Authentication of the R06E Fruit Bat Cell Line
Abstract
:1. Introduction
2. Results
2.1. Screen for Cross Species Contamination in Mammalian Cells
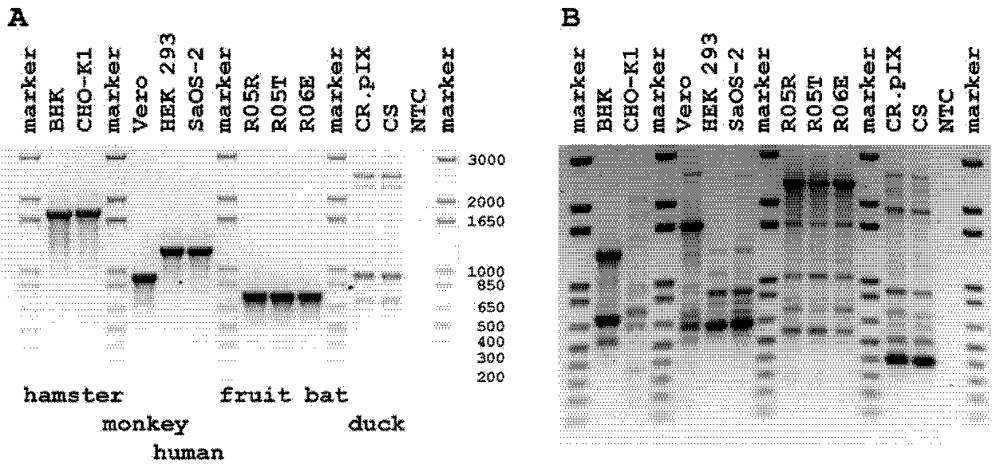
2.2. Authentication of R06E

2.3. Cocultivation of Vero.GFP and R06E Cells
2.4. Variable Permissivity of R06E Cells

3. Experimental Section
3.1. PCR and Primers
3.2. Cocultivation
3.3. Virus Infections
4. Discussion and Conclusions
Acknowledgments
Conflict of Interest
References
- Hayman, D.T.; Suu-Ire, R.; Breed, A.C.; McEachern, J.A.; Wang, L.; Wood, J.L.; Cunningham, A.A. Evidence of henipavirus infection in West African fruit bats. PLoS One 2008, 3, e2739. [Google Scholar]
- Yob, J.M.; Field, H.; Rashdi, A.M.; Morrissy, C.; van der Heide, B.; Rota, P.; bin Adzhar, A.; White, J.; Daniels, P.; Jamaluddin, A.; et al. Nipah virus infection in bats (order Chiroptera) in peninsular Malaysia. Emerg. Infect. Dis. 2001, 7, 439–441. [Google Scholar]
- Leroy, E.M.; Kumulungui, B.; Pourrut, X.; Rouquet, P.; Hassanin, A.; Yaba, P.; Delicat, A.; Paweska, J.T.; Gonzalez, J.P.; Swanepoel, R. Fruit bats as reservoirs of Ebola virus. Nature 2005, 438, 575–576. [Google Scholar]
- Towner, J.S.; Pourrut, X.; Albarino, C.G.; Nkogue, C.N.; Bird, B.H.; Grard, G.; Ksiazek, T.G.; Gonzalez, J.P.; Nichol, S.T.; Leroy, E.M. Marburg virus infection detected in a common African bat. PLoS One 2007, 2, e764. [Google Scholar]
- Towner, J.S.; Amman, B.R.; Sealy, T.K.; Carroll, S.A.; Comer, J.A.; Kemp, A.; Swanepoel, R.; Paddock, C.D.; Balinandi, S.; Khristova, M.L.; et al. Isolation of genetically diverse Marburg viruses from Egyptian fruit bats. PLoS Pathog. 2009, 5, e1000536. [Google Scholar]
- Chu, D.K.; Poon, L.L.; Chan, K.H.; Chen, H.; Guan, Y.; Yuen, K.Y.; Peiris, J.S. Coronaviruses in bent-winged bats (Miniopterus spp.). J. Gen. Virol. 2006, 87, 2461–2466. [Google Scholar]
- Li, W.; Shi, Z.; Yu, M.; Ren, W.; Smith, C.; Epstein, J.H.; Wang, H.; Crameri, G.; Hu, Z.; Zhang, H.; et al. Bats are natural reservoirs of SARS-like coronaviruses. Science 2005, 310, 676–679. [Google Scholar]
- Amengual, B.; Bourhy, H.; Lopez-Roig, M.; Serra-Cobo, J. Temporal dynamics of European bat Lyssavirus type 1 and survival of Myotis myotis bats in natural colonies. PLoS One 2007, 2, e566. [Google Scholar]
- Arguin, P.M.; Murray-Lillibridge, K.; Miranda, M.E.; Smith, J.S.; Calaor, A.B.; Rupprecht, C.E. Serologic evidence of Lyssavirus infections among bats, the Philippines. Emerg. Infect. Dis. 2002, 8, 258–262. [Google Scholar]
- van der Poel, W.H.; van der Heide, R.; Verstraten, E.R.; Takumi, K.; Lina, P.H.; Kramps, J.A. European bat lyssaviruses, The Netherlands. Emerg. Infect. Dis. 2005, 11, 1854–1859. [Google Scholar]
