Ovothiol Isolated from Sea Urchin Oocytes Induces Autophagy in the Hep-G2 Cell Line
Abstract
:1. Introduction
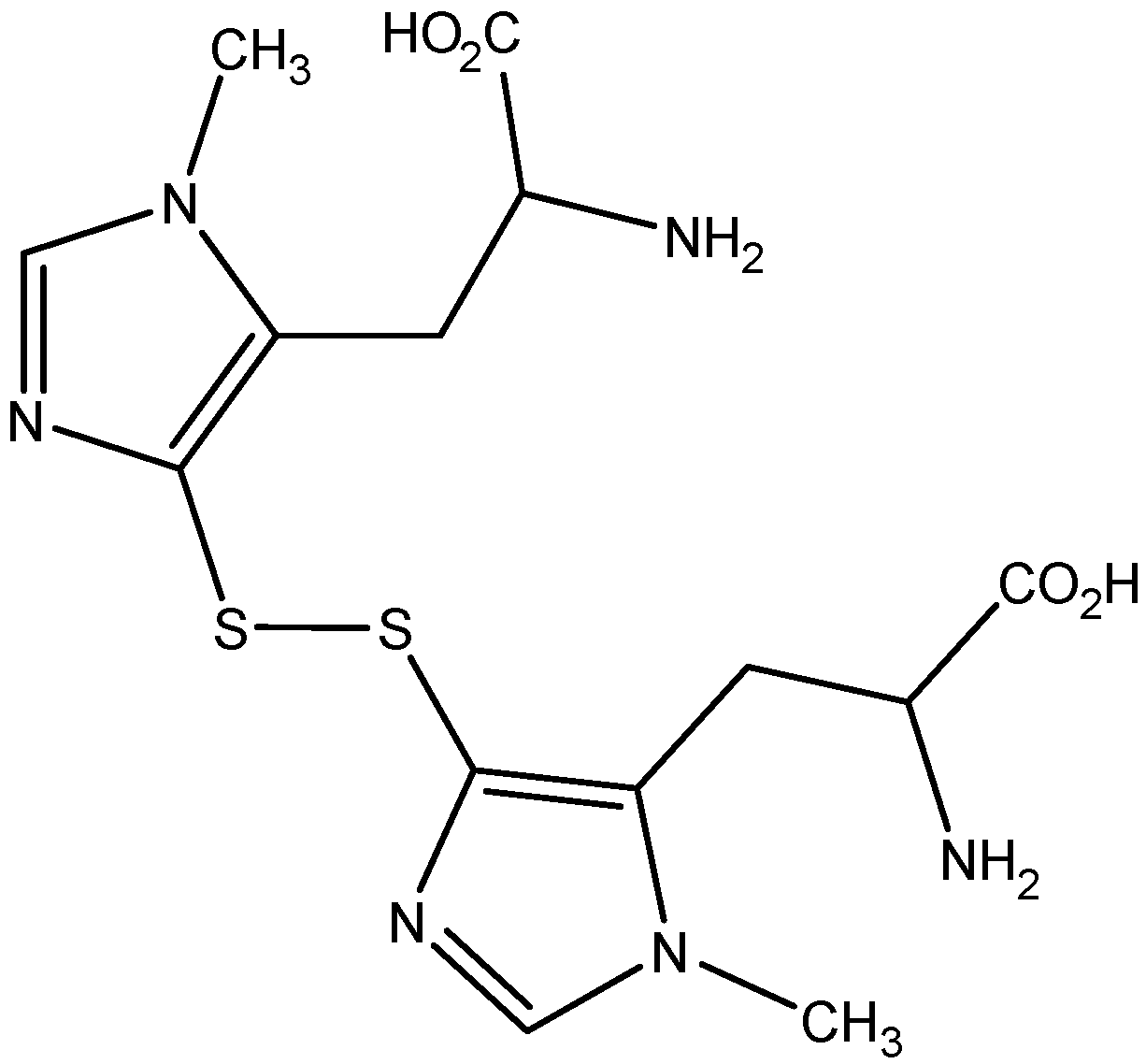
2. Results
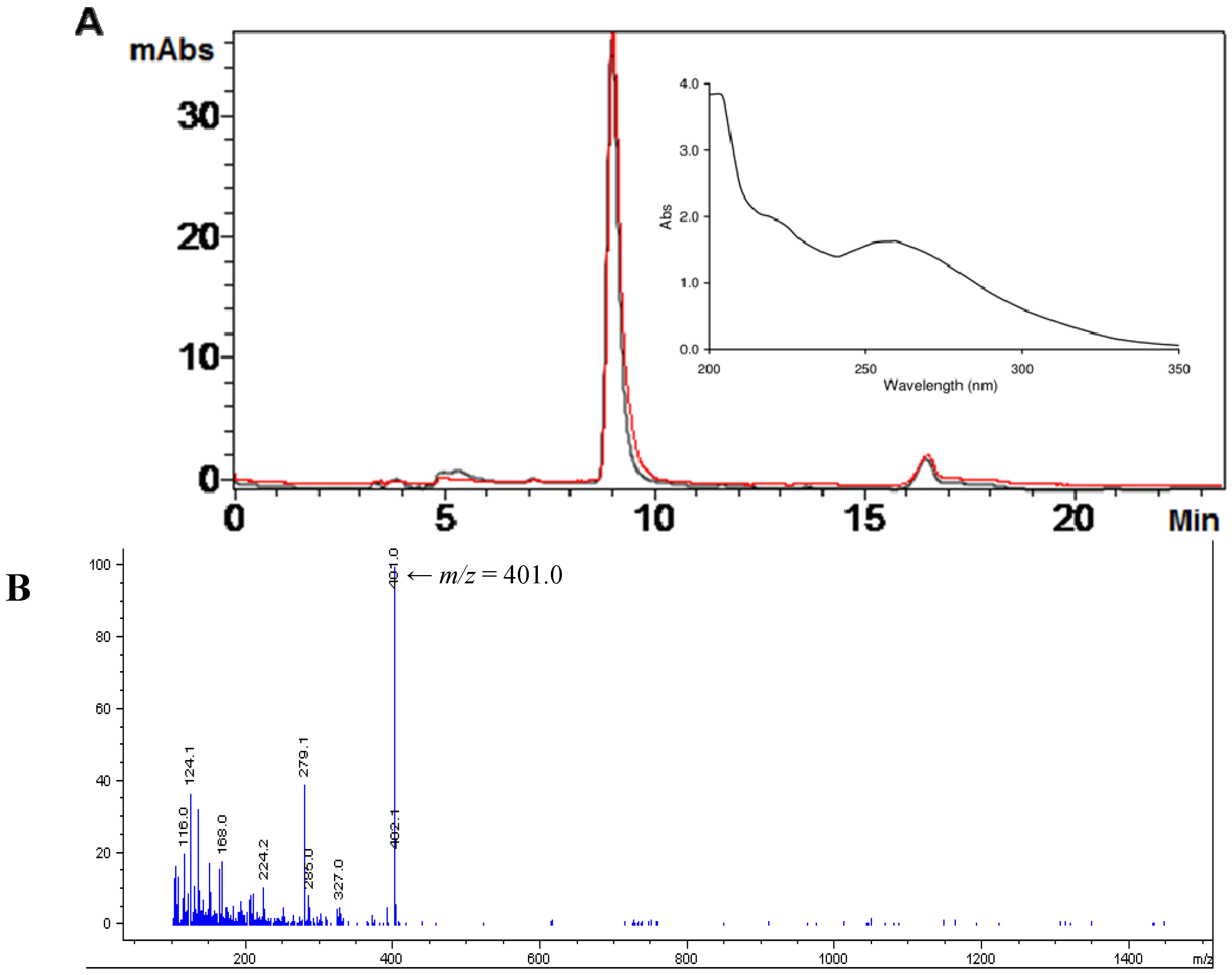
2.1. Isolation of Ovothiol A
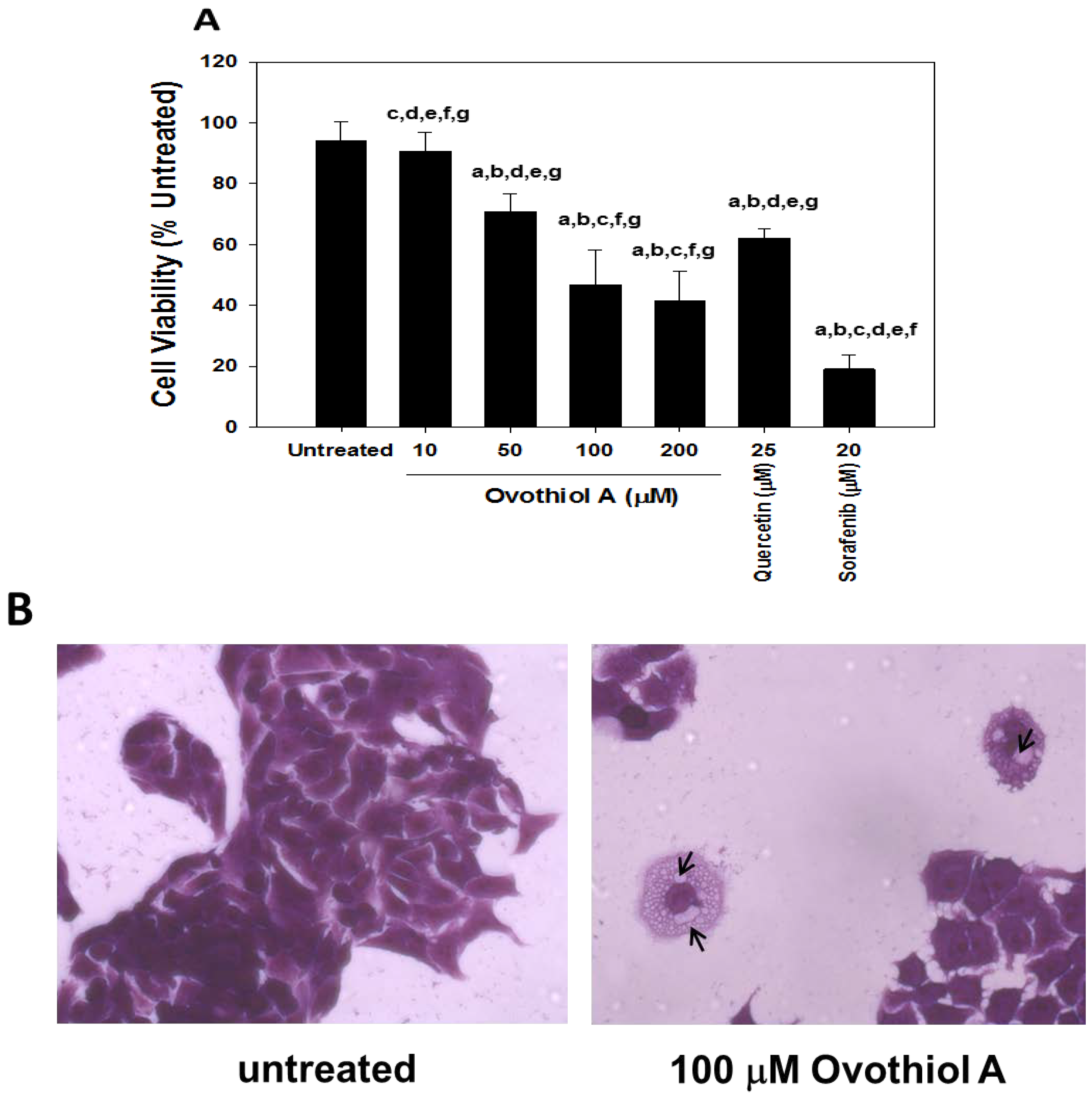
2.2. Anti-Proliferative Effects of Ovothiol A in the Hep-G2 Cell Line
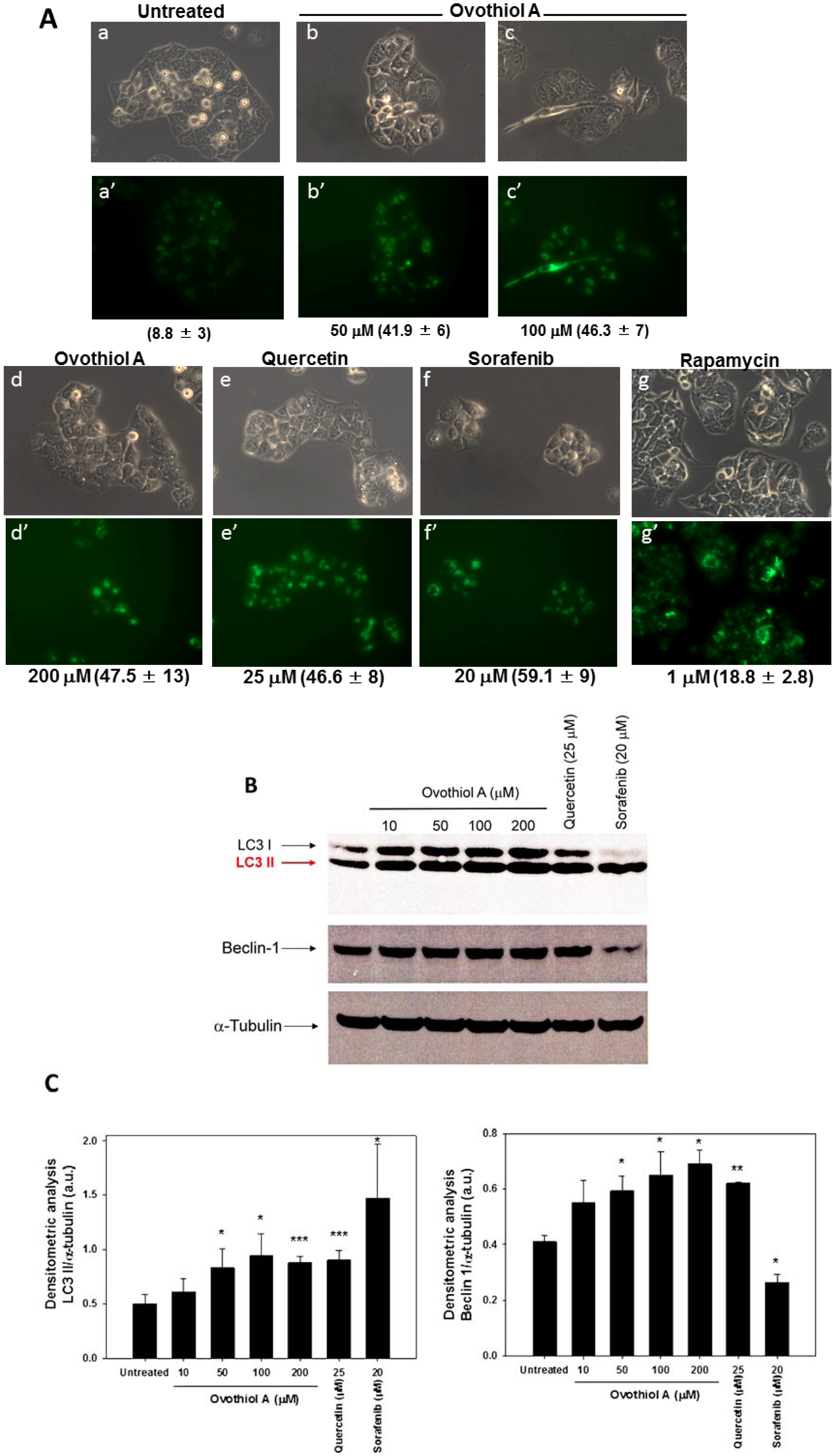
2.3. Sea Urchin Ovothiol A Activates Autophagic Processes in the Hep-G2 Cell Line
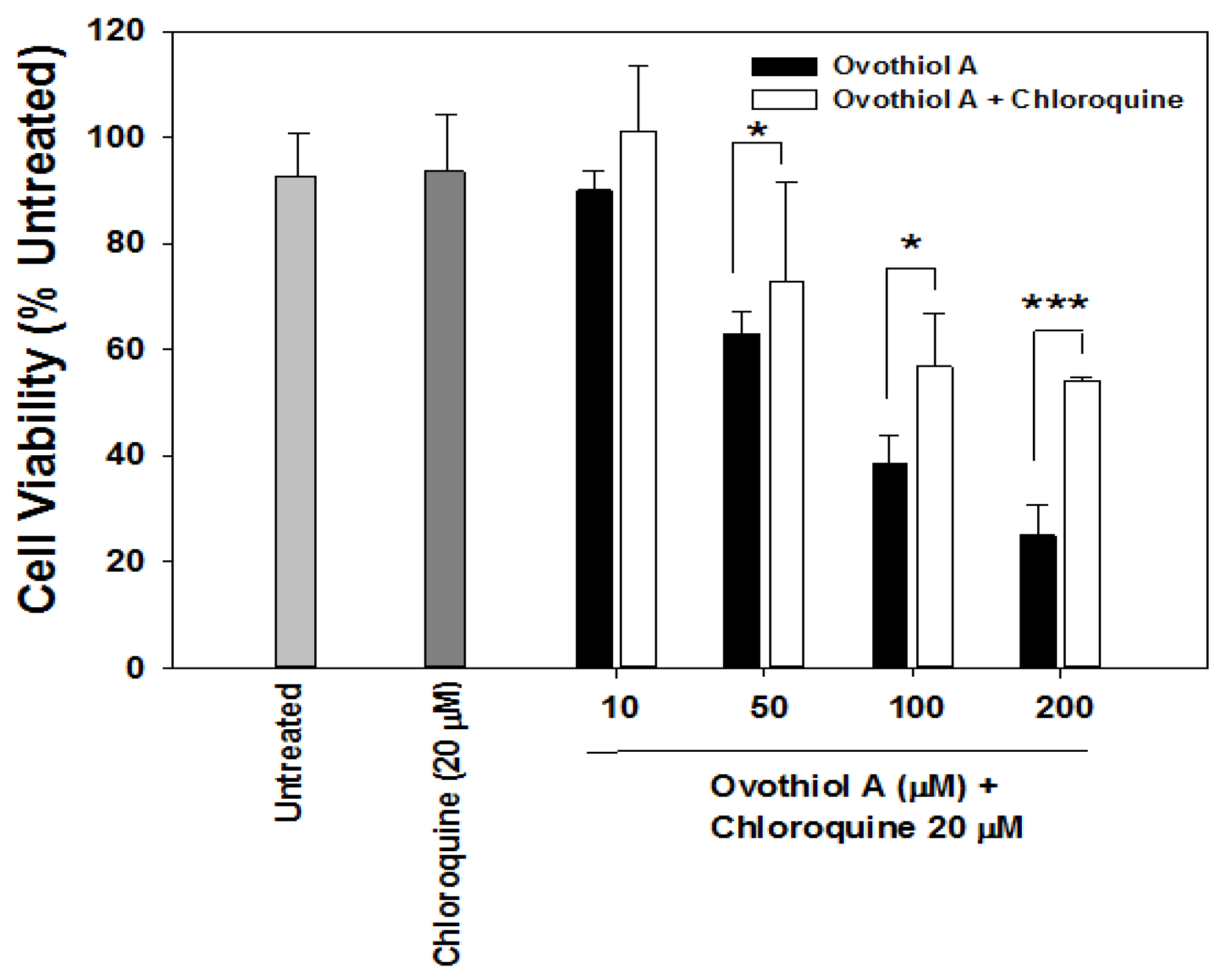
2.4. Ovothiol A Recovery in the Culture Media of the Hep-G2 Cell Line
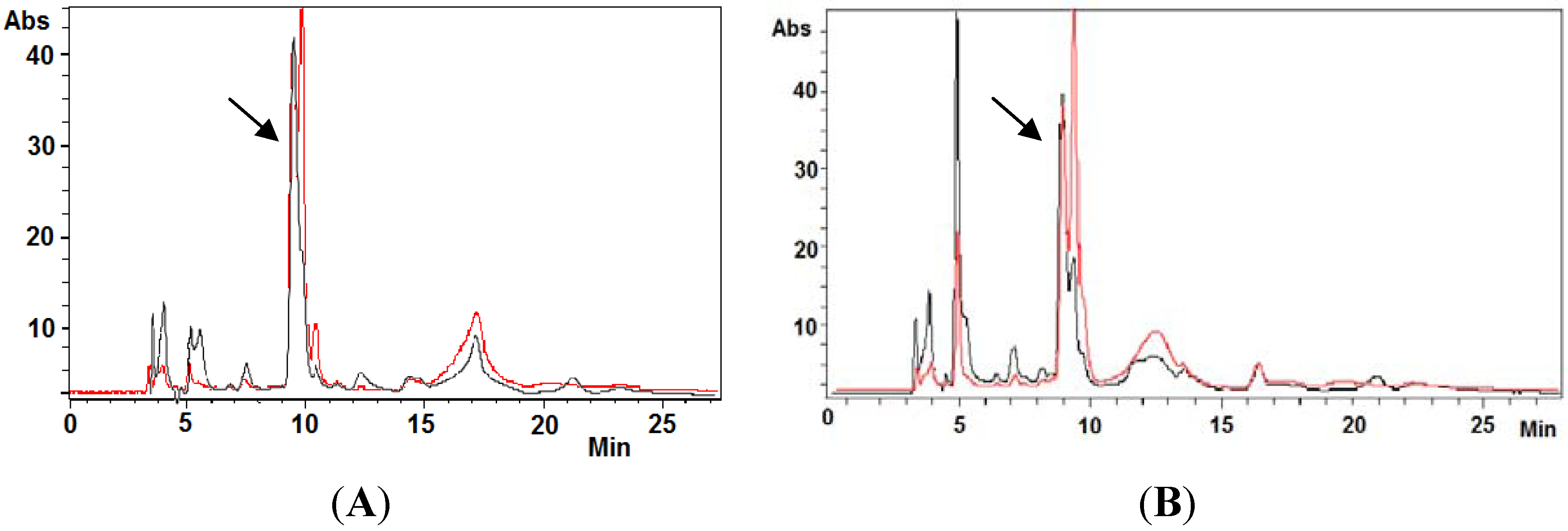
2.5. ROS Production in the Hep-G2 Cell Line
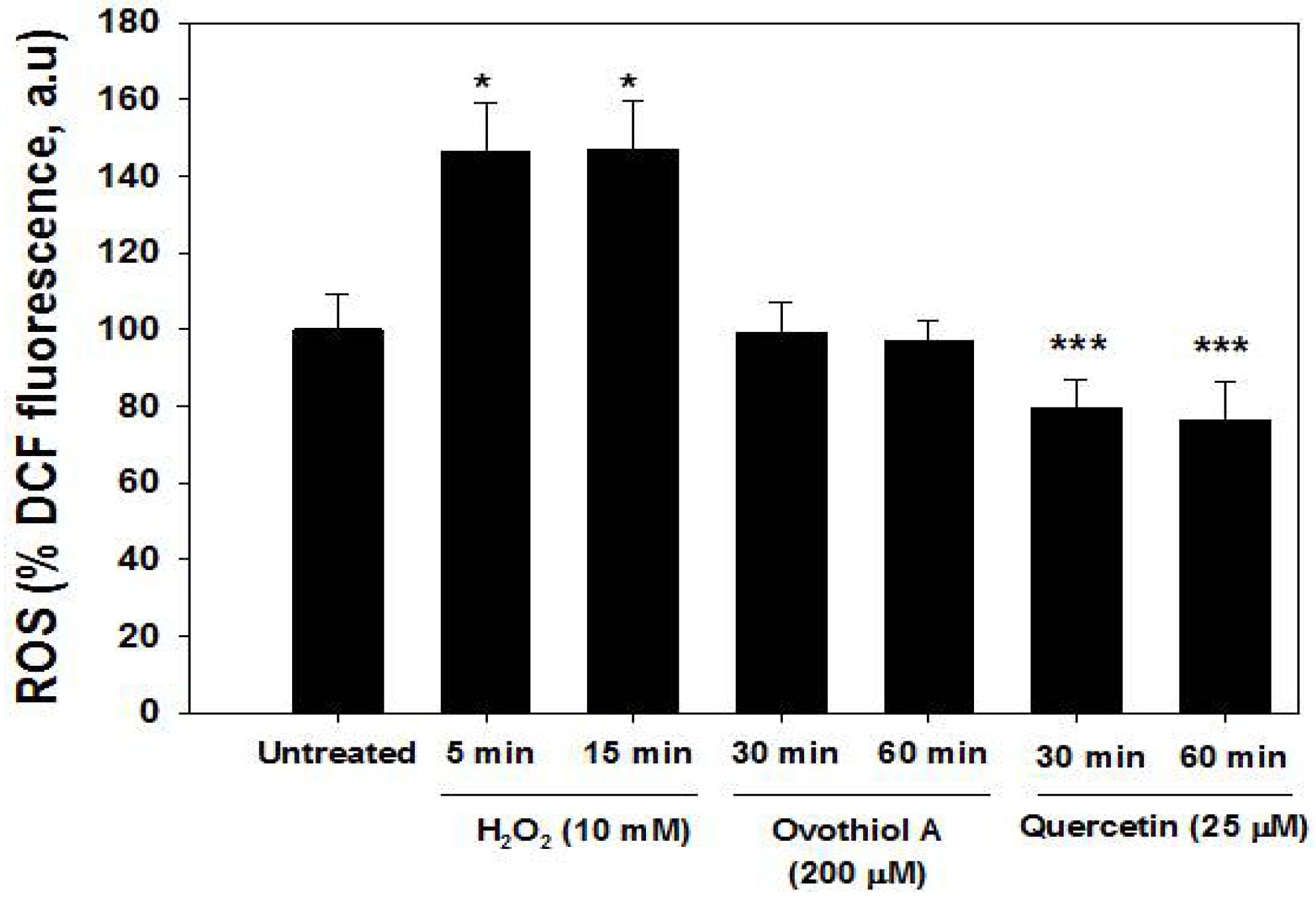
2.6. Ovothiol A Is Not Cytotoxic on Immortalized Fibroblast
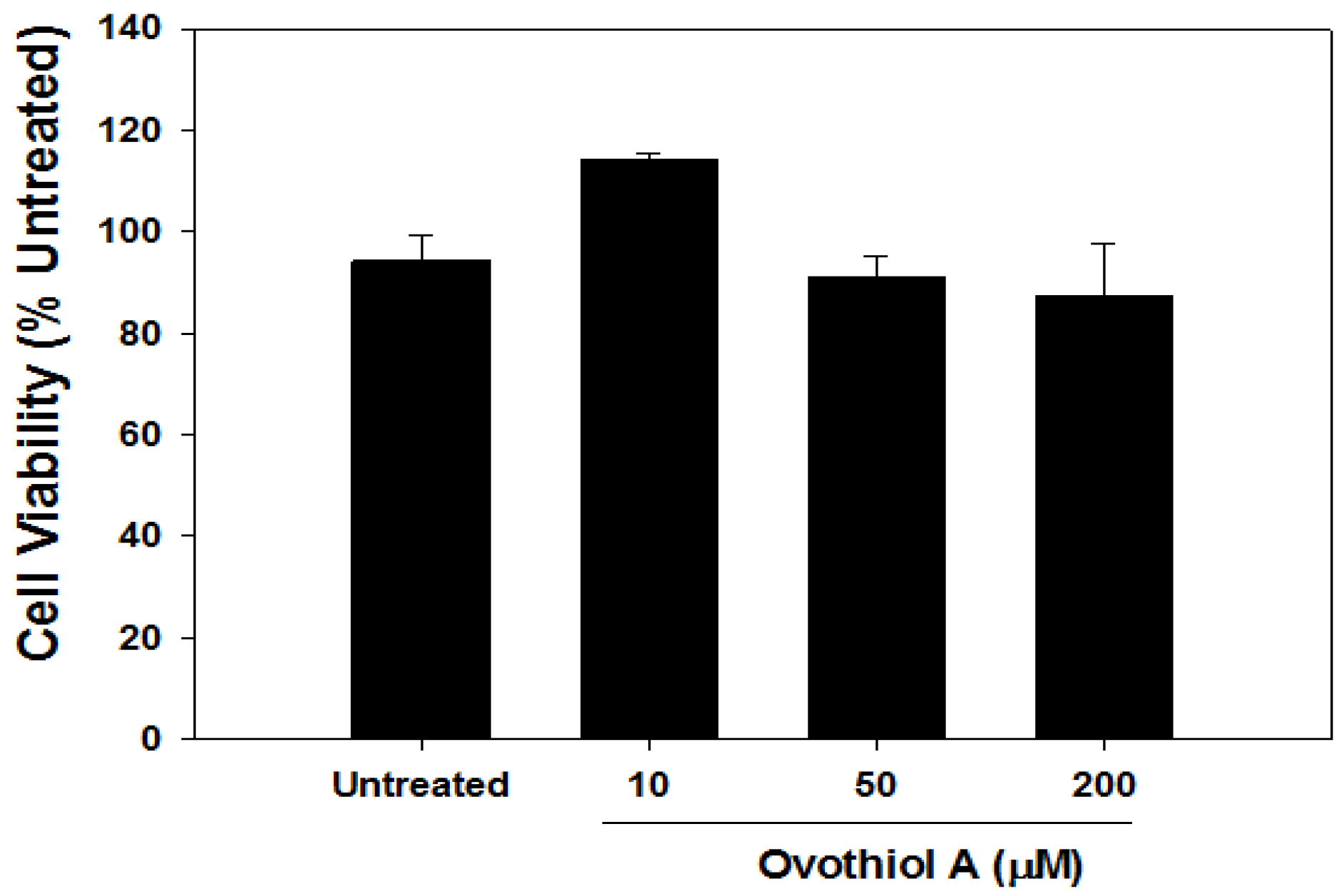
3. Experimental Section
3.1. Animals
3.2. Chemicals and Reagents
3.3. Isolation of Ovothiol A Disulfide
3.4. Cell Culture and Viability
3.5. Measurement of Autophagy
3.6. Immunoblots
3.7. Intracellular ROS Measurement
3.8. Ferrous Oxidation-Xylenol Orange (FOX) Assay
3.9. Analysis of Ovothiol A in Cell Cultures
4. Discussion
5. Conclusions
Abbreviations
| Ovo A | 5-histidylcysteine sulfoxide synthase |
| LC3 | Microtubule-associated protein 1A/1B-light chain 3 |
