phage gene VI. Biotechnology (N.Y.) 1995, 13, 378–382. [Google Scholar] [CrossRef]
- Somers, V.; Govarts, C.; Hellings, N.; Hupperts, R.; Stinissen, P. Profiling the autoantibody repertoire by serological antigen selection. J. Autoimmun. 2005, 25, 223–228. [Google Scholar] [CrossRef]
- Somers, V.A.; Brandwijk, R.J.; Joosten, B.; Moerkerk, P.T.; Arends, J.-W.; Menheere, P.; Pieterse, W.O.; Claessen, A.; Scheper, R.J.; Hoogenboom, H.R.; Hufton, S.E. A panel of candidate tumor antigens in colorectal cancer revealed by the serological selection of a phage displayed cDNA expression library. J. Immunol. 2002, 169, 2772–2780. [Google Scholar]
- Viaene, A.; Crab, A.; Meiring, M.; Pritchard, D.; Deckmyn, H. Identification of a collagen-binding protein from Necator americanus by using a cDNA-expression phage display library. J. Parasitol. 2001, 87, 619–625. [Google Scholar]
- Gao, C.; Mao, S.; Lo, C.H.; Wirsching, P.; Lerner, R.A.; Janda, K.D. Making artificial antibodies: a format for phage display of combinatorial heterodimeric arrays. Proc. Natl. Acad. Sci. U.S.A 1999, 96, 6025–6030. [Google Scholar]
- Løset, G.Å.; Bogen, B.; Sandlie, I. Expanding the versatility of phage display I: efficient display of peptide-tags on protein VII of the filamentous phage. PLoS ONE 2011, 6, e14702. [Google Scholar]
- Nilssen, N.R.; Frigstad, T.; Pollmann, S.; Roos, N.; Bogen, B.; Sandlie, I.; Løset, G.Å. DeltaPhage—a novel helper phage for high-valence pIX phagemid display. Nucleic Acids Res. 2012, 40, e120. [Google Scholar] [CrossRef]
- Shi, L.; Wheeler, J.C.; Sweet, R.W.; Lu, J.; Luo, J.; Tornetta, M.; Whitaker, B.; Reddy, R.; Brittingham, R.; Borozdina, L.; Chen, Q.; Amegadzie, B.; Knight, D.M.; Almagro, J.C.; Tsui, P. De novo selection of high-affinity antibodies from synthetic fab libraries displayed on phage as pIX fusion proteins. J. Mol. Biol. 2010, 397, 385–396. [Google Scholar] [CrossRef]
- Tornetta, M.; Reddy, R.; Wheeler, J.C. Selection and maturation of antibodies by phage display through fusion to pIX. Methods 2012, 58, 34–39. [Google Scholar] [CrossRef]
- Hust, M.; Dübel, S. Phage display vectors for the in vitro generation of human antibody fragments. Methods Mol Biol 2005, 295, 71–96. [Google Scholar]
- McCafferty, J.; Griffiths, A.D.; Winter, G.; Chiswell, D.J. Phage antibodies: Filamentous phage displaying antibody variable domains. Nature 1990, 348, 552–554. [Google Scholar] [CrossRef]
- Barbas, C.F., 3rd; Kang, A.S.; Lerner, R.A.; Benkovic, S.J. Assembly of combinatorial antibody libraries on phage surfaces: The gene III site. Proc. Natl. Acad. Sci. U.S.A. 1991, 88, 7978–7982. [Google Scholar] [CrossRef]
- Breitling, F.; Dübel, S.; Seehaus, T.; Klewinghaus, I.; Little, M. A surface expression vector for antibody screening. Gene 1991, 104, 147–153. [Google Scholar] [CrossRef]
- Clackson, T.; Hoogenboom, H.R.; Griffiths, A.D.; Winter, G. Making antibody fragments using phage display libraries. Nature 1991, 352, 624–628. [Google Scholar] [CrossRef]
- Hoogenboom, H.R.; Griffiths, A.D.; Johnson, K.S.; Chiswell, D.J.; Hudson, P.; Winter, G. Multi-subunit proteins on the surface of filamentous phage: Methodologies for displaying antibody (Fab) heavy and light chains. Nucleic Acids Res. 1991, 19, 4133–4137. [Google Scholar] [CrossRef]
