Updated Values for Molecular Diagnosis for Highly Pathogenic Avian Influenza Virus
Abstract
:1. Influenza Virus
2. Immunochromatography
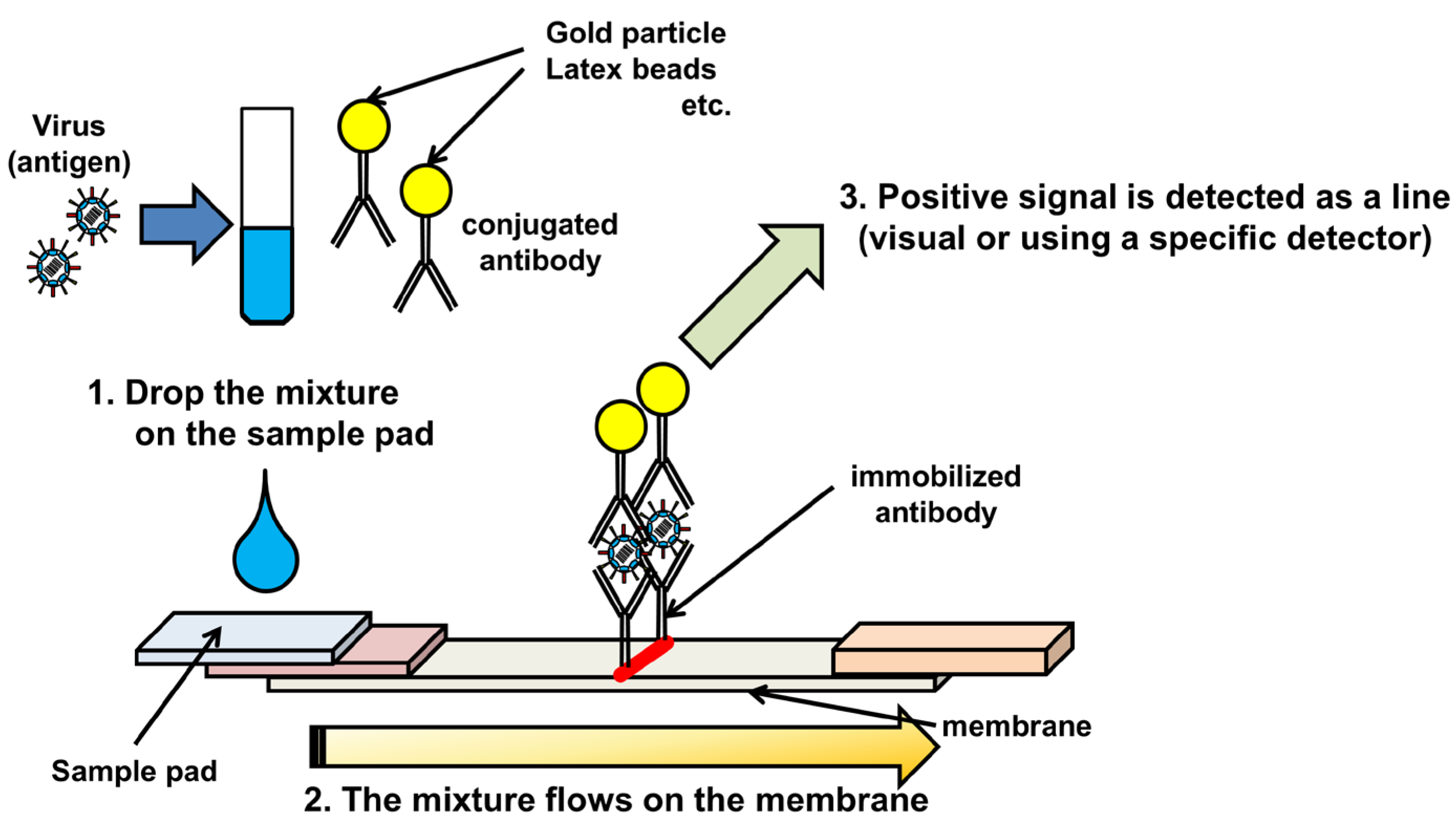
3. Sample preparation for RNA detection systems.
4. PCR-Based Methods
4.1. Reverse Transcriptase-PCR/Degenerate Reverse Transcriptase-PCR
4.2. Quantitative Real-Time PCR
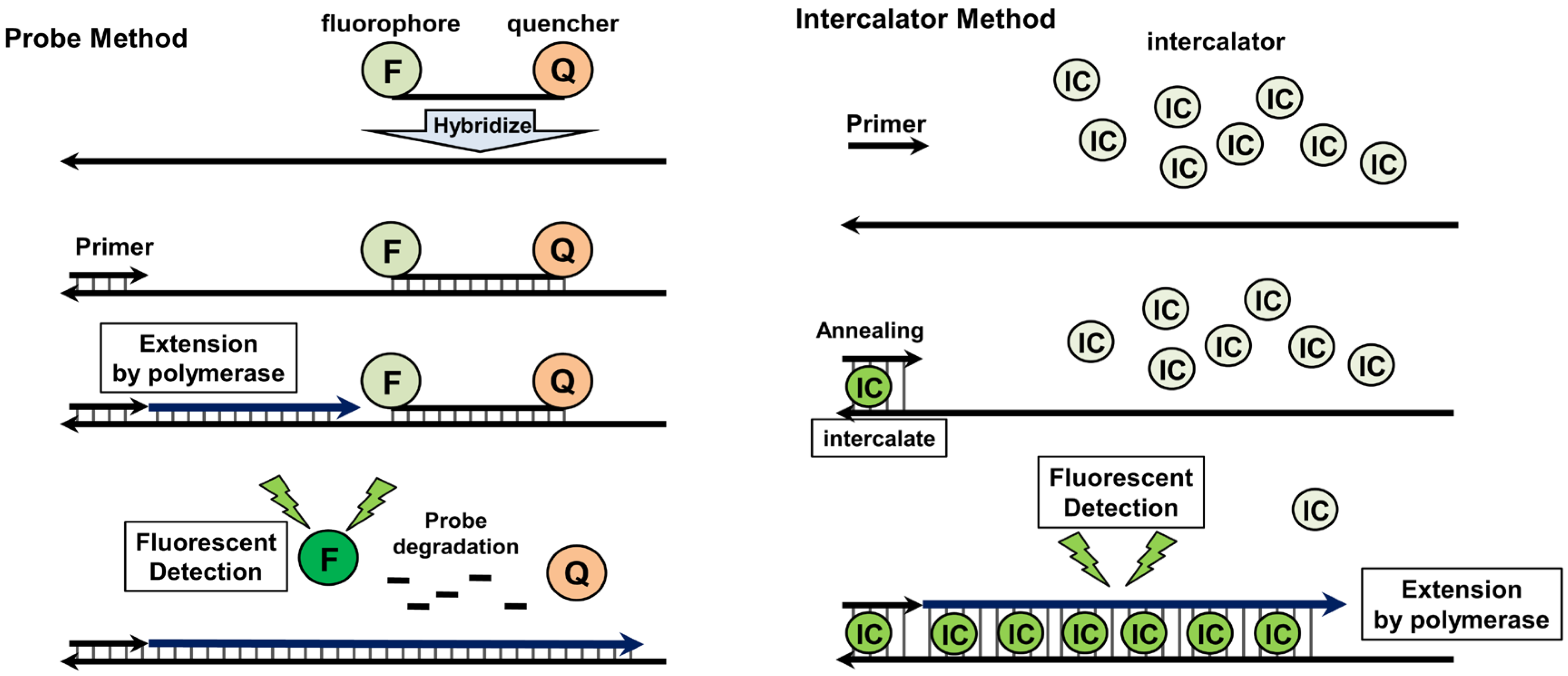
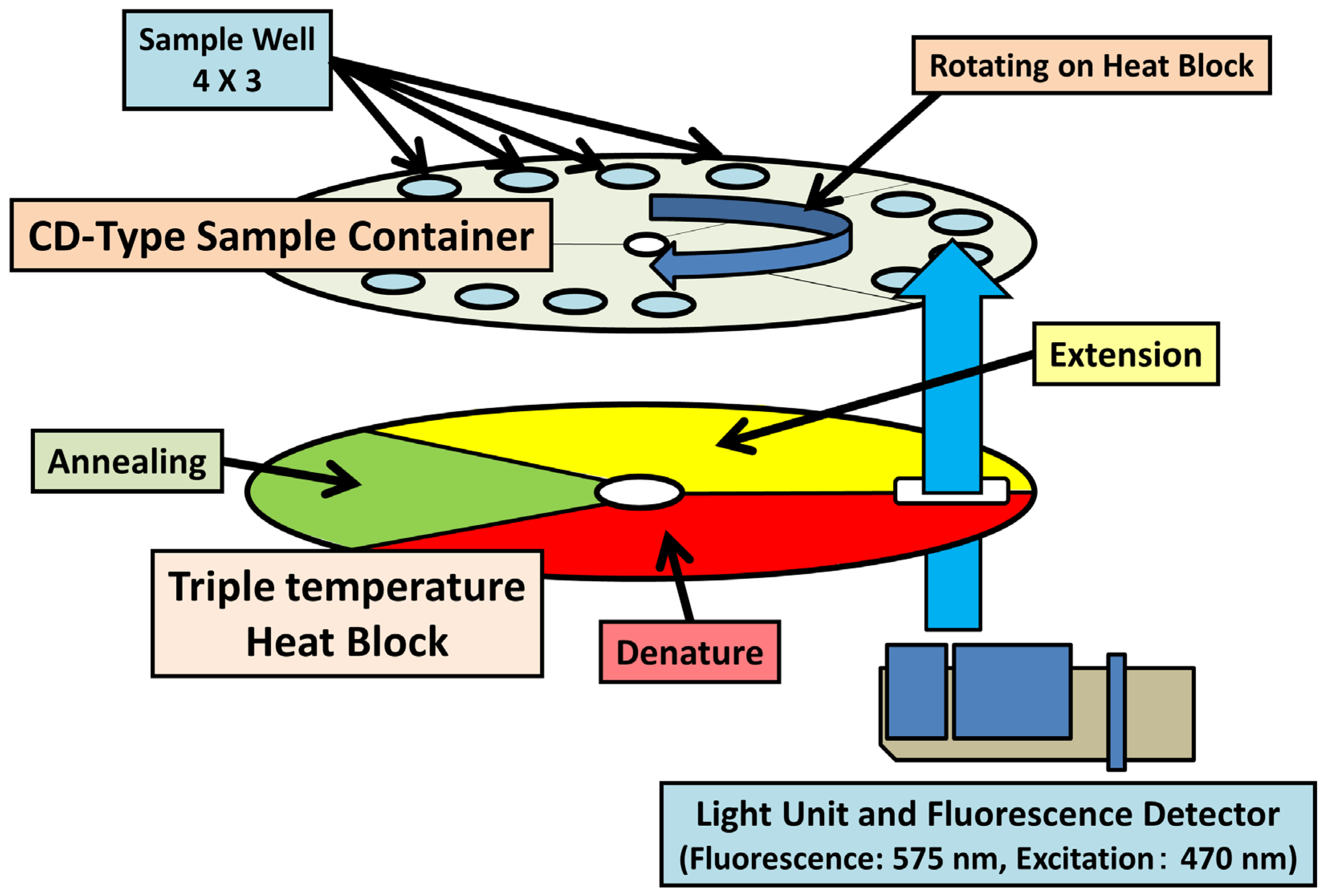
5. DNA Amplification under Isothermal Conditions
5.1. Loop-Mediated Isothermal Amplification
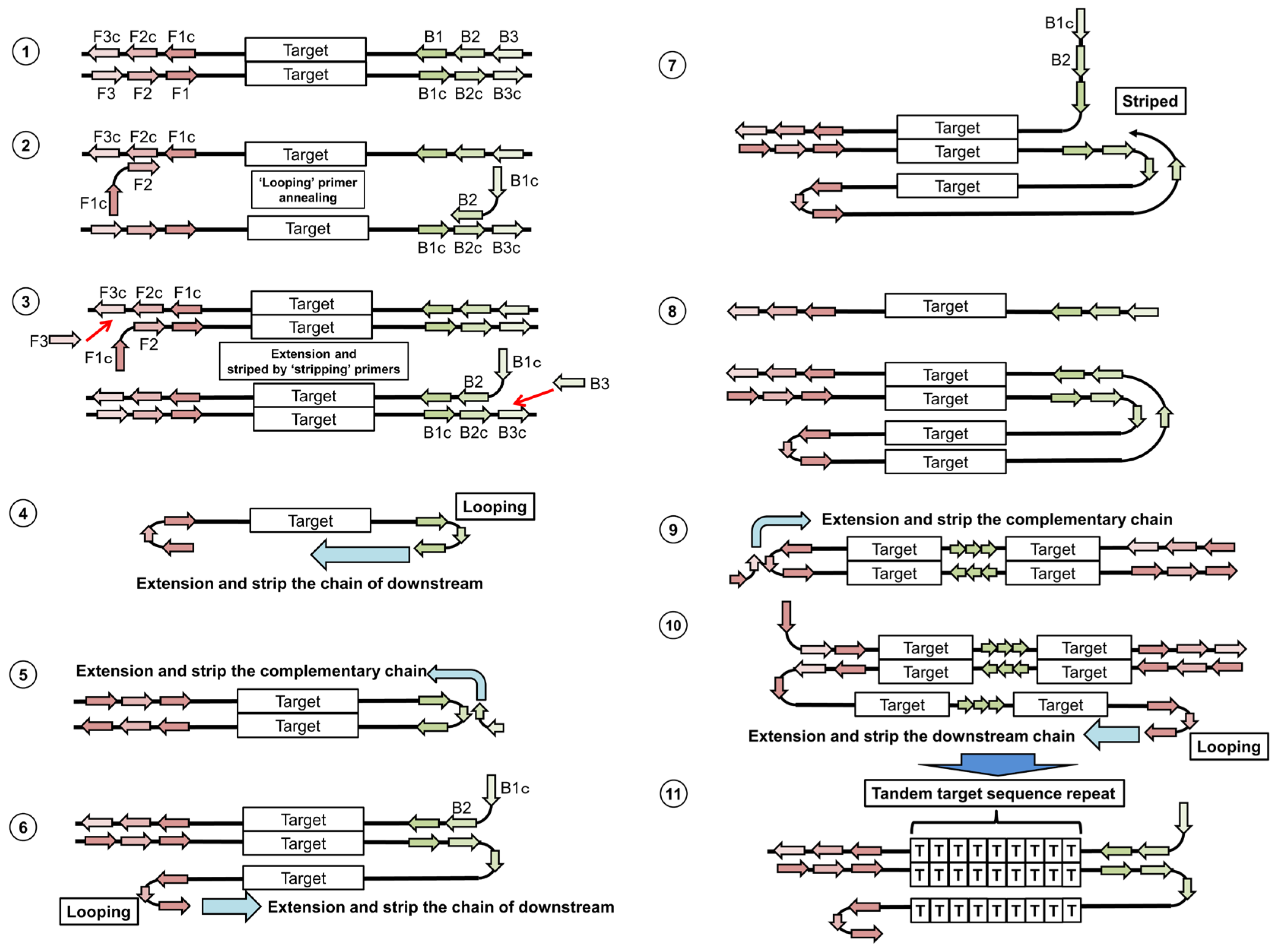
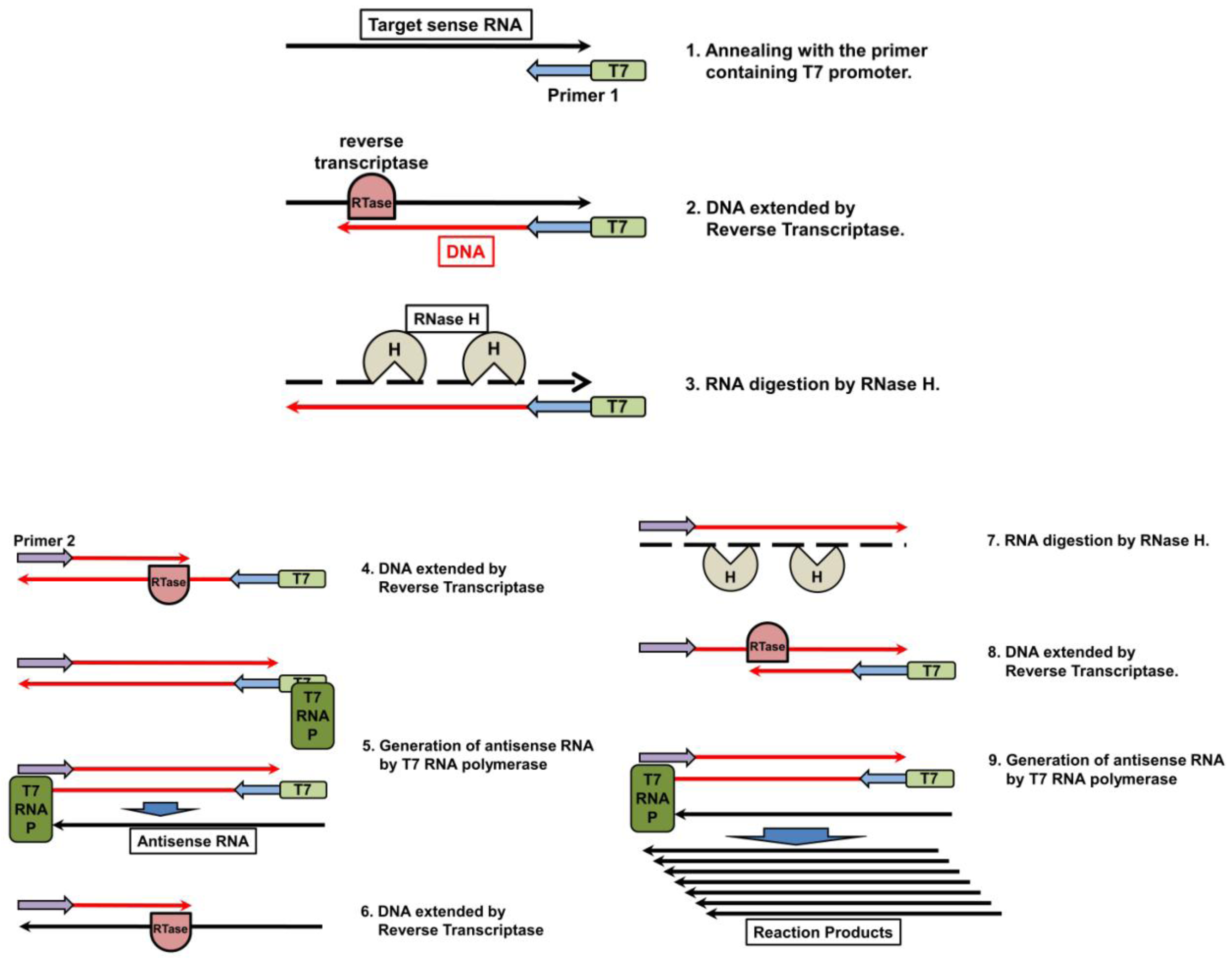
5.2. Nucleic Acid Sequencing-Based Amplification/Real-Time Nucleic Acid Sequencing-Based Amplification
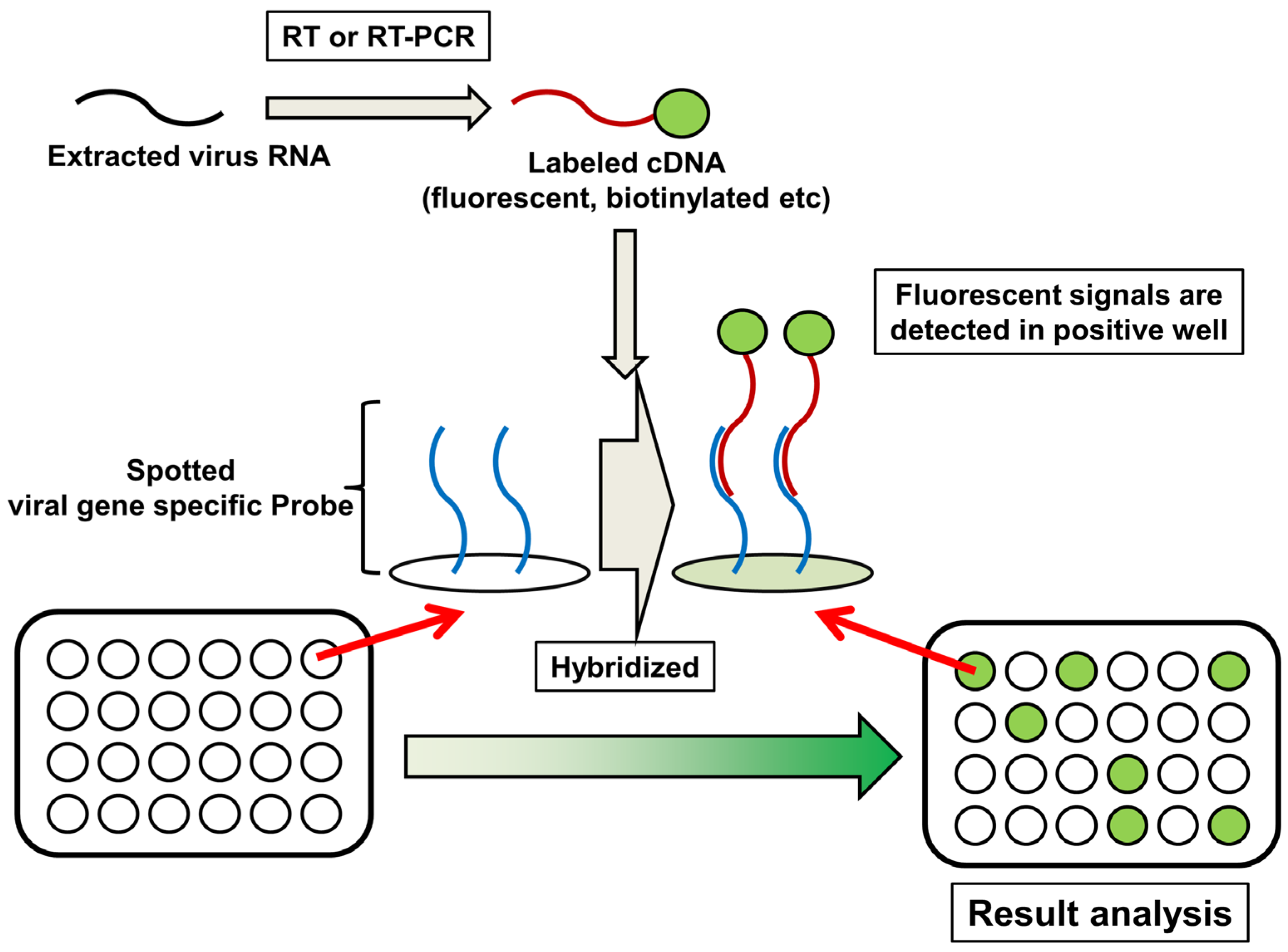
6. DNA Microarray
7. Concluding Remarks
 |
Acknowledgments
Conflict of Interest
References and Notes
- Wright, F.; Neumann, G.; Kawaoka, Y. Orthomyxoviruses. In Fields Virology; Wolters Kluwer, Lippincott Willams & Wilkins: Philadelphia, 2007; pp. 1691–1740. [Google Scholar]
- Kobasa, D.; Takada, A.; Shinya, K.; Hatta, M.; Halfmann, P.; Theriault, S.; Suzuki, H.; Nishimura, H.; Mitamura, K.; Sugaya, N.; Usui, T.; Murata, T.; Maeda, Y.; Watanabe, S.; Suresh, M.; Suzuki, T.; Suzuki, Y.; Feldmann, H.; Kawaoka, Y. Enhanced virulence of influenza A viruses with the haemagglutinin of the 1918 pandemic virus. Nature 2004, 431, 703–707. [Google Scholar] [CrossRef]
- Kawana, A.; Naka, G.; Fujikura, Y.; Kato, Y.; Mizuno, Y.; Kondo, T.; Kudo, K. Spanish influenza in Japanese armed forces, 1918-1920. Emerg. Infect. Dis. 2007, 13, 590–593. [Google Scholar] [CrossRef]
