Metabolomic Profiles of Dinophysis acuminata and Dinophysis acuta Using Non-Targeted High-Resolution Mass Spectrometry: Effect of Nutritional Status and Prey
Abstract
:1. Introduction
2. Results
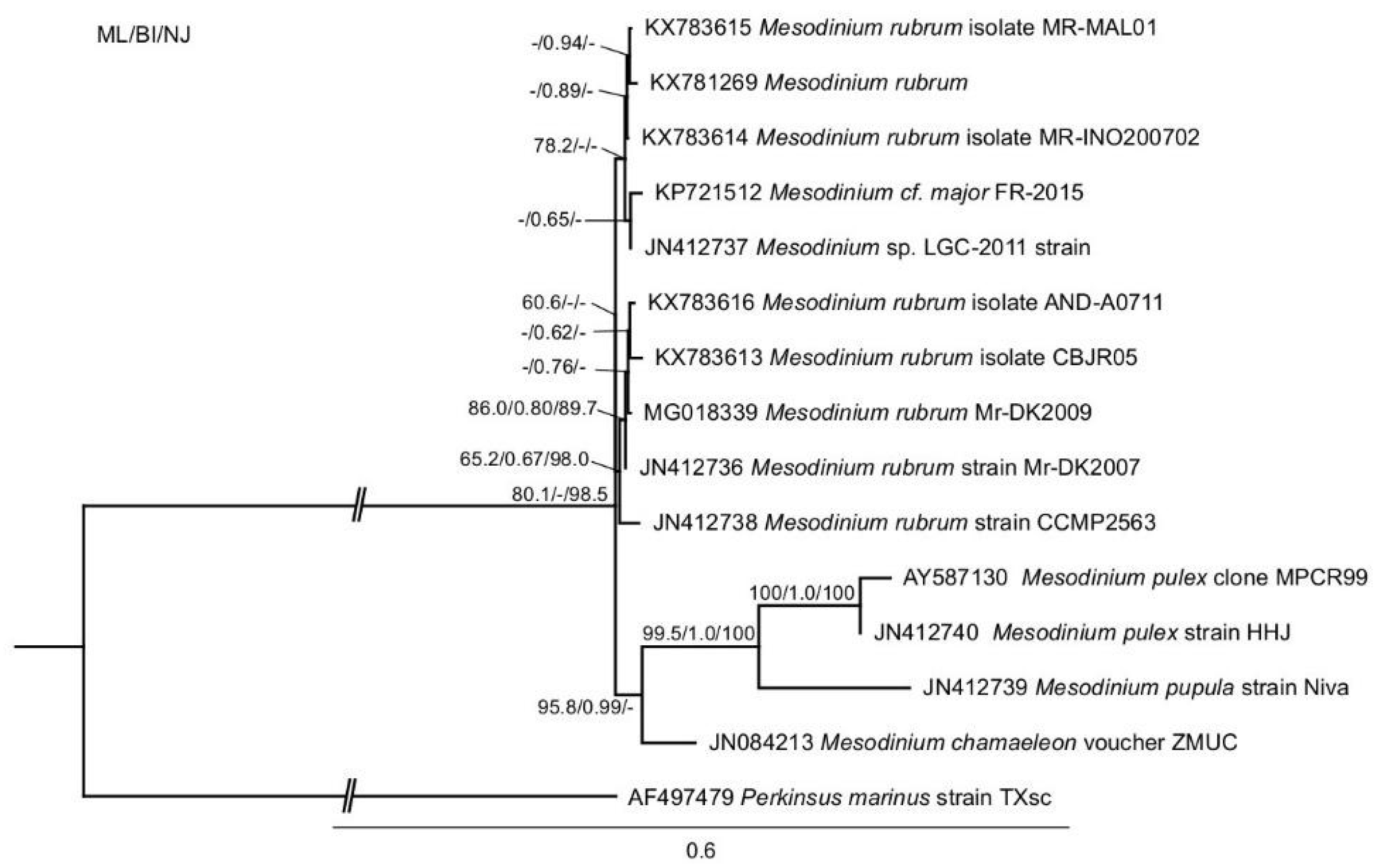
2.1. Phylogenetic Analysis
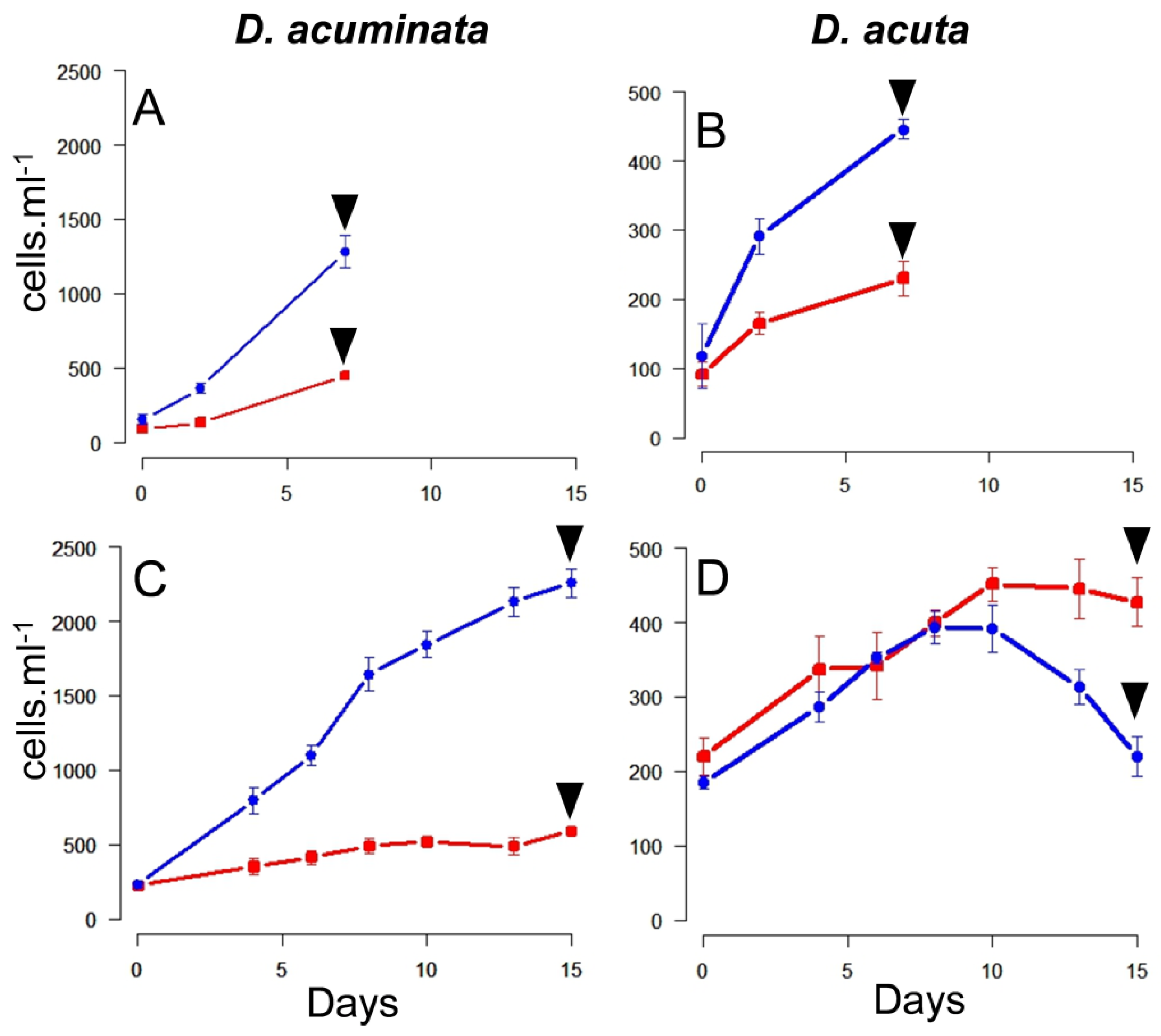
2.2. Growth Curves
2.3. Quantitative Toxin Analysis Using Low Resolution Mass Spectrometry
Esters of OA-Group Toxins
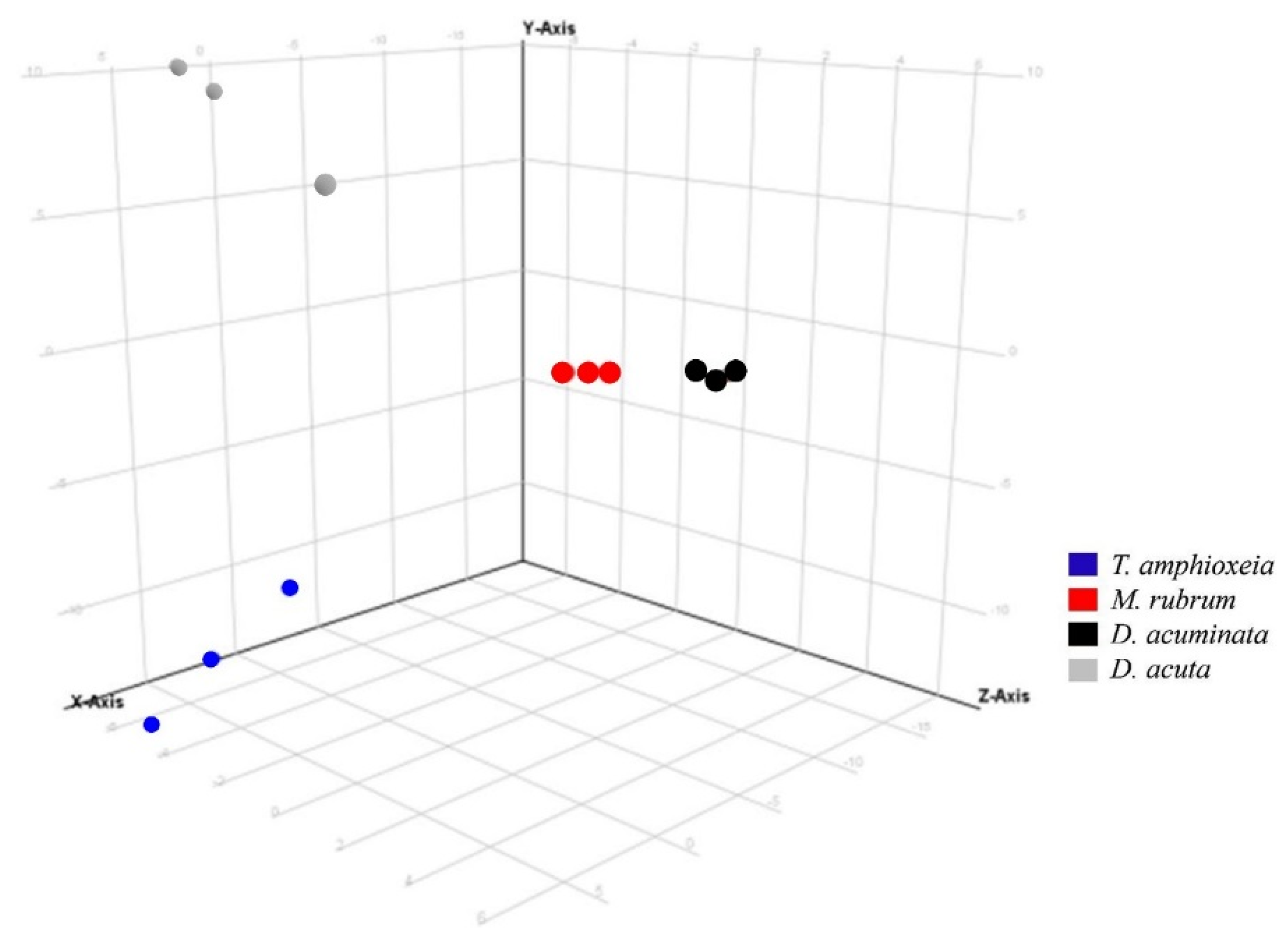
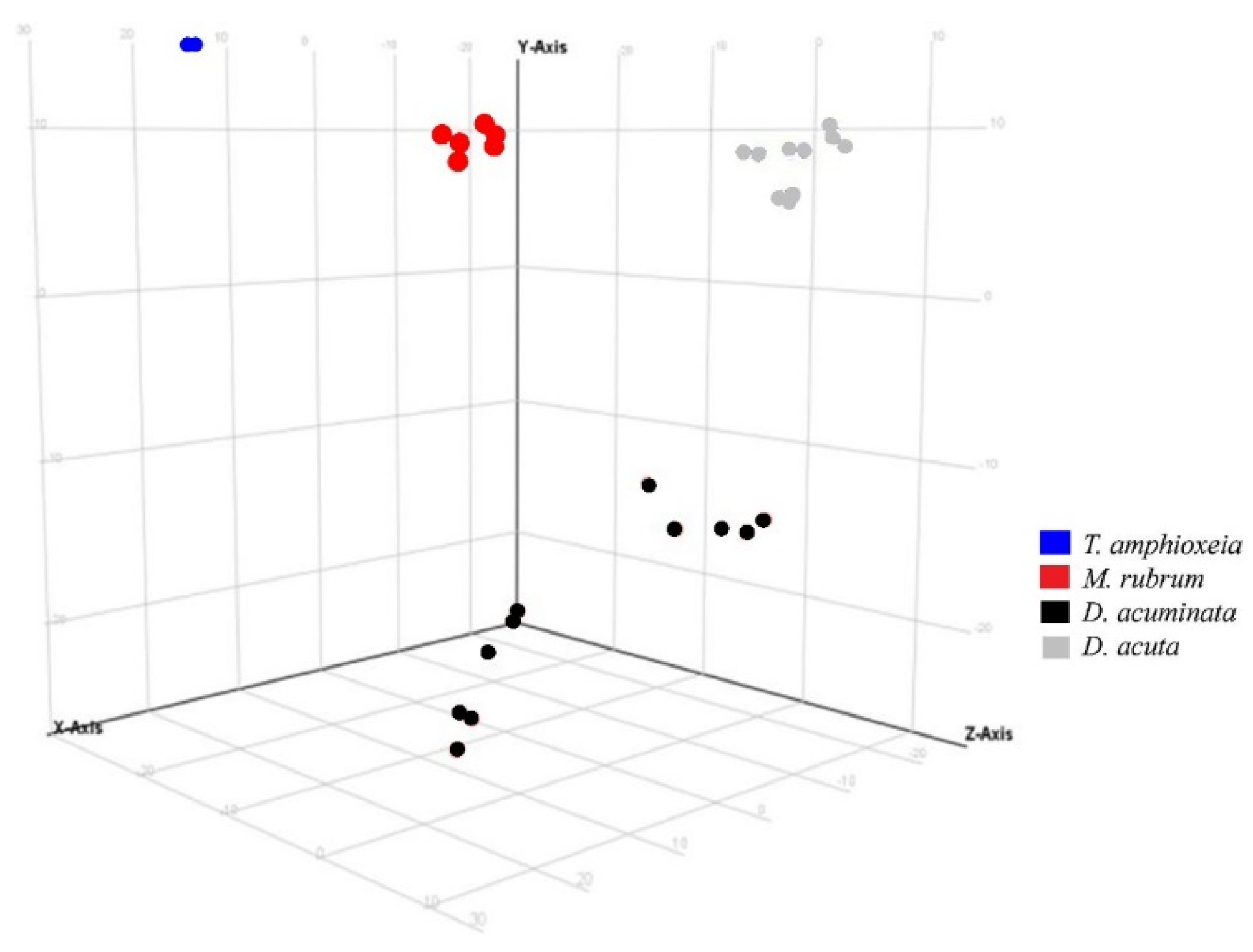
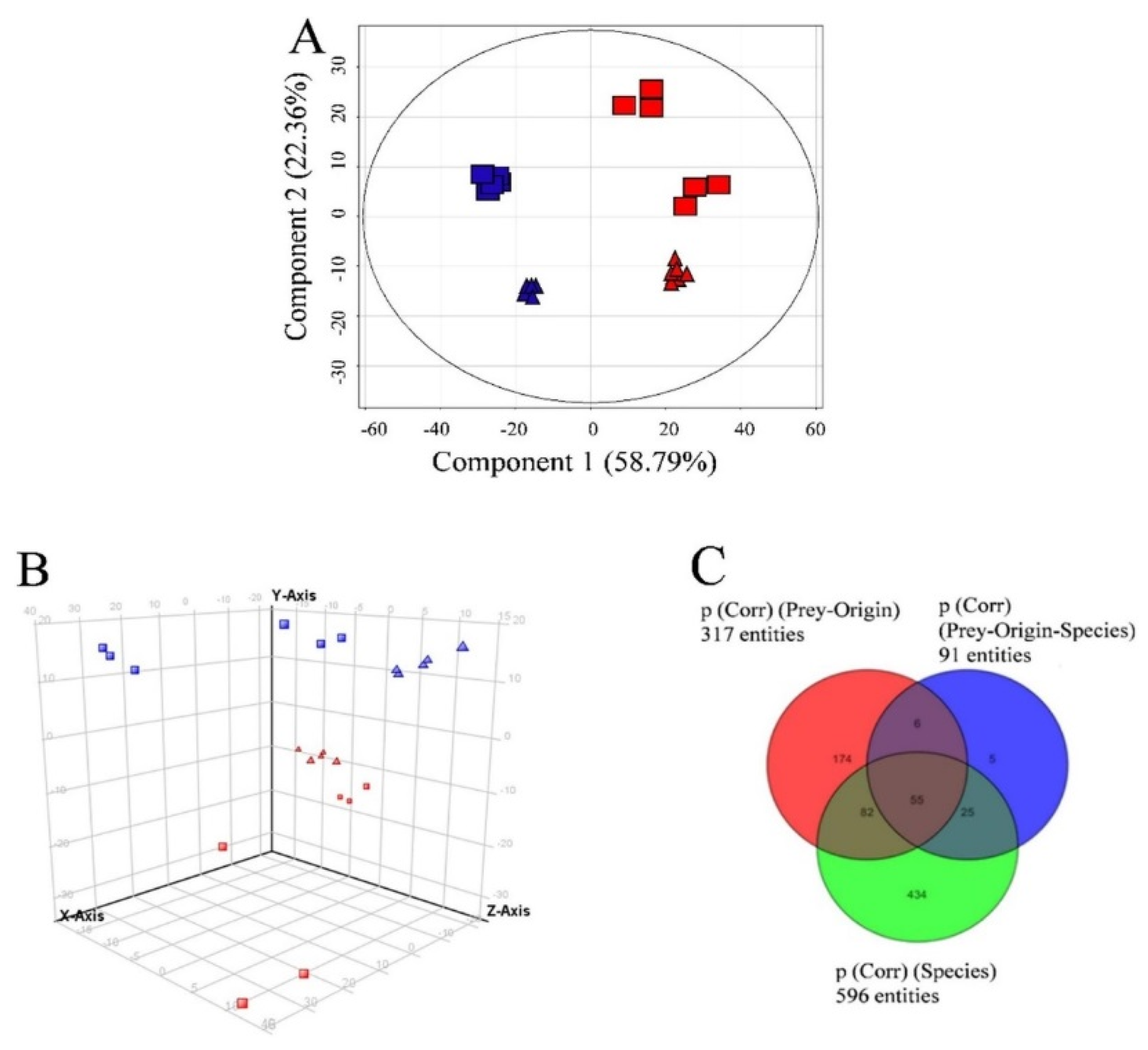
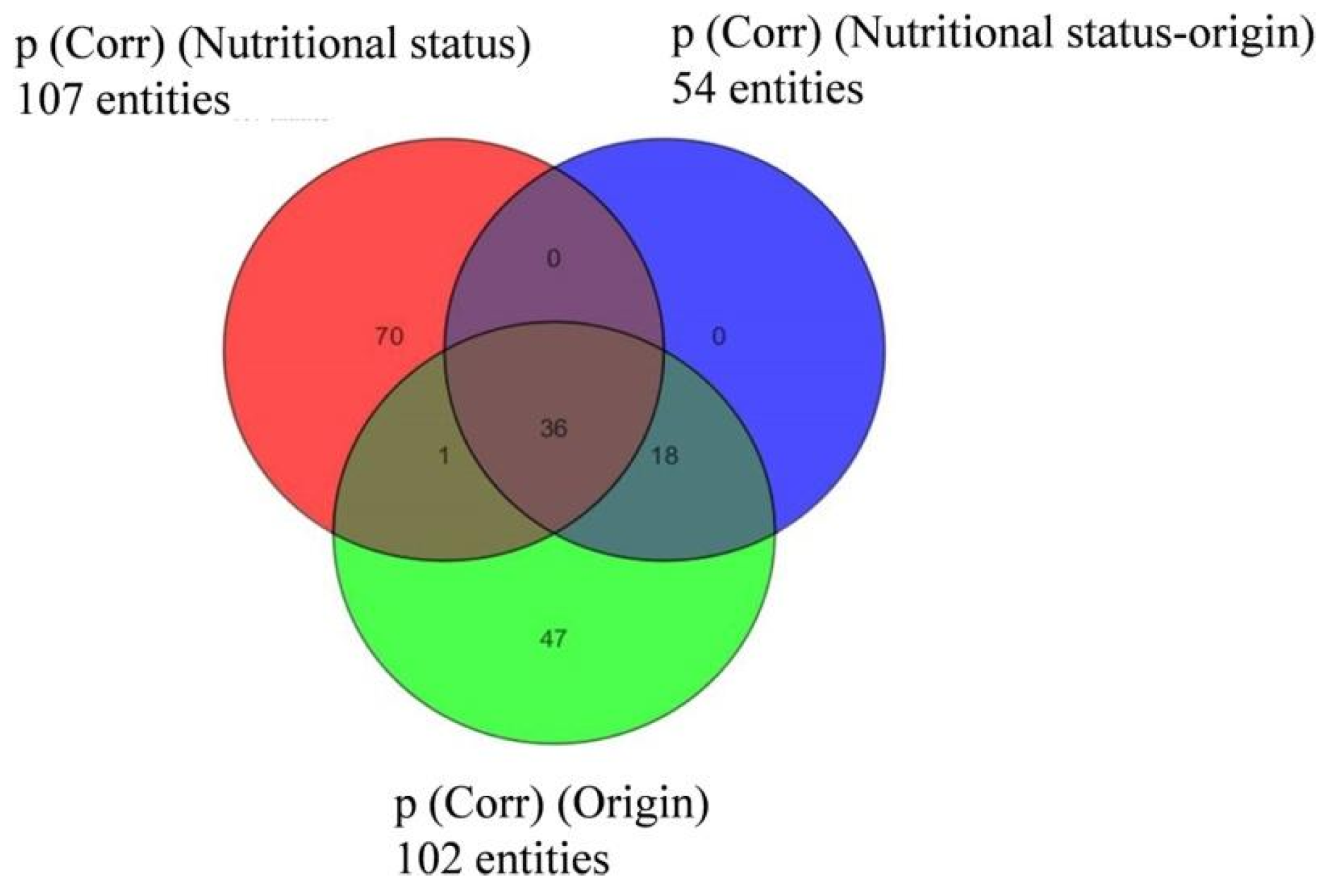
2.4. Data Treatment for Non-Targeted Liquid Chromatography High Resolution Mass Spectrometry
2.4.1. Compounds with an Already Known Physiological Meaning
Common to All Species
Compounds Exclusive to the Ciliate Mesodinium
Compounds Exclusive to Teleaulax amphioxeia
Compounds Exclusive to One of the Two Species of Dinophysis
2.4.2. Compounds First Found in Other Marine/Terrestrial Organisms
Compounds Present in Both Species
Compounds Exclusive to One of the Two Species of Dinophysis
3. Discussion
3.1. Growth Curves
3.2. Toxin Quota and Target Analysis
3.3. Semi-Quantitative Non-Targeted Analysis: Comparison of Larger Parts of Metabolomes
4. Material and Methods
4.1. Cultures
4.2. DNA Extraction, Polymerase Chain Reaction (PCR), 18S Sequencing and Phylogenetic Analysis of Mesodinium
4.3. Biovolume
4.4. Experiment 1 (Baseline Study)
4.5. Experiment 2 (Growth Curves)
4.6. Toxin Analysis
4.6.1. Extraction and Hydrolysis
4.6.2. High Resolution Mass Spectral Analysis—System A: (QTOF 6550)
4.6.3. Low Resolution Tandem Mass Spectrometry: System B (API 4000 QTrap)
4.7. Data Treatment for Non-Targeted Liquid Chromatography High Resolution Mass Spectrometry
4.7.1. Feature Identification and Variables’ Selection
5. Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Van Egmond, H.; Aune, T.; Lassus, P.; Speijers, G.; Waldock, M. Paralytic and diarrhoeic shellfish poisons: Occurrence in Europe, toxicity, analysis and regulation. J. Nat. Toxins 1993, 2, 41–83. [Google Scholar]
- Reguera, B.; Riobó, P.; Rodríguez, F.; Díaz, P.; Pizarro, G.; Paz, B.; Franco, J.; Blanco, J. Dinophysis toxins: Causative organisms, distribution and fate in shellfish. Mar. Drugs 2014, 12, 394–461. [Google Scholar] [CrossRef] [PubMed]
- López, B.D.; Methion, S. The impact of shellfish farming on common bottlenose dolphins’ use of habitat. Mar. Biol. 2017, 164, 83. [Google Scholar] [CrossRef]