- Vieira, J.; Messing, J. Production of single-stranded plasmid DNA. Meth. Enzymol. 1987, 153, 3–11. [Google Scholar] [CrossRef]
- Parmley, S.F.; Smith, G.P. Antibody-selectable filamentous fd phage vectors: affinity purification of target genes. Gene 1988, 73, 305–318. [Google Scholar] [CrossRef]
- Naseem, S.; Meens, J.; Jores, J.; Heller, M.; Dübel, S.; Hust, M.; Gerlach, G.-F. Phage display-based identification and potential diagnostic application of novel antigens from Mycoplasma mycoides subsp. mycoides small colony type. Vet. Microbiol 2010, 142, 285–292. [Google Scholar] [CrossRef]
- Rhyner, C.; Weichel, M.; Flückiger, S.; Hemmann, S.; Kleber-Janke, T.; Crameri, R. Cloning allergens via phage display. Methods 2004, 32, 212–218. [Google Scholar] [CrossRef]
- Schaffartzik, A.; Marti, E.; Torsteinsdottir, S.; Mellor, P.S.; Crameri, R.; Rhyner, C. Selective cloning, characterization, and production of the Culicoides nubeculosus salivary gland allergen repertoire associated with equine insect bite hypersensitivity. Vet. Immunol. Immunopathol 2010.
- Hottiger, M.; Gramatikoff, K.; Georgiev, O.; Chaponnier, C.; Schaffner, W.; Hübscher, U. The large subunit of HIV-1 reverse transcriptase interacts with beta-actin. Nucleic Acids Res. 1995, 23, 736–741. [Google Scholar] [CrossRef]
- Zhou, M.; Meyer, T.; Koch, S.; Koch, J.; von Briesen, H.; Benito, J.M.; Soriano, V.; Haberl, A.; Bickel, M.; Dübel, S.; Hust, M.; Dietrich, U. Identification of a new epitope for HIV-neutralizing antibodies in the gp41 membrane proximal external region by an Env-tailored phage display library. Eur. J. Immunol. 2013, 43, 499–509. [Google Scholar] [CrossRef]
- Jacobsson, K.; Rosander, A.; Bjerketorp, J.; Frykberg, L. Shotgun phage display—Selection for bacterial receptins or other exported proteins. Biol Proced Online 2003, 5, 123–135. [Google Scholar] [CrossRef] [Green Version]
- Gupta, S.; Arora, K.; Sampath, A.; Khurana, S.; Singh, S.S.; Gupta, A.; Chaudhary, V.K. Simplified gene-fragment phage display system for epitope mapping. BioTechniques 1999, 27, 328–330, 332–334. [Google Scholar]
- Hust, M.; Meysing, M.; Schirrmann, T.; Selke, M.; Meens, J.; Gerlach, G.-F.; Dübel, S. Enrichment of open reading frames presented on bacteriophage M13 using hyperphage. Biotechniques 2006, 41, 335–342. [Google Scholar] [CrossRef]
- Zacchi, P.; Sblattero, D.; Florian, F.; Marzari, R.; Bradbury, A.R.M. Selecting open reading frames from DNA. Genome Res. 2003, 13, 980–990. [Google Scholar] [CrossRef]
- Rondot, S.; Koch, J.; Breitling, F.; Dübel, S. A helper phage to improve single-chain antibody presentation in phage display. Nat Biotechnol 2001, 19, 75–78. [Google Scholar] [CrossRef]
- Soltes, G.; Hust, M.; Ng, K.K.Y.; Bansal, A.; Field, J.; Stewart, D.I.H.; Dübel, S.; Cha, S.; Wiersma, E.J. On the influence of vector design on antibody phage display. J Biotechnol 2007, 127, 626–637. [Google Scholar] [CrossRef]