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Ianora, A.; Bentley, M.G.; Caldwell, G.S.; Casotti, R.; Cembella, A.D.; Engstrom-Ost, J.; Halsband, C.; Sonnenschein, E.; Legrand, C.; Llewellyn, C.A.; et al. The relevance of marine chemical ecology to plankton and ecosystem function: An emerging field. Mar. Drugs 2011, 9, 1625–1648. [Google Scholar] [CrossRef] [Green Version]
- Sawadogo, W.R.; Schumacher, M.; Teiten, M.H.; Cerella, C.; Dicato, M.; Diederich, M. A survey of marine natural compounds and their derivatives with anti-cancer activity reported in 2011. Molecules 2013, 18, 3641–3673. [Google Scholar] [CrossRef]
- Palumbo, M.; d’Ischia, M.; Misuraca, G.; Prota, G. Isolation and structure of a new sulphur-containing aminoacid from sea urchin eggs. Tetrahedron Lett. 1982, 23, 3207–3208. [Google Scholar] [CrossRef]
- Palumbo, A.; Misuraca, G.; d’Ischia, M.; Donaudy, F.; Prota, G. Isolation and distribution of 1-methyl-5-thiol-l-histidine disulphide and a related metabolite in eggs from Echinoderms. Comp. Biochem. Physiol. 1984, 78B, 81–83. [Google Scholar]
- Rossi, F.; Nardi, G.; Palumbo, A.; Prota, G. 5-thiolhistidine, a new aminoacid from eggs of Octopus vulgaris. Comp. Biochem. Physiol. 1985, 80B, 843–845. [Google Scholar]
- Turner, E.; Klevit, R.; Hopkins, P.B.; Shapiro, B.M. Ovothiol: A novel thiohistidine compound from sea urchin eggs that confers NAD(P)H-O2 oxidoreductase activity on ovoperoxidase. J. Biol. Chem. 1986, 261, 13056–13063. [Google Scholar]