- Blanco, J.; Correa, J.; Muñíz, S.; Mariño, C.; Martín, H.; Arévalo, F. Evaluación del impacto de los métodos y niveles utilizados para el control de toxinas en el mejillón. Revista Galega dos Recursos Mariños 2013, 3, 1–55. (In Spanish) [Google Scholar]
- Park, M.G.; Kim, S.; Kim, H.S.; Myung, G.; Kang, Y.G.; Yih, W. First successful culture of the marine dinoflagellate Dinophysis acuminata. Aquat. Microb. Ecol. 2006, 45, 101–106. [Google Scholar] [CrossRef]
- Riisgaard, K.; Hansen, P.J. Role of food uptake for photosynthesis, growth and survival of the mixotrophic dinoflagellate Dinophysis acuminata. Mar. Ecol. Prog. Ser. 2009, 381, 51–62. [Google Scholar] [CrossRef]
- Kim, E.; Archibald, J.M. Plastid evolution: Gene transfer and the maintenance of ‘stolen’ organelles. BMC Biol. 2010, 8, 73. [Google Scholar] [CrossRef] [PubMed]
- Hattenrath-Lehmann, T.; Gobler, C.J. The contribution of inorganic and organic nutrients to the growth of a North American isolate of the mixotrophic dinoflagellate, Dinophysis acuminata. Limnol. Oceanogr. 2015, 60, 1588–1603. [Google Scholar] [CrossRef]
- Hansen, P.J. Dinophysis—A planktonic dinoflagellate genus which can act both as a prey and a predator of a ciliate. Mar. Ecol. Prog. Ser. 1991, 69, 201–204. [Google Scholar] [CrossRef]
- Wisecaver, J.H.; Hackett, J.D. Transcriptome analysis reveals nuclear-encoded proteins for the maintenance of temporary plastids in the dinoflagellate Dinophysis acuminata. BMC Genom. 2010, 11, 366. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Cuetos, L.; Moestrup, Ø.; Hansen, P.J.; Daugbjerg, N. The toxic dinoflagellate Dinophysis acuminata harbors permanent chloroplasts of cryptomonad origin, not kleptochloroplasts. Harmful Algae 2010, 9, 25–38. [Google Scholar] [CrossRef]
- Park, M.G.; Kim, M.; Kim, S.; Yih, W. Does Dinophysis caudata (Dinophyceae) have permanent plastids? J. Phycol. 2010, 46, 236–242. [Google Scholar] [CrossRef]
- Raho, N.; Jaén, D.; Mamán, L.; Rial, P.; Marín, I. psbA based molecular analysis of cross-feeding experiments suggests that Dinophysis acuta does not harbour permanent plastids. Harmful Algae 2014, 35, 20–28. [Google Scholar] [CrossRef]
- Takahashi, Y.; Takishita, K.; Koike, K.; Maruyama, T.; Nakayama, T.; Kobiyama, A.; Ogata, T. Development of molecular probes for Dinophysis (Dinophyceae) plastid: A tool to predict blooming and explore plastid origin. Mar. Biotechnol. 2005, 7, 95–103. [Google Scholar] [CrossRef] [PubMed]