- Crameri, R.; Suter, M. Display of biologically active proteins on the surface of filamentous phages: a cDNA cloning system for selection of functional gene products linked to the genetic information responsible for their production. Gene 1993, 137, 69–75. [Google Scholar] [CrossRef]
- Crameri, R.; Jaussi, R.; Menz, G.; Blaser, K. Display of expression products of cDNA libraries on phage surfaces. A versatile screening system for selective isolation of genes by specific gene-product/ligand interaction. Eur. J. Biochem. 1994, 226, 53–58. [Google Scholar] [CrossRef]
- Delvecchio, V.G.; Connolly, J.P.; Alefantis, T.G.; Walz, A.; Quan, M.A.; Patra, G.; Ashton, J.M.; Whittington, J.T.; Chafin, R.D.; Liang, X.; Grewal, P.; Khan, A.S.; Mujer, C.V. Proteomic profiling and identification of immunodominant spore antigens of Bacillus anthracis, Bacillus cereus, and Bacillus thuringiensis. Appl. Environ. Microbiol. 2006, 72, 6355–6363. [Google Scholar] [CrossRef]
- Huntley, J.F.; Conley, P.G.; Hagman, K.E.; Norgard, M.V. Characterization of Francisella tularensis outer membrane proteins. J. Bacteriol. 2007, 189, 561–574. [Google Scholar] [CrossRef]
- LaFrentz, B.R.; LaPatra, S.E.; Call, D.R.; Wiens, G.D.; Cain, K.D. Identification of immunogenic proteins within distinct molecular mass fractions of Flavobacterium psychrophilum. J. Fish Dis. 2011, 34, 823–830. [Google Scholar] [CrossRef]
- Meens, J.; Selke, M.; Gerlach, G.-F. Identification and immunological characterization of conserved Mycoplasma hyopneumoniae lipoproteins Mhp378 and Mhp651. Vet. Microbiol. 2006, 116, 85–95. [Google Scholar] [CrossRef]
- Zhao, Z.; Yan, F.; Ji, W.; Luo, D.; Liu, X.; Xing, L.; Duan, Y.; Yang, P.; Shi, X.; Lu, Z.; Wang, X. Identification of immunoreactive proteins of Brucella melitensis by immunoproteomics. Sci China Life Sci 2011, 54, 880–887. [Google Scholar] [CrossRef]
- Jacobsen, I.D.; Meens, J.; Baltes, N.; Gerlach, G.-F. Differential expression of non-cytoplasmic Actinobacillus pleuropneumoniae proteins induced by addition of bronchoalveolar lavage fluid. Vet. Microbiol. 2005, 109, 245–256. [Google Scholar] [CrossRef]
- Miltiadou, D.R.; Mather, A.; Vilei, E.M.; Du Plessis, D.H. Identification of genes coding for B cell antigens of Mycoplasma mycoides subsp. mycoides Small Colony (MmmSC) by using phage display. BMC Microbiol 2009, 9, 215. [Google Scholar] [CrossRef]
- Lauterbach, S.B.; Lanzillotti, R.; Coetzer, T.L. Construction and use of Plasmodium falciparum phage display libraries to identify host parasite interactions. Malar. J. 2003, 2, 47. [Google Scholar] [CrossRef]
- González, E.; Robles, Y.; Govezensky, T.; Bobes, R.J.; Gevorkian, G.; Manoutcharian, K. Isolation of neurocysticercosis-related antigens from a genomic phage display library of Taenia solium. J Biomol Screen 2010, 15, 1268–1273. [Google Scholar] [CrossRef]
- Crameri, R.; Walter, G. Selective enrichment and high-throughput screening of phage surface-displayed cDNA libraries from complex allergenic systems. Comb. Chem. High Throughput Screen. 1999, 2, 63–72. [Google Scholar]
- Crameri, R.; Kodzius, R.; Konthur, Z.; Lehrach, H.; Blaser, K.; Walter, G. Tapping allergen repertoires by advanced cloning technologies. Int Arch Allergy Immunol 2001, 124, 43–47. [Google Scholar] [CrossRef]