- Turner, E.; Hager, L.J.; Shapiro, B.M. Ovothiol replaces glutathione peroxidase as a hydrogen peroxide scavenger in sea urchin eggs. Science 1988, 242, 939–941. [Google Scholar] [CrossRef]
- Rohl, I.; Schneider, B.; Schmidt, B.; Zeeck, E. l-ovothiol A: The egg release pheromone of the marine polychaete Platynereis Dumerilii: Anellida: Polychaeta. Z. Naturforsch. 1999, 54c, 1145–1147. [Google Scholar]
- Hand, C.E.; Honek, J.F. Biological chemistry of naturally occurring thiols of microbial and marine origin. J. Nat. Prod. 2005, 68, 293–308. [Google Scholar] [CrossRef]
- Spies, H.S.; Steenkamp, D.J. Thiols of intracellular pathogens. Identification of ovothiol A in Leishmania donovani and structural analysis of a novel thiol from Mycobacterium bovis. Eur. J. Biochem. 1994, 224, 203–213. [Google Scholar] [CrossRef]
- Ariyanayagam, M.R.; Fairlamb, A.H. Ovothiol and trypanothione as antioxidants in trypanosomatids. Mol. Biochem. Parasitol. 2001, 115, 189–198. [Google Scholar] [CrossRef]
- Selman-Reimer, S.; Duhe, R.J.; Stockman, B.J.; Selman, B.R. l-1-N-methyl-4-mercaptohistidine disulfide, a potential endogenous regulator in the redox control of chloroplast coupling factor 1 in Dunaliella. J. Biol. Chem. 1991, 266, 182–188. [Google Scholar]
- Braunshausen, A.; Seebeck, F.P. Identification and characterization of the first ovothiol biosynthetic enzyme. J. Am. Chem. Soc. 2011, 133, 1757–1759. [Google Scholar] [CrossRef]