- Minnhagen, S.; Janson, S. Genetic analyses of Dinophysis spp. support kleptoplastidy. FEMS Microbiol. Ecol. 2006, 57, 47–54. [Google Scholar] [CrossRef] [PubMed]
- Nishitani, G.; Nagai, S.; Baba, K.; Kiyokawa, S.; Kosaka, Y.; Miyamura, K.; Nishikawa, T.; Sakurada, K.; Shinada, A.; Kamiyama, T. High-level congruence of Myrionecta rubra prey and Dinophysis species plastid identities as revealed by genetic analyses of isolates from Japanese coastal waters. Appl. Environ. Microbiol. 2010, 76, 2791–2798. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.; Nam, S.W.; Shin, W.; Coats, D.W.; Park, M.G. Dinophysis caudata (Dinophyceae) sequesters and retains plastids from the mixotrophic ciliate prey Mesodinium rubrum. J. Phycol. 2012, 48, 569–579. [Google Scholar] [CrossRef] [PubMed]
- Stern, R.; Amorim, A.; Bresnan, E. , Diversity and plastid types in Dinophysis acuminata complex (Dinophyceae) in Scottish waters. Harmful Algae 2014, 39, 223–231. [Google Scholar] [CrossRef]
- Rial, P.; Laza-Martínez, A.; Reguera, B.; Raho, N.; Rodríguez, F. Origin of cryptophyte plastids in Dinophysis from Galician waters: Results from field and culture experiments. Aquat. Microb. Ecol. 2015, 76, 163–174. [Google Scholar] [CrossRef]
- Herfort, L.; Maxey, K.; Voorhees, I.; Simon, H.M.; Grobler, K.; Peterson, T.D.; Zuber, P. Use of highly specific molecular markers reveals positive correlation between abundances of Mesodinium cf. major and its preferred prey, Teleaulax amphioxeia, during red water blooms in the Columbia river estuary. J. Eukaryot. Microbiol. 2017, 64, 740–755. [Google Scholar] [PubMed]
- Stoecker, D.K.; Johnson, M.D.; de Vargas, C.; Not, F. Acquired phototrophy in aquatic protists. Aquat. Microb. Ecol. 2009, 57, 279–310. [Google Scholar] [CrossRef]
- Reguera, B.; Velo-Suárez, L.; Raine, R.; Park, M.G. Harmful Dinophysis species: A. review. Harmful Algae 2012, 14, 87–106. [Google Scholar] [CrossRef]
- Gustafson, D.E.; Stoecker, D.K.; Johnson, M.D.; Van Heukelem, W.F.; Sneider, K. Cryptophyte algae are robbed of their organelles by the marine ciliate Mesodinium rubrum. Nature 2000, 405, 1049–1052. [Google Scholar] [CrossRef] [PubMed]
- Taylor, F.; Blackbourn, D.; Blackbourn, J. The red-water ciliate Mesodinium rubrum and its “incomplete symbionts”: A review including new ultrastructural observations. J. Fish. Res. Board Can. 1971, 28, 391–407. [Google Scholar] [CrossRef]