- Kodzius, R.; Rhyner, C.; Konthur, Z.; Buczek, D.; Lehrach, H.; Walter, G.; Crameri, R. Rapid identification of allergen-encoding cDNA clones by phage display and high-density arrays. Comb Chem High Throughput Screen 2003, 6, 147–154. [Google Scholar] [CrossRef]
- Govarts, C.; Somers, K.; Hupperts, R.; Stinissen, P.; Somers, V. Exploring cDNA phage display for autoantibody profiling in the serum of multiple sclerosis patients: optimization of the selection procedure. Ann N Y Acad Sci 2007, 1109, 372–384. [Google Scholar]
- Fosså, A.; Alsøe, L.; Crameri, R.; Funderud, S.; Gaudernack, G.; Smeland, E.B. Serological cloning of cancer/testis antigens expressed in prostate cancer using cDNA phage surface display. Cancer Immunol. Immunother. 2004, 53, 431–438. [Google Scholar] [CrossRef]
- Gazarian, K.G.; Palacios-Rodríguez, Y.; Gazarian, T.G.; Huerta, L. HIV-1 V3 loop crown epitope-focused mimotope selection by patient serum from random phage display libraries: implications for the epitope structural features. Mol. Immunol. 2013, 54, 148–156. [Google Scholar] [CrossRef]
- Larralde, O.G.; Martinez, R.; Camacho, F.; Amin, N.; Aguilar, A.; Talavera, A.; Stott, D.I.; Perez, E.M. Identification of hepatitis A virus mimotopes by phage display, antigenicity and immunogenicity. J. Virol. Methods 2007, 140, 49–58. [Google Scholar]
- Zhang, W.Y.; Wan, Y.; Li, D.G.; Tang, Y.; Zhou, W. A mimotope of pre-S2 region of surface antigen of viral hepatitis B screened by phage display. Cell Res. 2001, 11, 203–208. [Google Scholar] [CrossRef]
- Fehrsen, J.; du Plessis, D.H. Cross-reactive epitope mimics in a fragmented-genome phage display library derived from the rickettsia, Cowdria ruminantium. Immunotechnology 1999, 4, 175–184. [Google Scholar] [CrossRef]
- Nagata, R.; Kawaji, S.; Mori, Y. Use of enoyl coenzyme A hydratase of Mycobacterium avium subsp. paratuberculosis for the serological diagnosis of Johne’s disease. Vet. Immunol. Immunopathol. 2013. [Google Scholar]
- Liu, S.; Han, W.; Sun, C.; Lei, L.; Feng, X.; Yan, S.; Diao, Y.; Gao, Y.; Zhao, H.; Liu, Q.; Yao, C.; Li, M. Subtractive screening with the Mycobacterium tuberculosis surface protein phage display library. Tuberculosis (Edinb) 2011, 91, 579–586. [Google Scholar] [CrossRef]
- Rosander, A.; Guss, B.; Frykberg, L.; Björkman, C.; Näslund, K.; Pringle, M. Identification of immunogenic proteins in Treponema phagedenis-like strain V1 from digital dermatitis lesions by phage display. Vet. Microbiol. 2011, 153, 315–322. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Kügler, J.; Zantow, J.; Meyer, T.; Hust, M. Oligopeptide M13 Phage Display in Pathogen Research. Viruses 2013, 5, 2531-2545. https://doi.org/10.3390/v5102531
Kügler J, Zantow J, Meyer T, Hust M. Oligopeptide M13 Phage Display in Pathogen Research. Viruses. 2013; 5(10):2531-2545. https://doi.org/10.3390/v5102531
Chicago/Turabian StyleKügler, Jonas, Jonas Zantow, Torsten Meyer, and Michael Hust. 2013. "Oligopeptide M13 Phage Display in Pathogen Research" Viruses 5, no. 10: 2531-2545. https://doi.org/10.3390/v5102531