- Mashabela, G.T.; Seebeck, F.P. Substrate specificity of an oxygen dependent sulfoxide synthase in ovothiol biosynthesis. Chem. Commun. 2013, 49, 7714–7716. [Google Scholar] [CrossRef]
- Seebeck, F.P. Thiohistidine biosynthesis. Chimia 2013, 67, 333–336. [Google Scholar]
- Steenkamp, D.J.; Spies, H.S. Identification of a major low-molecular-mass thiol of the trypanosomatid Crithidia fasciculata as ovothiol A. Facile isolation and structural analysis of the bimane derivative. Eur. J. Biochem. 1994, 223, 43–50. [Google Scholar] [CrossRef]
- Holler, T.P.; Hopkins, P.B. Ovothiols as free-radical scavengers and the mechanism of ovothiol-promoted NAD(P)H-O2 oxidoreductase activity. Biochemistry 1990, 29, 1953–1961. [Google Scholar] [CrossRef]
- Zoete, V.; Bailly, F.; Catteau, J.P.; Bernier, J.L. Design, synthesis and antioxidant properties of ovothiolderived 4-mercaptoimidazoles. J. Chem. Soc. Perkin Trans. 1997, 1, 2983–2988. [Google Scholar]
- Bailly, F.; Zoete, V.; Vamecq, J.; Catteau, J.P.; Bernier, J.L. Antioxidant actions of ovothiol-derived 4-mercaptoimidazoles: Glutathione peroxidase activity and protection against peroxynitrite-induced damage. FEBS Lett. 2000, 486, 19–22. [Google Scholar] [CrossRef]
- Zoete, V.; Bailly, F.; Vezin, H.; Teissier, E.; Duriez, P.; Fruchart, J.C.; Catteau, J.P.; Bernier, J.L. 4-Mercaptoimidazoles derived from the naturally occurring antioxidant ovothiols 1. Antioxidant properties. Free Radic. Res. 2000, 32, 515–524. [Google Scholar] [CrossRef]