- Hansen, P.J.; Moldrup, M.; Tarangkoon, W.; Garcia-Cuetos, L.; Moestrup, Ø. Direct evidence for symbiont sequestration in the marine red tide ciliate Mesodinium rubrum. Aquat. Microb. Ecol. 2012, 66, 63–75. [Google Scholar] [CrossRef]
- Kim, G.H.; Han, J.H.; Kim, B.; Han, J.W.; Nam, S.W.; Shin, W.; Park, J.W.; Yih, W. Cryptophyte gene regulation in the kleptoplastidic, karyokleptic ciliate Mesodinium rubrum. Harmful Algae 2016, 52, 23–33. [Google Scholar] [CrossRef] [PubMed]
- Park, M.; Park, J.; Kim, M.; Yih, W. Plastid retention and functionality in the dinoflagellates Dinophysis acuminata and Dinophysis caudata. J. Phycol. 2007, 43, 7–8. [Google Scholar]
- Johnson, M.D.; Beaudoin, D.J.; Laza-Martinez, A.; Dyhrman, S.T.; Fensin, E.; Lin, S.; Merculief, A.; Nagai, S.; Pompeu, M.; Setälä, O. The genetic diversity of Mesodinium and associated Cryptophytes. Front. Microbiol. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Goulitquer, S.; Potin, P.; Tonon, T. Mass spectrometry-based metabolomics to elucidate functions in marine organisms and ecosystems. Mar. Drugs 2012, 10, 849–880. [Google Scholar] [CrossRef] [PubMed]
- Peng, B.; Li, H.; Peng, X.X. Functional metabolomics: From biomarker discovery to metabolome reprogramming. Protein Cell 2015, 6, 628–637. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Kate, B.N.; Banerjee, U. Bioactive compounds from cyanobacteria and microalgae: An overview. Crit. Rev. Biotechnol. 2005, 25, 73–95. [Google Scholar] [CrossRef] [PubMed]
- Bundy, J.G.; Davey, M.P.; Viant, M.R. Environmental metabolomics: A critical review and future perspectives. Metabolomics 2009, 5, 3. [Google Scholar] [CrossRef]
- Viant, M.R. Recent developments in environmental metabolomics. Mol. BioSyst. 2008, 4, 980–986. [Google Scholar] [CrossRef] [PubMed]
- Huseby, S.; Degerlund, M.; Zingone, A.; Hansen, E. Metabolic fingerprinting reveals differences between northern and southern strains of the cryptic diatom Chaetoceros socialis. Eur. J. Phycol. 2012, 47, 480–489. [Google Scholar] [CrossRef]
- Cachet, N.; Genta-Jouve, G.; Ivanisevic, J.; Chevaldonné, P.; Sinniger, F.; Culioli, G.; Pérez, T.; Thomas, O.P. Metabolomic profiling reveals deep chemical divergence between two morphotypes of the zoanthid Parazoanthus axinellae. Sci. Rep. 2015, 5, 8282. [Google Scholar] [CrossRef] [PubMed]
- Tachibana, K.; Scheuer, P.J.; Tsukitani, Y.; Kikuchi, H.; Van Engen, D.; Clardy, J.; Gopichand, Y.; Schmitz, F.J. Okadaic acid, a cytotoxic polyether from two marine sponges of the genus Halichondria. J. Am. Chem. Soc. 1981, 103, 2469–2471. [Google Scholar] [CrossRef]