- Best, S.M.; Kerr, P.J. Coevolution of host and virus: the pathogenesis of virulent and attenuated strains of myxoma virus in resistant and susceptible European rabbits. Virology 2000, 267, 36–48. [Google Scholar]
- Chua, K.B.; Koh, C.L.; Hooi, P.S.; Wee, K.F.; Khong, J.H.; Chua, B.H.; Chan, Y.P.; Lim, M.E.; Lam, S.K. Isolation of Nipah virus from Malaysian Island flying-foxes. Microbes Infect. 2002, 4, 145–151. [Google Scholar]
- Reynes, J.M.; Counor, D.; Ong, S.; Faure, C.; Seng, V.; Molia, S.; Walston, J.; Georges-Courbot, M.C.; Deubel, V.; Sarthou, J.L. Nipah virus in Lyle’s flying foxes, Cambodia. Emerg. Infect. Dis. 2005, 11, 1042–1047. [Google Scholar]
- Brookes, S.M.; Aegerter, J.N.; Smith, G.C.; Healy, D.M.; Jolliffe, T.A.; Swift, S.M.; Mackie, I.J.; Pritchard, J.S.; Racey, P.A.; Moore, N.P.; et al. European bat lyssavirus in Scottish bats. Emerg. Infect. Dis. 2005, 11, 572–578. [Google Scholar]
- Jordan, I.; Horn, D.; Oehmke, S.; Leendertz, F.H.; Sandig, V. Cell lines from the Egyptian fruit bat are permissive for modified vaccinia Ankara. Virus Res. 2009, 145, 54–62. [Google Scholar]
- Nakamura, Y.; Leppert, M.; O’Connell, P.; Wolff, R.; Holm, T.; Culver, M.; Martin, C.; Fujimoto, E.; Hoff, M.; Kumlin, E.; et al. Variable number of tandem repeat (VNTR) markers for human gene mapping. Science 1987, 235, 1616–1622. [Google Scholar]
- Housley, D.J.; Zalewski, Z.A.; Beckett, S.E.; Venta, P.J. Design factors that influence PCR amplification success of cross-species primers among 1147 mammalian primer pairs. BMC Genomics 2006, 7, 253. [Google Scholar]
- Weising, K.; Atkinson, R.G.; Gardner, R.C. Genomic fingerprinting by microsatellite-primed PCR: A critical evaluation. PCR Methods Appl. 1995, 4, 249–255. [Google Scholar]
- Gong, T.W.; Besirli, C.G.; Lomax, M.I. MACF1 gene structure: A hybrid of plectin and dystrophin. Mamm. Genome 2001, 12, 852–861. [Google Scholar]
- Meyer, H.; Sutter, G.; Mayr, A. Mapping of deletions in the genome of the highly attenuated vaccinia virus MVA and their influence on virulence. J. Gen. Virol. 1991, 72, 1031–1038. [Google Scholar]
- Mayr, A.; Munz, E. Changes in the vaccinia virus through continuing passages in chick embryo fibroblast cultures. Zentralbl. Bakteriol. Orig. 1964, 195, 24–35. [Google Scholar]
- Jordan, I.; Vos, A.; Beilfuss, S.; Neubert, A.; Breul, S.; Sandig, V. An avian cell line designed for production of highly attenuated viruses. Vaccine 2009, 27, 748–756. [Google Scholar]
- Kobune, F.; Sakata, H.; Sugiura, A. Marmoset lymphoblastoid cells as a sensitive host for isolation of measles virus. J. Virol. 1990, 64, 700–705. [Google Scholar]
- Suzuki, S.; Sawa, H.; Komagome, R.; Orba, Y.; Yamada, M.; Okada, Y.; Ishida, Y.; Nishihara, H.; Tanaka, S.; Nagashima, K. Broad distribution of the JC virus receptor contrasts with a marked cellular restriction of virus replication. Virology 2001, 286, 100–112. [Google Scholar]