- Munster, V.J.; Baas, C.; Lexmond, P.; Waldenstrom, J.; Wallensten, A.; Fransson, T.; Rimmelzwaan, G.F.; Beyer, W.E.; Schutten, M.; Olsen, B.; Osterhaus, A.D.; Fouchier, R.A. Spatial, temporal, and species variation in prevalence of influenza A viruses in wild migratory birds. PLoS Pathog. 2007, 3, e61. [Google Scholar] [CrossRef]
- Webster, R.G. Influenza virus: transmission between species and relevance to emergence of the next human pandemic. Arch. Virol. Suppl. 1997, 13, 105–113. [Google Scholar]
- Swayne, D.E.; Suarez, D.L. Highly pathogenic avian influenza. Rev. Sci. Tech. 2000, 19, 463–482. [Google Scholar]
- Neumann, G.; Chen, H.; Gao, G.F.; Shu, Y.; Kawaoka, Y. H5N1 influenza viruses: outbreaks and biological properties. Cell Res. 2010, 20, 51–61. [Google Scholar] [CrossRef]
- Horimoto, T.; Ito, T.; Alexander, D.J.; Kawaoka, Y. Cleavability of hemagglutinin from an extremely virulent strain of avian influenza virus containing a unique cleavage site sequence. J. Vet. Med. Sci. 1995, 57, 927–930. [Google Scholar] [CrossRef]
- Steinhauer, D.A. Role of hemagglutinin cleavage for the pathogenicity of influenza virus. Virology 1999, 258, 1–20. [Google Scholar] [CrossRef]
- White, J.; Matlin, K.; Helenius, A. Cell fusion by Semliki Forest, influenza, and vesicular stomatitis viruses. J. Cell. Biol. 1981, 89, 674–679. [Google Scholar] [CrossRef]
- Subbarao, K.; Klimov, A.; Katz, J.; Regnery, H.; Lim, W.; Hall, H.; Perdue, M.; Swayne, D.; Bender, C.; Huang, J.; Hemphill, M.; Rowe, T.; Shaw, M.; Xu, X.; Fukuda, K.; Cox, N. Characterization of an avian influenza A (H5N1) virus isolated from a child with a fatal respiratory illness. Science 1998, 279, 393–396. [Google Scholar]