- Murakami, Y.; Oshima, Y.; Yasumoto, T. Identification of okadaic acid as a toxic component of a marine dinoflagellate Prorocentrum lima. Bull. Jpn. Soc. Sci. Fish. 1982, 48, 69–72. [Google Scholar] [CrossRef]
- Nagai, H.; Torigoe, K.; Satake, M.; Murata, M.; Yasumoto, T.; Hirota, H. Gambieric acids: Unprecedented potent antifungal substances isolated from cultures of a marine dinoflagellate Gambierdiscus toxicus. J. Am. Chem. Soc. 1992, 114, 1102–1103. [Google Scholar] [CrossRef]
- Sogawa, K.; Matsuda, M.; Okutani, K. Induction of apoptosis by a marine microalgal polysaccharide in a human leukemic cell line. J. Mar. Biotechnol. 1998, 6, 241–243. [Google Scholar] [PubMed]
- Umemura, K.; Yanase, K.; Suzuki, M.; Okutani, K.; Yamori, T.; Andoh, T. Inhibition of DNA topoisomerases I and II, and growth inhibition of human cancer cell lines by a marine microalgal polysaccharide. Biochem. Pharmacol. 2003, 66, 481–487. [Google Scholar] [CrossRef]
- Kobayashi, J.; Tsuda, M. Bioactive products from Okinawan marine micro-and macroorganisms. Phytochem. Rev. 2004, 3, 267–274. [Google Scholar] [CrossRef]
- Kerr, R.G.; Kerr, S.S. Marine natural products as therapeutic agents. Expert. Opin. Ther. Pat. 1999, 9, 1207–1222. [Google Scholar] [CrossRef]
- Blunt, J.W.; Munro, M.H. Dictionary of Marine Natural Products with CD-ROM, 1st ed.; Chapman and Hall/CRC: Boca Raton, FL, USA, 2007; pp. 1–2536. ISBN 978-0-8493-8216-1. [Google Scholar]
- Smith, C.A.; O’Maille, G.; Want, E.J.; Qin, C.; Trauger, S.A.; Brandon, T.R.; Custodio, D.E.; Abagyan, R.; Siuzdak, G. METLIN: A metabolite mass spectral database. Ther. Drug. Monit. 2005, 27, 747–751. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Cuetos, L.; Moestrup, Ø.; Hansen, P.J. Studies on the genus Mesodinium II. Ultrastructural and molecular investigations of five marine species help clarifying the taxonomy. J. Eukaryot. Microbiol. 2012, 59, 374–400. [Google Scholar] [CrossRef] [PubMed]
- Vale, P.; Sampayo, M.A.D.M. Esterification of DSP toxins by Portuguese bivalves from the Northwest coast determined by LC-MS—A widespread phenomenon. Toxicon 2002, 40, 33–42. [Google Scholar] [CrossRef]
- Sibat, M.; García-Portela, M.; Hess, P. First identification of a C9-diol-ester of okadaic acid in Dinophysis acuta from Galician Rías Baixas (NW Spain). Toxicon. (under review).