- Zoete, V.; Vezin, H.; Bailly, F.; Vergoten, G.; Catteau, J.P.; Bernier, J.L. 4-Mercaptoimidazoles derived from the naturally occurring antioxidant ovothiols 2. Computational and experimental approach of the radical scavenging mechanism. Free Radic. Res. 2000, 32, 525–533. [Google Scholar] [CrossRef]
- Vamecq, J.; Maurois, P.; Bac, P.; Bailly, F.; Bernier, J.L.; Stables, J.P.; Husson, I.; Gressens, P. Potent mammalian cerebroprotection and neuronal cell death inhibition are afforded by a synthetic antioxidant analogue of marine invertebrate cell protectant ovothiols. Eur. J. Neurosci. 2003, 18, 1110–1120. [Google Scholar] [CrossRef]
- Klappan, A.K.; Hones, S.; Mylonas, I.; Bruning, A. Proteasome inhibition by quercetin triggers macroautophagy and blocks mTOR activity. Histochem. Cell Biol. 2012, 137, 25–36. [Google Scholar] [CrossRef]
- Tai, W.T.; Shiau, C.W.; Chen, H.L.; Liu, C.Y.; Lin, C.S.; Cheng, A.L.; Chen, P.J.; Chen, K.F. Mcl-1-dependent activation of Beclin 1 mediates autophagic cell death induced by sorafenib and SC-59 in hepatocellular carcinoma cells. Cell Death Dis. 2013, 4, e485. [Google Scholar] [CrossRef]
- Klionsky, D.J.; Abdalla, F.C.; Abeliovich, H.; Abraham, R.T.; Acevedo-Arozena, A.; Adeli, K.; Agholme, L.; Agnello, M.; Agostinis, P.; Aguirre-Ghiso, J.A.; et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy 2012, 8, 445–544. [Google Scholar] [CrossRef]