- Claas, E.C.; Osterhaus, A.D.; van Beek, R.; De Jong, J.C.; Rimmelzwaan, G.F.; Senne, D.A.; Krauss, S.; Shortridge, K.F.; Webster, R.G. Human influenza A H5N1 virus related to a highly pathogenic avian influenza virus. Lancet 1998, 351, 472–477. [Google Scholar]
- de Jong, J.C.; Claas, E.C.; Osterhaus, A.D.; Webster, R.G.; Lim, W.L. A pandemic warning? Nature 1997, 389, 554. [Google Scholar]
- Hoffmann, E.; Stech, J.; Leneva, I.; Krauss, S.; Scholtissek, C.; Chin, P.S.; Peiris, M.; Shortridge, K.F.; Webster, R.G. Characterization of the influenza A virus gene pool in avian species in southern China: was H6N1 a derivative or a precursor of H5N1? J. Virol. 2000, 74, 6309–6315. [Google Scholar]
- Cheung, C.L.; Vijaykrishna, D.; Smith, G.J.; Fan, X.H.; Zhang, J.X.; Bahl, J.; Duan, L.; Huang, K.; Tai, H.; Wang, J.; Poon, L.L.; Peiris, J.S.; Chen, H.; Guan, Y. Establishment of influenza A virus (H6N1) in minor poultry species in southern China. J. Virol. 2007, 81, 10402–10412. [Google Scholar]
- Chen, H.; Deng, G.; Li, Z.; Tian, G.; Li, Y.; Jiao, P.; Zhang, L.; Liu, Z.; Webster, R.G.; Yu, K. The evolution of H5N1 influenza viruses in ducks in southern China. Proc. Natl. Acad. Sci. U S A 2004, 101, 10452–10457. [Google Scholar]
- Guan, Y.; Peiris, J.S.; Lipatov, A.S.; Ellis, T.M.; Dyrting, K.C.; Krauss, S.; Zhang, L.J.; Webster, R.G.; Shortridge, K.F. Emergence of multiple genotypes of H5N1 avian influenza viruses in Hong Kong SAR. Proc. Natl. Acad. Sci. U S A 2002, 99, 8950–8955. [Google Scholar]