- Hiraga, Y.; Kaku, K.; Omoda, D.; Sugihara, K.; Hosoya, H.; Hino, M. A new digalactosyl diacylglycerol from a cultured marine dinoflagellate Heterocapsa circularisquama. J. Nat. Prod. 2002, 65, 1494–1496. [Google Scholar] [CrossRef] [PubMed]
- Boudière, L.; Michaud, M.; Petroutsos, D.; Rébeillé, F.; Falconet, D.; Bastien, O.; Roy, S.; Finazzi, G.; Rolland, N.; Jouhet, J. Glycerolipids in photosynthesis: Composition, synthesis and trafficking. Biochim. Biophys. Acta Bioenerg. 2014, 1837, 470–480. [Google Scholar] [CrossRef] [PubMed]
- LeBlond, J.D.; Menn, F.M.; Schultz, T.W.; Sayler, G.S. Structure-toxicity assessment of metabolites of the aerobic bacterial transformation of substituted naphthalenes. Environ. Toxicol. Chem. 2000, 19, 1235–1246. [Google Scholar]
- Joyard, J.; Teyssier, E.; Miège, C.; Berny-Seigneurin, D.; Maréchal, E.; Block, M.A.; Dorne, A.-J.; Rolland, N.; Ajlani, G.; Douce, R. The biochemical machinery of plastid envelope membranes. Plant. Physiol. 1998, 118, 715–723. [Google Scholar] [CrossRef] [PubMed]
- Sakurai, I.; Hagio, M.; Gombos, Z.; Tyystjärvi, T.; Paakkarinen, V.; Aro, E.-M.; Wada, H. Requirement of phosphatidylglycerol for maintenance of photosynthetic machinery. Plant. Physiol. 2003, 133, 1376–1384. [Google Scholar] [CrossRef] [PubMed]
- Härtel, H.; Dörmann, P.; Benning, C. DGD1-independent biosynthesis of extraplastidic galactolipids after phosphate deprivation in Arabidopsis. Proc. Natl. Acad. Sci. USA 2000, 97, 10649–10654. [Google Scholar] [CrossRef] [PubMed]
- Van Mooy, B.A.; Fredricks, H.F.; Pedler, B.E.; Dyhrman, S.T.; Karl, D.M.; Koblížek, M.; Lomas, M.W.; Mincer, T.J.; Moore, L.R.; Moutin, T. Phytoplankton in the ocean use non-phosphorus lipids in response to phosphorus scarcity. Nature 2009, 458, 69–72. [Google Scholar] [CrossRef] [PubMed]
- Lasek-Nesselquist, E.; Wisecaver, J.H.; Hackett, J.D.; Johnson, M.D. Insights into transcriptional changes that accompany organelle sequestration from the stolen nucleus of Mesodinium rubrum. BMC Genom. 2015, 16, 805. [Google Scholar] [CrossRef] [PubMed]
- Dawaliby, R.; Trubbia, C.; Delporte, C.; Noyon, C.; Ruysschaert, J.-M.; Van Antwerpen, P.; Govaerts, C. Phosphatidylethanolamine is a key regulator of membrane fluidity in eukaryotic cells. J. Biol. Chem. 2016, 291, 3658–3667. [Google Scholar] [CrossRef] [PubMed]
- Hakomori, S. Structure, organization, and function of glycosphingolipids in membrane. Curr. Opin. Hematol. 2003, 10, 16–24. [Google Scholar] [CrossRef] [PubMed]
- Gong, W.; Browne, J.; Hall, N.; Schruth, D.; Paerl, H.; Marchetti, A. Molecular insights into a dinoflagellate bloom. ISME J. 2016, 11, 439. [Google Scholar] [CrossRef] [PubMed]
- Mansour, M.P.; Volkman, J.K.; Blackburn, S.I. The effect of growth phase on the lipid class, fatty acid and sterol composition in the marine dinoflagellate, Gymnodinium sp. in batch culture. Phytochemistry 2003, 63, 145–153. [Google Scholar] [CrossRef]
- Fujita, Y. Protochlorophyllide reduction: A key step in the greening of plants. Plant. Cell. Physiol. 1996, 37, 411–421. [Google Scholar] [CrossRef] [PubMed]
- Snyder, R.; Gibbs, P.; Palacios, A.; Abiy, L.; Dickey, R.; Lopez, J.V.; Rein, K. Polyketide synthase genes from marine dinoflagellates. Mar. Biotechnol. 2003, 5, 1–12. [Google Scholar] [PubMed]
- Snyder, R.V.; Guerrero, M.A.; Sinigalliano, C.D.; Winshell, J.; Perez, R.; Lopez, J.V.; Rein, K.S. Localization of polyketide synthase encoding genes to the toxic dinoflagellate Karenia brevis. Phytochemistry 2005, 66, 1767–1780. [Google Scholar] [CrossRef] [PubMed]
- Yu, S.; Deng, Z.; van Ofwegen, L.; Proksch, P.; Lin, W. 5,8-Epidioxysterols and related derivatives from a Chinese Soft Coral Sinularia flexibilis. Steroids 2006, 71, 955–959. [Google Scholar] [CrossRef] [PubMed]
- Imbs, A.B.; Latyshev, N.A. Fatty acid composition as an indicator of possible sources of nutrition for soft corals of the genus Sinularia (Alcyoniidae). J. Mar. Biol. Assoc. UK 2012, 92, 1341–1347. [Google Scholar] [CrossRef]
- Shimizu, Y.; Chou, H.N.; Bando, H.; Van Duyne, G.; Clardy, J. Structure of brevetoxin A (GB-1 toxin), the most potent toxin in the Florida red tide organism Gymnodinium breve (Ptychodiscus brevis). J. Am. Chem. Soc. 1986, 108, 514–515. [Google Scholar] [CrossRef] [PubMed]
- Larsen, K.; Petersen, D.; Wilkins, A.L.; Samdal, I.A.; Sandvik, M.; Rundberget, T.; Goldstone, D.; Arcus, V.; Hovgaard, P.; Rise, F. Clarification of the C-35 stereochemistries of dinophysistoxin-1 and dinophysistoxin-2 and its consequences for binding to protein phosphatase. Chem. Res. Toxicol. 2007, 20, 868–875. [Google Scholar] [CrossRef] [PubMed]
- Lu, C.-K.; Chou, H.N.; Lee, C.K.; Lee, T.H. Prorocentin, a new polyketide from the marine dinoflagellate Prorocentrum lima. Org. Lett. 2005, 7, 3893–3896. [Google Scholar]
- Chill, L.; Berrer, N.; Benayahu, Y.; Kashman, Y. Eunicellin diterpenes from two Kenyan soft corals. J. Nat. Prod. 2005, 68, 19–25. [Google Scholar] [CrossRef] [PubMed]
- Ovenden, S.P.; Capon, R.J. Nuapapuin A and sigmosceptrellins D and E: New norterpene cyclic peroxides from a southern Australian marine sponge, Sigmosceptrella sp. J. Nat. Prod. 1999, 62, 214–218. [Google Scholar] [CrossRef] [PubMed]
- Cardellina, J.H.; Marner, F.J.; Moore, R.E. Malyngamide A, a novel chlorinated metabolite of the marine cyanophyte Lyngbya majuscula. J. Am. Chem. Soc. 1979, 101, 240–242. [Google Scholar] [CrossRef]