- Tanida, I.; Ueno, T.; Kominami, E. LC3 and Autophagy. Methods Mol. Biol. 2008, 445, 77–88. [Google Scholar] [CrossRef]
- Kang, R.; Zeh, H.J.; Lotze, M.T.; Tang, D. The Beclin 1 network regulates autophagy and apoptosis. Cell Death Differ. 2011, 18, 571–580. [Google Scholar] [CrossRef]
- Rubinsztein, D.C.; Codogno, P.; Beth Levine, B. Autophagy modulation as a potential therapeutic target for diverse diseases. Nat. Rev. Drug Discov. 2012, 11, 709–730. [Google Scholar] [CrossRef]
- Tedesco, I.; Russo, M.; Bilotto, S.; Spagnuolo, C.; Scognamiglio, A.; Palumbo, R.; Nappo, A.; Iacomino, G.; Moio, L.; Russo, G.L. Dealcoholated red wine induces autophagic and apoptotic cell death in an osteosarcoma cell line. Food Chem. Toxicol. 2013, 60, 377–384. [Google Scholar] [CrossRef]
- Kimura, T.; Takabatake, Y.; Takahashi, A.; Isaka, Y. Chloroquine in cancer therapy: A double-edged sword of autophagy. Cancer Res. 2013, 73, 3–7. [Google Scholar] [CrossRef]
- Choi, A.M.; Ryter, S.W.; Levine, B. Autophagy in Human Health and Disease. N. Engl. J. Med. 2013, 368, 651–662. [Google Scholar] [CrossRef]
- Halliwell, B.; Whiteman, M. Measuring reactive species and oxidative damage in vivo and in cell culture: How should you do it and what do the results mean? Br. J. Pharmacol. 2004, 142, 231–255. [Google Scholar] [CrossRef]