- Guan, Y.; Poon, L.L.; Cheung, C.Y.; Ellis, T.M.; Lim, W.; Lipatov, A.S.; Chan, K.H.; Sturm-Ramirez, K.M.; Cheung, C.L.; Leung, Y.H.; Yuen, K.Y.; Webster, R.G.; Peiris, J.S. H5N1 influenza: a protean pandemic threat. Proc. Natl. Acad. Sci. U S A 2004, 101, 8156–8161. [Google Scholar]
- Li, K.S.; Guan, Y.; Wang, J.; Smith, G.J.; Xu, K.M.; Duan, L.; Rahardjo, A.P.; Puthavathana, P.; Buranathai, C.; Nguyen, T.D.; Estoepangestie, A.T.; Chaisingh, A.; Auewarakul, P.; Long, H.T.; Hanh, N.T.; Webby, R.J.; Poon, L.L.; Chen, H.; Shortridge, K.F.; Yuen, K.Y.; Webster, R.G.; Peiris, J.S. Genesis of a highly pathogenic and potentially pandemic H5N1 influenza virus in eastern Asia. Nature 2004, 430, 209–213. [Google Scholar]
- Webster, R.G.; Govorkova, E.A. H5N1 influenza–continuing evolution and spread. N. Engl. J. Med. 2006, 355, 2174–2177. [Google Scholar] [CrossRef]
- Gambotto, A.; Barratt-Boyes, S.M.; de Jong, M.D.; Neumann, G.; Kawaoka, Y. Human infection with highly pathogenic H5N1 influenza virus. Lancet 2008, 371, 1464–1475. [Google Scholar] [CrossRef]
- Maines, T.R.; Lu, X.H.; Erb, S.M.; Edwards, L.; Guarner, J.; Greer, P.W.; Nguyen, D.C.; Szretter, K.J.; Chen, L.M.; Thawatsupha, P.; Chittaganpitch, M.; Waicharoen, S.; Nguyen, D.T.; Nguyen, T.; Nguyen, H.H.; Kim, J.H.; Hoang, L.T.; Kang, C.; Phuong, L.S.; Lim, W.; Zaki, S.; Donis, R.O.; Cox, N.J.; Katz, J.M.; Tumpey, T.M. Avian influenza (H5N1) viruses isolated from humans in Asia in 2004 exhibit increased virulence in mammals. J. Virol. 2005, 79, 11788–11800. [Google Scholar]
- Cumulative number of confirmed human cases of avian influenza A(H5N1) reported to WHO, 2003-2012. Available online: http://www.who.int/influenza/human_animal_interface/EN_GIP_20120305CumulativeNumberH5N1cases.pdf (accessed on 7 March 2012).