- Kouomou, D.W.; Wild, T.F. Adaptation of wild-type measles virus to tissue culture. J. Virol. 2002, 76, 1505–1509. [Google Scholar]
- Chen, W.; Baric, R.S. Molecular anatomy of mouse hepatitis virus persistence: coevolution of increased host cell resistance and virus virulence. J. Virol. 1996, 70, 3947–3960. [Google Scholar]
- Cagle, C.; To, T.L.; Nguyen, T.; Wasilenko, J.; Adams, S.C.; Cardona, C.J.; Spackman, E.; Suarez, D.L.; Pantin-Jackwood, M.J. Pekin and Muscovy ducks respond differently to vaccination with a H5N1 highly pathogenic avian influenza (HPAI) commercial inactivated vaccine. Vaccine 2011, 29, 6549–6557. [Google Scholar]
- Barber, M.R.; Aldridge, J.R., Jr.; Webster, R.G.; Magor, K.E. Association of RIG-I with innate immunity of ducks to influenza. Proc. Natl. Acad. Sci. USA 2010, 107, 5913–5918. [Google Scholar]
- Boonstra, J.J.; van der Velden, A.W.; Beerens, E.C.; van Marion, R.; Morita-Fujimura, Y.; Matsui, Y.; Nishihira, T.; Tselepis, C.; Hainaut, P.; Lowe, A.W.; et al. Mistaken identity of widely used esophageal adenocarcinoma cell line TE-7. Cancer Res. 2007, 67, 7996–8001. [Google Scholar]
- Perkel, J.M. Curing cell lines. Biotechniques 2011, 51, 85–90. [Google Scholar]
- MacLeod, R.A.; Dirks, W.G.; Matsuo, Y.; Kaufmann, M.; Milch, H.; Drexler, H.G. Widespread intraspecies cross-contamination of human tumor cell lines arising at source. Int. J. Cancer 1999, 83, 555–563. [Google Scholar]
- Lucey, B.P.; Nelson-Rees, W.A.; Hutchins, G.M. Henrietta Lacks, HeLa cells, and cell culture contamination. Arch. Pathol. Lab. Med. 2009, 133, 1463–1467. [Google Scholar]
- Phuchareon, J.; Ohta, Y.; Woo, J.M.; Eisele, D.W.; Tetsu, O. Genetic profiling reveals cross-contamination and misidentification of 6 adenoid cystic carcinoma cell lines: ACC2, ACC3, ACCM, ACCNS, ACCS and CAC2 . PLoS One 2009, 4, e6040. [Google Scholar]
- Krahling, V.; Dolnik, O.; Kolesnikova, L.; Schmidt-Chanasit, J.; Jordan, I.; Sandig, V.; Gunther, S.; Becker, S. Establishment of fruit bat cells (Rousettus aegyptiacus) as a model system for the investigation of filoviral infection. PLoS Negl. Trop. Dis. 2010, 4, e802. [Google Scholar]
- Nims, R.W.; Shoemaker, A.P.; Bauernschub, M.A.; Rec, L.J.; Harbell, J.W. Sensitivity of isoenzyme analysis for the detection of interspecies cell line cross-contamination. In Vitro Cell. Dev. Biol. Anim. 1998, 34, 35–39. [Google Scholar]
- Emeny, J.M.; Morgan, M.J. Regulation of the interferon system: Evidence that Vero cells have a genetic defect in interferon production. J. Gen. Virol. 1979, 43, 247–252. [Google Scholar]
- Teferedegne, B.; Murata, H.; Quinones, M.; Peden, K.; Lewis, A.M. Patterns of microRNA expression in non-human primate cells correlate with neoplastic development in vitro. PLoS One 2010, 5, e14416. [Google Scholar]