- Halliwell, B. The wanderings of a free radical. Free Radic. Biol. Med. 2009, 46, 531–542. [Google Scholar] [CrossRef]
- Namba, M.; Nishitani, K.; Kimoto, T. Carcinogenesis in tissue culture. 29: Neoplastic transformation of a normal human diploid cell strain, WI-38, with Co-60 gamma rays. Jpn. J. Exp. Med. 1978, 48, 303–311. [Google Scholar]
- Kim, J.S.; Song, K.S.; Joo, H.J.; Lee, J.H.; Yu, I.J. Determination of cytotoxicity attributed to multiwall carbon nanotubes (MWCNT) in normal human embryonic lung cell (WI-38) line. J. Toxicol. Environ. Health A 2010, 73, 1521–1529. [Google Scholar] [CrossRef]
- Tedesco, I.; Russo, M.; Russo, G.L. Commentary on resveratrol commonly displays hormesis: Occurrence and biomedical significance. Hum. Exp. Toxicol. 2010, 29, 1029–1031. [Google Scholar] [CrossRef]
- Spagnuolo, C.; Russo, M.; Bilotto, S.; Tedesco, I.; Laratta, B.; Russo, G.L. Dietary polyphenols in cancer prevention: The example of the flavonoid quercetin in leukemia. Ann. N. Y. Acad. Sci. 2012, 1259, 95–103. [Google Scholar] [CrossRef]
- Knowles, B.B.; Howe, C.C.; Aden, D.P. Human hepatocellular carcinoma cell lines secrete the major plasma proteins and hepatitis B surface antigen. Science 1980, 209, 497–499. [Google Scholar]
- Hayflick, L.; Moorhead, P.S. The serial cultivation of human diploid cell strains. Exp. Cell Res. 1961, 25, 585–621. [Google Scholar] [CrossRef]