- Belser, J.A.; Bridges, C.B.; Katz, J.M.; Tumpey, T.M. Past, present, and possible future human infection with influenza virus A subtype H7. Emerg. Infect. Dis. 2009, 15, 859–865. [Google Scholar] [CrossRef]
- Kurtz, J.; Manvell, R.J.; Banks, J. Avian influenza virus isolated from a woman with conjunctivitis. Lancet 1996, 348, 901–902. [Google Scholar] [CrossRef]
- Koopmans, M.; Wilbrink, B.; Conyn, M.; Natrop, G.; van der Nat, H.; Vennema, H.; Meijer, A.; van Steenbergen, J.; Fouchier, R.; Osterhaus, A.; Bosman, A. Transmission of H7N7 avian influenza A virus to human beings during a large outbreak in commercial poultry farms in the Netherlands. Lancet 2004, 363, 587–593. [Google Scholar]
- Fouchier, R.A.; Schneeberger, P.M.; Rozendaal, F.W.; Broekman, J.M.; Kemink, S.A.; Munster, V.; Kuiken, T.; Rimmelzwaan, G.F.; Schutten, M.; Van Doornum, G.J.; Koch, G.; Bosman, A.; Koopmans, M.; Osterhaus, A.D. Avian influenza A virus (H7N7) associated with human conjunctivitis and a fatal case of acute respiratory distress syndrome. Proc. Natl. Acad. Sci. U S A 2004, 101, 1356–1361. [Google Scholar]
- Tweed, S.A.; Skowronski, D.M.; David, S.T.; Larder, A.; Petric, M.; Lees, W.; Li, Y.; Katz, J.; Krajden, M.; Tellier, R.; Halpert, C.; Hirst, M.; Astell, C.; Lawrence, D.; Mak, A. Human illness from avian influenza H7N3, British Columbia. Emerg. Infect. Dis. 2004, 10, 2196–2199. [Google Scholar]
- Avian influenza in the Netherlands. Available online: http://www.who.int/csr/don/2003_04_24/en/ (accessed on 22 May 2012).
- Capua, I.; Alexander, D.J. Avian influenza: recent developments. Avian. Pathol. 2004, 33, 393–404. [Google Scholar] [CrossRef]
- Dawood, F.S.; Jain, S.; Finelli, L.; Shaw, M.W.; Lindstrom, S.; Garten, R.J.; Gubareva, L.V.; Xu, X.; Bridges, C.B.; Uyeki, T.M. Emergence of a novel swine-origin influenza A (H1N1) virus in humans. N. Engl. J. Med. 2009, 360, 2605–2615. [Google Scholar]
- Manual of Diagnostic Tests and Vaccines for Terrestrial Animals: CHAPTER 2.3.4 AVIAN INFLUENZA. Available online: http://www.oie.int/fileadmin/Home/eng/Health_standards/tahm/2.03.04_AI.pdf (accessed on 23 May 2012).
- Manual of Diagnostic Tests and Vaccines for Terrestrial Animals: CHAPTER 2.8.8 SWINE INFLUENZA. Available online: http://www.oie.int/fileadmin/Home/eng/Health_standards/tahm/2.08.08_SWINE_INFLUENZA.pdf (accessed on 23 May 2012).
- A revision of the system of nomenclature for influenza viruses: a WHO memorandum. Bull. World Health Organ. 1980, 58, 585–591.
- Schild, G.C.; Newman, R.W.; Webster, R.G.; Major, D.; Hinshaw, V.S. Antigenic analysis of influenza A virus surface antigens: considerations for the nomenclature of influenza virus. Comp. Immunol. Microbiol. Infect. Dis. 1980, 3, 5–18. [Google Scholar] [CrossRef]
- Takimoto, S.; Grandien, M.; Ishida, M.A.; Pereira, M.S.; Paiva, T.M.; Ishimaru, T.; Makita, E.M.; Martinez, C.H. Comparison of enzyme-linked immunosorbent assay, indirect immunofluorescence assay, and virus isolation for detection of respiratory viruses in nasopharyngeal secretions. Journal of clinical microbiology 1991, 29, 470–474. [Google Scholar]
- Wang, R.; Taubenberger, J.K. Methods for molecular surveillance of influenza. Expert Rev. Anti Infect. Ther. 2010, 8, 517–527. [Google Scholar] [CrossRef]
- Sakai-Tagawa, Y.; Ozawa, M.; Tamura, D.; Le, M.Q.; Nidom, C.A.; Sugaya, N.; Kawaoka, Y. Sensitivity of influenza rapid diagnostic tests to H5N1 and 2009 pandemic H1N1 viruses. Journal of clinical microbiology 2010, 48, 2872–2877. [Google Scholar] [CrossRef]
- Fader, R.C. Comparison of the Binax NOW Flu A enzyme immunochromatographic assay and R-Mix shell vial culture for the 2003-2004 influenza season. Journal of clinical microbiology 2005, 43, 6133–6135. [Google Scholar] [CrossRef]
- Cruz, A.T.; Cazacu, A.C.; McBride, L.J.; Greer, J.M.; Demmler, G.J. Performance characteristics of a rapid immunochromatographic assay for detection of influenza virus in children during the 2003 to 2004 influenza season. Ann. Emerg. Med. 2006, 47, 250–254. [Google Scholar] [CrossRef]
- Keitel, K.; Wagner, N.; Lacroix, L.; Manzano, S.; Gervaix, A. Performance characteristics of a rapid immunochromatographic assay for detection of pandemic influenza A (H1N1) virus in children. Eur. J. Pediatr. 2011, 170, 511–517. [Google Scholar] [CrossRef]
- Tsuda, Y.; Sakoda, Y.; Sakabe, S.; Mochizuki, T.; Namba, Y.; Kida, H. Development of an immunochromatographic kit for rapid diagnosis of H5 avian influenza virus infection. Microbiol. Immunol. 2007, 51, 903–907. [Google Scholar]
- Miyagawa, E.; Kogaki, H.; Uchida, Y.; Fujii, N.; Shirakawa, T.; Sakoda, Y.; Kida, H. Development of a novel rapid immunochromatographic test specific for the H5 influenza virus. J. Virol. Methods 2011, 173, 213–219. [Google Scholar] [CrossRef]
- Wada, A.; Sakoda, Y.; Oyamada, T.; Kida, H. Development of a highly sensitive immunochromatographic detection kit for H5 influenza virus hemagglutinin using silver amplification. J. Virol. Methods 2011, 178, 82–86. [Google Scholar] [CrossRef]
- Zou, Z.; Du, D.; Wang, J.; Smith, J.N.; Timchalk, C.; Li, Y.; Lin, Y. Quantum dot-based immunochromatographic fluorescent biosensor for biomonitoring trichloropyridinol, a biomarker of exposure to chlorpyrifos. Anal. Chem. 2010, 82, 5125–5133. [Google Scholar] [CrossRef]
- Khreich, N.; Lamourette, P.; Lagoutte, B.; Ronco, C.; Franck, X.; Creminon, C.; Volland, H. A fluorescent immunochromatographic test using immunoliposomes for detecting microcystins and nodularins. Anal. Bioanal. Chem. 2010, 397, 1733–1742. [Google Scholar] [CrossRef]
- Dhumpa, R.; Bu, M.; Handberg, K.J.; Wolff, A.; Bang, D.D. Rapid sample preparation for detection and identification of avian influenza virus from chicken faecal samples using magnetic bead microsystem. J. Virol. Methods 2010, 169, 228–231. [Google Scholar] [CrossRef]
- Boom, R.; Sol, C.J.; Salimans, M.M.; Jansen, C.L.; Wertheim-van Dillen, P.M.; van der Noordaa, J. Rapid and simple method for purification of nucleic acids. Journal of clinical microbiology 1990, 28, 495–503. [Google Scholar]