- Hansen, F.C.; Witte, H.J.; Passarge, J. Grazing in the heterotrophic dinoflagellate Oxyrrhis marina: Size selectivity and preference for calcified Emiliania huxleyi cells. Aquat. Microb. Ecol. 1996, 10, 307–313. [Google Scholar] [CrossRef]
- Naustvoll, L.J. Prey size spectra and food preferences in thecate heterotrophic dinoflagellates. Phycologia 2000, 39, 187–198. [Google Scholar] [CrossRef]
- Tong, M.; Zhou, Q.; David, K.M.; Jiang, T.; Qi, Y.; Donald, A.M. Culture techniques and growth characteristics of Dinophysis acuminata and its prey. Chin. J. Oceanol. Limnol. 2010, 28, 1230–1239. [Google Scholar] [CrossRef]
- Nagai, S.; Suzuki, T.; Nishikawa, T.; Kamiyama, T. Differences in the production and excretion kinetics of okadaic acid, dinophysistoxin-1, and pectenotoxin-2 between cultures of Dinophysis acuminata and Dinophysis fortii isolated from Western Japan. J. Phycol. 2011, 47, 1326–1337. [Google Scholar] [CrossRef] [PubMed]
- Pizarro, G.; Escalera, L.; González-Gil, S.; Franco, J.; Reguera, B. Growth, behaviour and cell toxin quota of Dinophysis acuta during a daily cycle. Mar. Ecol. Prog. Ser. 2008, 353, 89–105. [Google Scholar] [CrossRef]
- Fernández, M.L.; Reguera, B.; González-Gil, S.; Míguez, A. Pectenotoxin-2 in single-cell isolates of Dinophysis caudata and Dinophysis acuta from the Galician Rías (NW Spain). Toxicon 2006, 48, 477–490. [Google Scholar] [CrossRef] [PubMed]
- Martins, C.A.; Kulis, D.; Franca, S.; Anderson, D.M. The loss of PSP toxin production in a formerly toxic Alexandrium lusitanicum clone. Toxicon 2004, 43, 195–205. [Google Scholar] [CrossRef] [PubMed]
- Fiehn, O. Metabolomics—The link between genotypes and phenotypes. In Functional Genomics; Springer: Berlin/Heidelberg, Germany, 2002; pp. 155–171. [Google Scholar]
- Sardans, J.; Penuelas, J.; Rivas-Ubach, A. Ecological metabolomics: Overview of current developments and future challenges. Chemoecology 2011, 21, 191–225. [Google Scholar] [CrossRef]
- Nielsen, J.; Jewett, M.C. The role of metabolomics in systems biology. In Metabolomics; Springer: Berlin/Heidelberg, Germany, 2007; pp. 1–10. [Google Scholar]
- Legrand, C.; Rengefors, K.; Fistarol, G.O.; Graneli, E. Allelopathy in phytoplankton-biochemical, ecological and evolutionary aspects. Phycologia 2003, 42, 406–419. [Google Scholar] [CrossRef]
- Arzul, G.; Gentien, P. Allelopathic interactions among marine microalgae. In Algal Cultures, Analogues of Blooms and Applications; Subba Rao, D.V., Ed.; Science Publishers: Enfield, NH, USA, 2006; pp. 131–161. [Google Scholar]
- Thacker, R.W.; Nagle, D.G.; Paul, V.J. Effects of repeated exposures to marine cyanobacterial secondary metabolites on feeding by juvenile rabbitfish and parrotfish. Mar. Ecol. Prog. Ser. 1997, 21–29. [Google Scholar] [CrossRef]
- Waggett, R.J.; Tester, P.A.; Place, A.R. Anti-grazing properties of the toxic dinoflagellate Karlodinium veneficum during predator–prey interactions with the copepod Acartia tonsa. Mar. Ecol. Prog. Ser. 2008, 366, 31–42. [Google Scholar] [CrossRef]
- Wood-Charlson, E.M.; Hollingsworth, L.L.; Krupp, D.A.; Weis, V.M. Lectin/glycan interactions play a role in recognition in a coral/dinoflagellate symbiosis. Cell. Microbiol. 2006, 8, 1985–1993. [Google Scholar] [CrossRef] [PubMed]
- Mafra, L.L., Jr.; Nagai, S.; Uchida, H.; Tavares, C.P.; Escobar, B.P.; Suzuki, T. Harmful effects of Dinophysis to the ciliate Mesodinium rubrum: Implications for prey capture. Harmful Algae 2016, 59, 82–90. [Google Scholar] [CrossRef] [PubMed]
- Ojamäe, K.; Hansen, P.J.; Lips, I. Mass entrapment and lysis of Mesodinium rubrum cells in mucus threads observed in cultures with Dinophysis. Harmful Algae 2016, 55, 77–84. [Google Scholar] [CrossRef] [PubMed]
- Papiol, G.G.; Beuzenberg, V.; Selwood, A.I.; MacKenzie, L.; Packer, M.A. The use of a mucus trap by Dinophysis acuta for the capture of Mesodinium rubrum prey under culture conditions. Harmful Algae 2016, 58, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Richlen, M.; Barber, P. A technique for the rapid extraction of microalgal DNA from single live and preserved cells. Mol. Ecol. Resour. 2005, 5, 688–691. [Google Scholar] [CrossRef]
- Guindon, S.; Dufayard, J.-F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of PhyML 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Lefort, V.; Longueville, J.-E.; Gascuel, O. SMS: Smart Model Selection in PhyML. Mol. Biol. Evol. 2017, 34, 2422–2424. [Google Scholar] [CrossRef] [PubMed]
- Huelsenbeck, J.P.; Ronquist, F. MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 2001, 17, 754–755. [Google Scholar] [CrossRef] [PubMed]
- Bowers, H.A.; Tomas, C.; Tengs, T.; Kempton, J.W.; Lewitus, A.J.; Oldach, D.W. Raphidophyceae (chadefaud ex silva) systematics and rapid identification: Sequence analyses and real-time PCR assays. J. Phycol. 2006, 42, 1333–1348. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.H.; Ivanisevic, J.; Siuzdak, G. Metabolomics: Beyond biomarkers and towards mechanisms. Nat. Rev. Mol. Cell Biol. 2016, 17, 451–459. [Google Scholar] [CrossRef] [PubMed]
- Johnson, M.D.; Tengs, T.; Oldach, D.W.; Delwiche, C.F.; Stoecker, D.K. Highly divergent SSU rRNA genes found in the marine ciliates Myrionecta rubra and Mesodinium pulex. Protist 2004, 155, 347–359. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protoc. Guide Methods Appl. 1990, 18, 315–322. [Google Scholar]
- Scholin, C.A.; Herzog, M.; Sogin, M.; Anderson, D.M. Identification of group-and strain-specific genetic markers for globally distributed Alexandrium (Dinophyceae). II. Sequence analysis of a fragment of the LSU rRNA gene. J. Phycol. 1994, 30, 999–1011. [Google Scholar] [CrossRef]