- Tanaka, T.; Takahashi, M.; Takahashi, H.; Ichiyama, K.; Hoshino, Y.; Nagashima, S.; Mizuo, H.; Okamoto, H. Development and characterization of a genotype 4 hepatitis E virus cell culture system using a HE-JF5/15F strain recovered from a fulminant hepatitis patient. J. Clin. Microbiol. 2009, 47, 1906–1910. [Google Scholar]
- Lohmann, V.; Korner, F.; Dobierzewska, A.; Bartenschlager, R. Mutations in hepatitis C virus RNAs conferring cell culture adaptation. J. Virol. 2001, 75, 1437–1449. [Google Scholar]
- Chaurushiya, M.S.; Weitzman, M.D. Viral manipulation of DNA repair and cell cycle checkpoints. DNA Repair (Amst) 2009, 8, 1166–1176. [Google Scholar]
- Barre, B.; Perkins, N.D. A cell cycle regulatory network controlling NF-kappaB subunit activity and function. EMBO J. 2007, 26, 4841–4855. [Google Scholar]
- Joyce, D.; Albanese, C.; Steer, J.; Fu, M.; Bouzahzah, B.; Pestell, R.G. NF-kappaB and cell-cycle regulation: the cyclin connection. Cytokine Growth Factor Rev. 2001, 12, 73–90. [Google Scholar]
- Lin, S.; Wu, M.; Xu, Y.; Xiong, W.; Yi, Z.; Zhang, X.; Zhenghong, Y. Inhibition of hepatitis B virus replication by MyD88 is mediated by nuclear factor-kappaB activation. Biochim. Biophys. Acta 2007, 1772, 1150–1157. [Google Scholar]
- Nimmerjahn, F.; Dudziak, D.; Dirmeier, U.; Hobom, G.; Riedel, A.; Schlee, M.; Staudt, L.M.; Rosenwald, A.; Behrends, U.; Bornkamm, G.W.; et al. Active NF-kappaB signalling is a prerequisite for influenza virus infection. J. Gen. Virol. 2004, 85, 2347–2356. [Google Scholar]
- Brown, H.J.; Song, M.J.; Deng, H.; Wu, T.T.; Cheng, G.; Sun, R. NF-kappaB inhibits gammaherpesvirus lytic replication. J. Virol. 2003, 77, 8532–8540. [Google Scholar]
- Manohar, M.; Orrison, B.; Peden, K.; Lewis, A.M., Jr. Assessing the tumorigenic phenotype of VERO cells in adult and newborn nude mice. Biologicals 2008, 36, 65–72. [Google Scholar]
- Omeir, R.L.; Teferedegne, B.; Foseh, G.S.; Beren, J.J.; Snoy, P.J.; Brinster, L.R.; Cook, J.L.; Peden, K.; Lewis, A.M., Jr. Heterogeneity of the tumorigenic phenotype expressed by Madin-Darby canine kidney cells. Comp. Med. 2011, 61, 243–250. [Google Scholar]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Jordan, I.; Munster, V.J.; Sandig, V. Authentication of the R06E Fruit Bat Cell Line. Viruses 2012, 4, 889-900. https://doi.org/10.3390/v4050889
Jordan I, Munster VJ, Sandig V. Authentication of the R06E Fruit Bat Cell Line. Viruses. 2012; 4(5):889-900. https://doi.org/10.3390/v4050889
Chicago/Turabian StyleJordan, Ingo, Vincent J. Munster, and Volker Sandig. 2012. "Authentication of the R06E Fruit Bat Cell Line" Viruses 4, no. 5: 889-900. https://doi.org/10.3390/v4050889