- Perandin, F.; Pollara, P.C.; Gargiulo, F.; Bonfanti, C.; Manca, N. Performance evaluation of the automated NucliSens easyMAG nucleic acid extraction platform in comparison with QIAamp Mini kit from clinical specimens. Diagn. Microbiol. Infect. Dis. 2009, 64, 158–165. [Google Scholar] [CrossRef]
- Loens, K.; Bergs, K.; Ursi, D.; Goossens, H.; Ieven, M. Evaluation of NucliSens easyMAG for automated nucleic acid extraction from various clinical specimens. Journal of clinical microbiology 2007, 45, 421–425. [Google Scholar] [CrossRef]
- Yang, G.; Erdman, D.E.; Kodani, M.; Kools, J.; Bowen, M.D.; Fields, B.S. Comparison of commercial systems for extraction of nucleic acids from DNA/RNA respiratory pathogens. J. Virol. Methods 2011, 171, 195–199. [Google Scholar] [CrossRef]
- Chandler, D.P.; Griesemer, S.B.; Cooney, C.G.; Holmberg, R.; Thakore, N.; Mokhiber, B.; Belgrader, P.; Knickerbocker, C.; Schied, J.; St George, K. Rapid, simple influenza RNA extraction from nasopharyngeal samples. J. Virol. Methods 2012, 183, 8–13. [Google Scholar] [CrossRef]
- Herrmann, B.; Larsson, C.; Zweygberg, B.W. Simultaneous detection and typing of influenza viruses A and B by a nested reverse transcription-PCR: comparison to virus isolation and antigen detection by immunofluorescence and optical immunoassay (FLU OIA). Journal of clinical microbiology 2001, 39, 134–138. [Google Scholar] [CrossRef]
- Hoffmann, E.; Stech, J.; Guan, Y.; Webster, R.G.; Perez, D.R. Universal primer set for the full-length amplification of all influenza A viruses. Arch. Virol. 2001, 146, 2275–2289. [Google Scholar]
- Zhou, B.; Donnelly, M.E.; Scholes, D.T.; St George, K.; Hatta, M.; Kawaoka, Y.; Wentworth, D.E. Single-reaction genomic amplification accelerates sequencing and vaccine production for classical and Swine origin human influenza a viruses. J. Virol. 2009, 83, 10309–10313. [Google Scholar] [CrossRef]
- Gall, A.; Hoffmann, B.; Harder, T.; Grund, C.; Beer, M. Universal primer set for amplification and sequencing of HA0 cleavage sites of all influenza A viruses. Journal of clinical microbiology 2008, 46, 2561–2567. [Google Scholar]
- Michael, K.; Harder, T.C.; Mettenleiter, T.C.; Karger, A. Diagnosis and strain differentiation of avian influenza viruses by restriction fragment mass analysis. J. Virol. Methods 2009, 158, 63–69. [Google Scholar] [CrossRef]
- Alvarez, A.C.; Brunck, M.E.; Boyd, V.; Lai, R.; Virtue, E.; Chen, W.; Bletchly, C.; Heine, H.G.; Barnard, R. A broad spectrum, one-step reverse-transcription PCR amplification of the neuraminidase gene from multiple subtypes of influenza A virus. Virol. J. 2008, 5, 77. [Google Scholar] [CrossRef]
- Qiu, B.F.; Liu, W.J.; Peng, D.X.; Hu, S.L.; Tang, Y.H.; Liu, X.F. A reverse transcription-PCR for subtyping of the neuraminidase of avian influenza viruses. J. Virol. Methods 2009, 155, 193–198. [Google Scholar] [CrossRef]
- van Elden, L.J.; Nijhuis, M.; Schipper, P.; Schuurman, R.; van Loon, A.M. Simultaneous detection of influenza viruses A and B using real-time quantitative PCR. Journal of clinical microbiology 2001, 39, 196–200. [Google Scholar] [CrossRef]
- Alexander, D.J. Avian influenza - diagnosis. Zoonoses Public Health 2008, 55, 16–23. [Google Scholar] [CrossRef]
- Spackman, E.; Senne, D.A.; Myers, T.J.; Bulaga, L.L.; Garber, L.P.; Perdue, M.L.; Lohman, K.; Daum, L.T.; Suarez, D.L. Development of a real-time reverse transcriptase PCR assay for type A influenza virus and the avian H5 and H7 hemagglutinin subtypes. Journal of clinical microbiology 2002, 40, 3256–3260. [Google Scholar]
- Habib-Bein, N.F.; Beckwith, W.H., 3rd; Mayo, D.; Landry, M.L. Comparison of SmartCycler real-time reverse transcription-PCR assay in a public health laboratory with direct immunofluorescence and cell culture assays in a medical center for detection of influenza A virus. Journal of clinical microbiology 2003, 41, 3597–3601. [Google Scholar] [CrossRef]
- Giglio, S.; Monis, P.T.; Saint, C.P. Demonstration of preferential binding of SYBR Green I to specific DNA fragments in real-time multiplex PCR. Nucleic. Acids. Res. 2003, 31, e136. [Google Scholar] [CrossRef]
- Krafft, A.E.; Russell, K.L.; Hawksworth, A.W.; McCall, S.; Irvine, M.; Daum, L.T.; Connoly, J.L.; Reid, A.H.; Gaydos, J.C.; Taubenberger, J.K. Evaluation of PCR testing of ethanol-fixed nasal swab specimens as an augmented surveillance strategy for influenza virus and adenovirus identification. Journal of clinical microbiology 2005, 43, 1768–1775. [Google Scholar]
- Aguero, M.; Sanchez, A.; San Miguel, E.; Gomez-Tejedor, C.; Jimenez-Clavero, M.A. A real-time TaqMan RT-PCR method for neuraminidase type 1 (N1) gene detection of H5N1 Eurasian strains of avian influenza virus. Avian. Dis. 2007, 51, 378–381. [Google Scholar] [CrossRef]
- Ellis, J.S.; Smith, J.W.; Braham, S.; Lock, M.; Barlow, K.; Zambon, M.C. Design and validation of an H5 TaqMan real-time one-step reverse transcription-PCR and confirmatory assays for diagnosis and verification of influenza A virus H5 infections in humans. Journal of clinical microbiology 2007, 45, 1535–1543. [Google Scholar]
- Di Trani, L.; Bedini, B.; Donatelli, I.; Campitelli, L.; Chiappini, B.; De Marco, M.A.; Delogu, M.; Buonavoglia, C.; Vaccari, G. A sensitive one-step real-time PCR for detection of avian influenza viruses using a MGB probe and an internal positive control. BMC Infect. Dis. 2006, 6, 87. [Google Scholar] [CrossRef]
- Wahlestedt, C.; Salmi, P.; Good, L.; Kela, J.; Johnsson, T.; Hokfelt, T.; Broberger, C.; Porreca, F.; Lai, J.; Ren, K.; Ossipov, M.; Koshkin, A.; Jakobsen, N.; Skouv, J.; Oerum, H.; Jacobsen, M.H.; Wengel, J. Potent and nontoxic antisense oligonucleotides containing locked nucleic acids. Proc. Natl. Acad. Sci. U S A 2000, 97, 5633–5638. [Google Scholar]
- Braasch, D.A.; Corey, D.R. Locked nucleic acid (LNA): fine-tuning the recognition of DNA and RNA. Chem. Biol. 2001, 8, 1–7. [Google Scholar] [CrossRef]
- Thanh, T.T.; Pawestri, H.A.; Ngoc, N.M.; Hien, V.M.; Syahrial, H.; Trung, N.V.; van Doorn, R.H.; Wertheim, H.F.; Chau, N.V.; Ha do, Q.; Farrar, J.J.; Hien, T.T.; Sedyaningsih, E.R.; de Jong, M.D. A real-time RT-PCR for detection of clade 1 and 2 H5N1 influenza A virus using locked nucleic acid (LNA) TaqMan probes. Virol. J. 2010, 7, 46. [Google Scholar] [CrossRef]