- Olenina, I. Biovolumes and size-classes of phytoplankton in the Baltic Sea. HELCOM Balt. Sea Environ. Proc. 2006, 106, 1–144. [Google Scholar]
- Guillard, R.; Hargraves, P. Stichochrysis immobilis is a diatom, not a chrysophyte. Phycologia 1993, 32, 234–236. [Google Scholar] [CrossRef]
- Mountfort, D.O.; Suzuki, T.; Truman, P. Protein phosphatase inhibition assay adapted for determination of total DSP in contaminated mussels. Toxicon 2001, 39, 383–390. [Google Scholar] [CrossRef]
| Sample | Day | Cells (N) ± Std Dev. | Pellet Biomass % Average | Volume (mL) | ||
|---|---|---|---|---|---|---|
| Dinophysis | M. rubrum | T. amphioxeia | ||||
| T. amphioxeia | 3 | 283,333 ± 28,867 | 100 | 10 | ||
| Danish M. rubrum | 7 | 89,500 ± 866 | 100 | 15 | ||
| Spanish M. rubrum | 7 | 80,000 ± 8660 | 100 | 30 | ||
| D. acuta + Danish M. rubrum | 7 | 40,140 ± 1247 | 93.48 | 6.52 | 90 | |
| D. acuta + Spanish M. rubrum | 7 | 29,943 ± 3264 | 97.26 | 2.74 | 130 | |
| D. acuminata + Danish M. rubrum | 7 | 115,470 ± 9490 | 99.40 | 0.60 | 90 | |
| D. acuminata + Spanish M. rubrum | 7 | 58,413 ± 3134 | 86.62 | 10.38 | 130 | |
| D. acuta + Danish M. rubrum | 15 | 21,967 ± 2627 | 81.62 | 18.38 | 100 | |
| D. acuta + Spanish M. rubrum | 15 | 42,733 ± 3225 | 100 | 100 | ||
| D. acuminata + Danish M. rubrum | 15 | 225,800 ± 9616 | 100 | 100 | ||
| D. acuminata + Spanish M. rubrum | 15 | 58,900 ± 265 | 100 | 100 | ||
| Sp | PO | GP/NS | Toxin Contents pb = per Biomass (fg µm−3), pc = per Cell (pg cell−1) ± Standard Deviation | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Free OA | Total OA | Free DTX2 | Total DTX2 | PTX2 | ||||||||
| pb | pc | pb | pc | pb | pc | pb | pc | pb | pc | |||
| D. acuta | ES | ME | 0.17 ± 0.03 | 9.2 ± 2.0 | 0.22 ± 0.03 | 12.2 ± 2.3 | 0.07 ± 0.01 | 3.8 ± 0.6 | 0.08 ± 0.01 | 4.4 ± 0.9 | 0.41 ± 0.14 | 22.2 ± 9.4 |
| D. acuminata | ES | ME | 1.05 ± 0.34 | 21.5 ± 0.5 | 2.1 ± 0.3 | 35.2 ± 6.8 | ||||||
| D. acuta | ES | Well-Fed | 0.55 ± 0.06 | 30.0 ± 3.7 | 0.75 ± 0.07 | 41.0 ± 4.9 | 0.24 ± 0.03 | 13.5 ± 1.7 | 0.32 ± 0.06 | 17.4 ± 4.1 | 0.69 ± 0.12 | 38.0 ± 8.2 |
| Prey-limited | 1.49 ± 0.24 | 76.7 ± 18.6 | 1.36 ± 0.11 | 74.1 ± 8.2 | 0.56 ± 0.11 | 30.2 ± 8.5 | 0.60 ± 0.05 | 32.4 ± 3.8 | 1.29 ± 0.15 | 59.3 ± 11.8 | ||
| DK | Well-Fed | 0.53 ± 0.12 | 28.7 ± 8.1 | 0.66 ± 0.11 | 35.9 ± 7.07 | 0.26 ± 0.03 | 14.2 ± 2.2 | 0.30 ± 0.01 | 16.5 ± 0.8 | 0.78 ± 0.01 | 70.0 ± 0.8 | |
| Prey-limited | 0.64 ± 0.08 | 34.6 ± 5.4 | 0.71 ± 0.07 | 38.6 ± 4.5 | 0.32 ± 0.04 | 17.4 ± 2.7 | 0.35 ± 0.03 | 19.0 ± 2.3 | 0.8 ± 0.09 | 43.6 ± 6.2 | ||
| D. acuminata | ES | Well-Fed | 0.37 ± 0.14 | 6.0 ± 3.0 | 0.36 ± 0.13 | 6.0 ± 2.8 | ||||||
| Prey-limited | 1.02 ± 0.20 | 17.1 ± 4.6 | 1.28 ± 0.03 | 21.5 ± 0.7 | ||||||||
| DK | Well-Fed | 0.49 ± 0.09 | 8.3 ± 1.5 | 0.59 ± 0.08 | 9.8 ± 1.7 | |||||||
| Prey-limited | 1.44 ± 0.32 | 24.0 ± 6.5 | 1.93 ± 0.23 | 32.3 ± 4.7 | ||||||||
| Name | Sequence 5′–3′ | Reference |
|---|---|---|
| 4617F | TCCTGCCAGTAGTCATATGC | [93] |
| Meso580R | GACGTACAGACTACGGACG | [94] |
| Meso245F | CGACTCGACGTCCCG | [94] |
| UNIDEUK1416R | GTTTCAGACTTGTGTCCATACTA | [95] |
| Meso580F | CGTCCGTAGTCTGTACGTC | [94] |
| Meso1480R | CTAAACACTCGATCGGTAGG | [94] |
| Meso1200F | ATTCCGGTAACGAACGAGAC | [94] |
| Meso28S_R | AGACTTGGATGACTTTTATCACC | [94] |
| ITS1 | TCCGTAGGTGAACCTGCGG | [96] |
| Dir-2CR | CCTTGGTCCGTGTTTCAAGA | [97] |
| Toxin | Ionization Mode | Ion | Parent Ion (m/z) | Fragment Ions (m/z) | DP (V) | CE (eV) |
|---|---|---|---|---|---|---|
| OA | ESI− | [M − H]− | 803.4 | 255.1 * | −170 | −62 |
| 113.1 | −92 | |||||
| DTX2 | ESI− | [M − H]− | 803.4 | 255.1 * | −170 | −62 |
| 113.1 | −92 | |||||
| DTX1 | ESI− | [M − H]− | 817.5 | 255.1 * | −170 | −68 |
| 113.1 | −92 | |||||
| PTX2 | ESI+ | [M + NH4]+ | 876.6 | 823.6 * | 91 | 31 |
| 805.6 | 37 | |||||
| 213.6 | 55 | |||||
| PTX2sa and 7-epi PTX2sa | ESI+ | [M + NH4]+ | 894.6 | 823.6 * | 91 | 31 |
| 805.6 | 37 | |||||
| 213.6 | 55 | |||||
| PTX1, PTX4 and PTX11 | ESI+ | [M + NH4]+ | 892.6 | 839.6 * | 91 | 31 |
| 821.4 | 37 | |||||
| 213.6 | 55 | |||||
| PTX3 | ESI+ | [M + NH4]+ | 890.5 | 873.6 * | 91 | 31 |
| 856.6 | 37 | |||||
| 213.6 | 55 | |||||
| PTX6 and PTX7 | ESI+ | [M + NH4]+ | 906.6 | 871.6 * | 91 | 31 |
| 853.6 | 37 | |||||
| 213.6 | 55 | |||||
| PTX12 and PTX14 | ESI+ | [M + NH4]+ | 874.6 | 857.6 * | 91 | 31 |
| 840.6 | 37 | |||||
| 213.6 | 55 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
García-Portela, M.; Reguera, B.; Sibat, M.; Altenburger, A.; Rodríguez, F.; Hess, P. Metabolomic Profiles of Dinophysis acuminata and Dinophysis acuta Using Non-Targeted High-Resolution Mass Spectrometry: Effect of Nutritional Status and Prey. Mar. Drugs 2018, 16, 143. https://doi.org/10.3390/md16050143
García-Portela M, Reguera B, Sibat M, Altenburger A, Rodríguez F, Hess P. Metabolomic Profiles of Dinophysis acuminata and Dinophysis acuta Using Non-Targeted High-Resolution Mass Spectrometry: Effect of Nutritional Status and Prey. Marine Drugs. 2018; 16(5):143. https://doi.org/10.3390/md16050143
Chicago/Turabian StyleGarcía-Portela, María, Beatriz Reguera, Manoella Sibat, Andreas Altenburger, Francisco Rodríguez, and Philipp Hess. 2018. "Metabolomic Profiles of Dinophysis acuminata and Dinophysis acuta Using Non-Targeted High-Resolution Mass Spectrometry: Effect of Nutritional Status and Prey" Marine Drugs 16, no. 5: 143. https://doi.org/10.3390/md16050143