- Das, A.; Spackman, E.; Thomas, C.; Swayne, D.E.; Suarez, D.L. Detection of H5N1 high-pathogenicity avian influenza virus in meat and tracheal samples from experimentally infected chickens. Avian. Dis. 2008, 52, 40–48. [Google Scholar] [CrossRef]
- Tsukamoto, K.; Javier, P.C.; Shishido, M.; Noguchi, D.; Pearce, J.; Kang, H.M.; Jeong, O.M.; Lee, Y.J.; Nakanishi, K.; Ashizawa, T. SYBR green-based real-time reverse transcription-PCR for typing and subtyping of all hemagglutinin and neuraminidase genes of avian influenza viruses and comparison to standard serological subtyping tests. Journal of clinical microbiology 2012, 50, 37–45. [Google Scholar] [CrossRef]
- Hoffmann, B.; Harder, T.; Starick, E.; Depner, K.; Werner, O.; Beer, M. Rapid and highly sensitive pathotyping of avian influenza A H5N1 virus by using real-time reverse transcription-PCR. Journal of clinical microbiology 2007, 45, 600–603. [Google Scholar] [CrossRef]
- Sakurai, A.; Nomura, N.; Nanba, R.; Sinkai, T.; Iwaki, T.; Obayashi, T.; Hashimoto, K.; Hasegawa, M.; Sakoda, Y.; Naito, A.; Morizane, Y.; Hosaka, M.; Tsuboi, K.; Kida, H.; Kai, A.; Shibasaki, F. Rapid typing of influenza viruses using super high-speed quantitative real-time PCR. J. Virol. Methods 2011, 178, 75–81. [Google Scholar] [CrossRef]
- Chaharaein, B.; Omar, A.R.; Aini, I.; Yusoff, K.; Hassan, S.S. Detection of H5, H7 and H9 subtypes of avian influenza viruses by multiplex reverse transcription-polymerase chain reaction. Microbiol. Res. 2009, 164, 174–179. [Google Scholar] [CrossRef]
- Boonsuk, P.; Payungporn, S.; Chieochansin, T.; Samransamruajkit, R.; Amonsin, A.; Songserm, T.; Chaisingh, A.; Chamnanpood, P.; Chutinimitkul, S.; Theamboonlers, A.; Poovorawan, Y. Detection of influenza virus types A and B and type A subtypes (H1, H3, and H5) by multiplex polymerase chain reaction. Tohoku J. Exp. Med. 2008, 215, 247–255. [Google Scholar] [CrossRef]
- Spackman, E.; Senne, D.A.; Bulaga, L.L.; Trock, S.; Suarez, D.L. Development of multiplex real-time RT-PCR as a diagnostic tool for avian influenza. Avian. Dis. 2003, 47, 1087–1090. [Google Scholar] [CrossRef]
- Payungporn, S.; Chutinimitkul, S.; Chaisingh, A.; Damrongwantanapokin, S.; Buranathai, C.; Amonsin, A.; Theamboonlers, A.; Poovorawan, Y. Single step multiplex real-time RT-PCR for H5N1 influenza A virus detection. J. Virol. Methods. 2006, 131, 143–147. [Google Scholar] [CrossRef]
- Wu, C.; Cheng, X.; He, J.; Lv, X.; Wang, J.; Deng, R.; Long, Q.; Wang, X. A multiplex real-time RT-PCR for detection and identification of influenza virus types A and B and subtypes H5 and N1. J. Virol. Methods 2008, 148, 81–88. [Google Scholar] [CrossRef]
- Abdelwhab el, S.M.; Erfan, A.M.; Grund, C.; Ziller, M.; Arafa, A.S.; Beer, M.; Aly, M.M.; Hafez, H.M.; Harder, T.C. Simultaneous detection and differentiation by multiplex real time RT-PCR of highly pathogenic avian influenza subtype H5N1 classic (clade 2.2.1 proper) and escape mutant (clade 2.2.1 variant) lineages in Egypt. Virol. J. 2010, 7, 260. [Google Scholar]
- Suwannakarn, K.; Payungporn, S.; Chieochansin, T.; Samransamruajkit, R.; Amonsin, A.; Songserm, T.; Chaisingh, A.; Chamnanpood, P.; Chutinimitkul, S.; Theamboonlers, A.; Poovorawan, Y. Typing (A/B) and subtyping (H1/H3/H5) of influenza A viruses by multiplex real-time RT-PCR assays. J. Virol. Methods 2008, 152, 25–31. [Google Scholar] [CrossRef]
- Wang, W.; Ren, P.; Mardi, S.; Hou, L.; Tsai, C.; Chan, K.H.; Cheng, P.; Sheng, J.; Buchy, P.; Sun, B.; Toyoda, T.; Lim, W.; Peiris, J.S.; Zhou, P.; Deubel, V. design of multiplexed detection assays for identification of avian influenza a virus subtypes pathogenic to humans by SmartCycler real-time reverse transcription-PCR. Journal of clinical microbiology 2009, 47, 86–92. [Google Scholar] [CrossRef]
- He, J.; Bose, M.E.; Beck, E.T.; Fan, J.; Tiwari, S.; Metallo, J.; Jurgens, L.A.; Kehl, S.C.; Ledeboer, N.; Kumar, S.; Weisburg, W.; Henrickson, K.J. Rapid multiplex reverse transcription-PCR typing of influenza A and B virus, and subtyping of influenza A virus into H1, 2, 3, 5, 7, 9, N1 (human), N1 (animal), N2, and N7, including typing of novel swine origin influenza A (H1N1) virus, during the 2009 outbreak in Milwaukee, Wisconsin. Journal of clinical microbiology 2009, 47, 2772–2778. [Google Scholar]
- Notomi, T.; Okayama, H.; Masubuchi, H.; Yonekawa, T.; Watanabe, K.; Amino, N.; Hase, T. Loop-mediated isothermal amplification of DNA. Nucleic. Acids. Res. 2000, 28, E63. [Google Scholar] [CrossRef]
- Imai, M.; Ninomiya, A.; Minekawa, H.; Notomi, T.; Ishizaki, T.; Van Tu, P.; Tien, N.T.; Tashiro, M.; Odagiri, T. Rapid diagnosis of H5N1 avian influenza virus infection by newly developed influenza H5 hemagglutinin gene-specific loop-mediated isothermal amplification method. J. Virol. Methods 2007, 141, 173–180. [Google Scholar] [CrossRef]
- Dinh, D.T.; Le, M.T.; Vuong, C.D.; Hasebe, F.; Morita, K. An Updated Loop-Mediated Isothermal Amplification Method for Rapid Diagnosis of H5N1 Avian Influenza Viruses. Trop. Med. Health. 2011, 39, 3–7. [Google Scholar] [CrossRef]
- Poon, L.L.; Leung, C.S.; Chan, K.H.; Lee, J.H.; Yuen, K.Y.; Guan, Y.; Peiris, J.S. Detection of human influenza A viruses by loop-mediated isothermal amplification. Journal of clinical microbiology 2005, 43, 427–430. [Google Scholar] [CrossRef]
- Ito, M.; Watanabe, M.; Nakagawa, N.; Ihara, T.; Okuno, Y. Rapid detection and typing of influenza A and B by loop-mediated isothermal amplification: comparison with immunochromatography and virus isolation. J. Virol. Methods 2006, 135, 272–275. [Google Scholar] [CrossRef]
- Chen, H.T.; Zhang, J.; Sun, D.H.; Ma, L.N.; Liu, X.T.; Cai, X.P.; Liu, Y.S. Development of reverse transcription loop-mediated isothermal amplification for rapid detection of H9 avian influenza virus. J. Virol. Methods 2008, 151, 200–203. [Google Scholar] [CrossRef]
- Postel, A.; Letzel, T.; Frischmann, S.; Grund, C.; Beer, M.; Harder, T. Evaluation of two commercial loop-mediated isothermal amplification assays for detection of avian influenza H5 and H7 hemagglutinin genes. J. Vet. Diagn. Invest. 2010, 22, 61–66. [Google Scholar] [CrossRef]
- Nagatani, N.; Yamanaka, K.; Saito, M.; Koketsu, R.; Sasaki, T.; Ikuta, K.; Miyahara, T.; Tamiya, E. Semi-real time electrochemical monitoring for influenza virus RNA by reverse transcription loop-mediated isothermal amplification using a USB powered portable potentiostat. Analyst 2011, 136, 5143–5150. [Google Scholar] [CrossRef]
- Compton, J. Nucleic acid sequence-based amplification. Nature 1991, 350, 91–92. [Google Scholar] [CrossRef]
- Collins, R.A.; Ko, L.S.; So, K.L.; Ellis, T.; Lau, L.T.; Yu, A.C. Detection of highly pathogenic and low pathogenic avian influenza subtype H5 (Eurasian lineage) using NASBA. J. Virol. Methods 2002, 103, 213–225. [Google Scholar] [CrossRef]
- Collins, R.A.; Ko, L.S.; So, K.L.; Ellis, T.; Lau, L.T.; Yu, A.C. A NASBA method to detect high- and low-pathogenicity H5 avian influenza viruses. Avian Dis 2003, 47, 1069–1074. [Google Scholar] [CrossRef]
- Collins, R.A.; Ko, L.S.; Fung, K.Y.; Chan, K.Y.; Xing, J.; Lau, L.T.; Yu, A.C. Rapid and sensitive detection of avian influenza virus subtype H7 using NASBA. Biochem. Biophys. Res. Commun. 2003, 300, 507–515. [Google Scholar] [CrossRef]
- Lau, L.T.; Banks, J.; Aherne, R.; Brown, I.H.; Dillon, N.; Collins, R.A.; Chan, K.Y.; Fung, Y.W.; Xing, J.; Yu, A.C. Nucleic acid sequence-based amplification methods to detect avian influenza virus. Biochem. Biophys. Res. Commun. 2004, 313, 336–342. [Google Scholar] [CrossRef]
- Chantratita, W.; Sukasem, C.; Kaewpongsri, S.; Srichunrusami, C.; Pairoj, W.; Thitithanyanont, A.; Chaichoune, K.; Ratanakron, P.; Songserm, T.; Damrongwatanapokin, S.; Landt, O. Qualitative detection of avian influenza A (H5N1) viruses: a comparative evaluation of four real-time nucleic acid amplification methods. Mol. Cell Probes 2008, 22, 287–293. [Google Scholar] [CrossRef]
- Moore, C.; Telles, J.N.; Corden, S.; Gao, R.B.; Vernet, G.; Van Aarle, P.; Shu, Y.L. Development and validation of a commercial real-time NASBA assay for the rapid confirmation of influenza A H5N1 virus in clinical samples. J. Virol. Methods 2010, 170, 173–176. [Google Scholar] [CrossRef]
- Moore, C.; Corden, S.; Sinha, J.; Jones, R. Dry cotton or flocked respiratory swabs as a simple collection technique for the molecular detection of respiratory viruses using real-time NASBA. J. Virol. Methods 2008, 153, 84–89. [Google Scholar] [CrossRef]
- van Deursen, P.B.; Gunther, A.W.; van Riel, C.C.; van der Eijnden, M.M.; Vos, H.L.; van Gemen, B.; van Strijp, D.A.; Tackent, N.M.; Bertina, R.M. A novel quantitative multiplex NASBA method: application to measuring tissue factor and CD14 mRNA levels in human monocytes. Nucleic. Acids. Res. 1999, 27, e15. [Google Scholar]
- Lau, L.T.; Feng, X.Y.; Lam, T.Y.; Hui, H.K.; Yu, A.C. Development of multiplex nucleic acid sequence-based amplification for detection of human respiratory tract viruses. J. Virol. Methods 2010, 168, 251–254. [Google Scholar] [CrossRef]
- Pease, A.C.; Solas, D.; Sullivan, E.J.; Cronin, M.T.; Holmes, C.P.; Fodor, S.P. Light-generated oligonucleotide arrays for rapid DNA sequence analysis. Proc. Natl. Acad. Sci. U S A 1994, 91, 5022–5026. [Google Scholar]
- Schena, M.; Shalon, D.; Davis, R.W.; Brown, P.O. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 1995, 270, 467–470. [Google Scholar]
- Li, J.; Chen, S.; Evans, D.H. Typing and subtyping influenza virus using DNA microarrays and multiplex reverse transcriptase PCR. Journal of clinical microbiology 2001, 39, 696–704. [Google Scholar] [CrossRef]
- Kessler, N.; Ferraris, O.; Palmer, K.; Marsh, W.; Steel, A. Use of the DNA flow-thru chip, a three-dimensional biochip, for typing and subtyping of influenza viruses. Journal of clinical microbiology 2004, 42, 2173–2185. [Google Scholar] [CrossRef]
- Ryabinin, V.A.; Kostina, E.V.; Maksakova, G.A.; Neverov, A.A.; Chumakov, K.M.; Sinyakov, A.N. Universal oligonucleotide microarray for sub-typing of Influenza A virus. PLoS One 2011, 6, e17529. [Google Scholar]
- Townsend, M.B.; Dawson, E.D.; Mehlmann, M.; Smagala, J.A.; Dankbar, D.M.; Moore, C.L.; Smith, C.B.; Cox, N.J.; Kuchta, R.D.; Rowlen, K.L. Experimental evaluation of the FluChip diagnostic microarray for influenza virus surveillance. Journal of clinical microbiology 2006, 44, 2863–2871. [Google Scholar]
- Mehlmann, M.; Dawson, E.D.; Townsend, M.B.; Smagala, J.A.; Moore, C.L.; Smith, C.B.; Cox, N.J.; Kuchta, R.D.; Rowlen, K.L. Robust sequence selection method used to develop the FluChip diagnostic microarray for influenza virus. Journal of clinical microbiology 2006, 44, 2857–2862. [Google Scholar] [CrossRef]
- Huang, Y.; Tang, H.; Duffy, S.; Hong, Y.; Norman, S.; Ghosh, M.; He, J.; Bose, M.; Henrickson, K.J.; Fan, J.; Kraft, A.J.; Weisburg, W.G.; Mather, E.L. Multiplex assay for simultaneously typing and subtyping influenza viruses by use of an electronic microarray. Journal of clinical microbiology 2009, 47, 390–396. [Google Scholar]
- Dawson, E.D.; Moore, C.L.; Smagala, J.A.; Dankbar, D.M.; Mehlmann, M.; Townsend, M.B.; Smith, C.B.; Cox, N.J.; Kuchta, R.D.; Rowlen, K.L. MChip: a tool for influenza surveillance. Anal. Chem. 2006, 78, 7610–7615. [Google Scholar]
- Dawson, E.D.; Moore, C.L.; Dankbar, D.M.; Mehlmann, M.; Townsend, M.B.; Smagala, J.A.; Smith, C.B.; Cox, N.J.; Kuchta, R.D.; Rowlen, K.L. Identification of A/H5N1 influenza viruses using a single gene diagnostic microarray. Anal. Chem. 2007, 79, 378–384. [Google Scholar]
- Moore, C.L.; Smagala, J.A.; Smith, C.B.; Dawson, E.D.; Cox, N.J.; Kuchta, R.D.; Rowlen, K.L. Evaluation of MChip with historic subtype H1N1 influenza A viruses, including the 1918 "Spanish Flu" strain. Journal of clinical microbiology 2007, 45, 3807–3810. [Google Scholar]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Sakurai, A.; Shibasaki, F. Updated Values for Molecular Diagnosis for Highly Pathogenic Avian Influenza Virus. Viruses 2012, 4, 1235-1257. https://doi.org/10.3390/v4081235
Sakurai A, Shibasaki F. Updated Values for Molecular Diagnosis for Highly Pathogenic Avian Influenza Virus. Viruses. 2012; 4(8):1235-1257. https://doi.org/10.3390/v4081235
Chicago/Turabian StyleSakurai, Akira, and Futoshi Shibasaki. 2012. "Updated Values for Molecular Diagnosis for Highly Pathogenic Avian Influenza Virus" Viruses 4, no. 8: 1235-1257. https://doi.org/10.3390/v4081235