- Serrano, M.; Lin, A.W.; McCurrach, M.E.; Beach, D.; Lowe, S.W. Oncogenic ras provokes premature cell senescence associated with accumulation of p53 and p16INK4a. Cell 1997, 88, 593–602. [Google Scholar] [CrossRef]
- Nourooz-Zadeh, J.; Tajaddini-Sarmadi, J.; Wolff, S.P. Measurement of plasma hydroperoxide concentrations by the ferrous oxidation-xylenol orange assay in conjunction with triphenylphosphine. Anal. Biochem. 1994, 220, 403–409. [Google Scholar] [CrossRef]
- Nelson, D.A.; White, E. Exploiting different ways to die. Gene Dev. 2004, 18, 1223–1226. [Google Scholar] [CrossRef]
- Lorin, S.; Hamai, A.; Mehrpour, M.; Codogno, P. Autophagy regulation and its role in cancer. Semin. Cancer Biol. 2013, 23, 361–379. [Google Scholar] [CrossRef]
- Maiuri, M.C.; Zalckvar, E.; Kimchi, A.; Kroemer, G. Self-eating and self-killing: Crosstalk between autophagy and apoptosis. Nat. Rev. Mol. Cell Biol. 2007, 8, 741–752. [Google Scholar] [CrossRef]
- Ouyang, L.; Shi, Z.; Zhao, S.; Wang, F.T.; Zhou, T.T.; Liu, B.; Bao, J.K. Programmed cell death pathways in cancer: A review of apoptosis, autophagy and programmed necrosis. Cell Prolif. 2012, 45, 487–498. [Google Scholar] [CrossRef]
- Gibson, S.B. Investigating the role of reactive oxygen species in regulating autophagy. Methods Enzymol. 2013, 528, 217–235. [Google Scholar] [CrossRef]
- Castellano, I.; Ruocco, M.R.; Cecere, F.; Di Maro, A.; Chambery, A.; Michniewicz, A.; Parlato, G.; Masullo, M.; de Vendittis, E. Glutathionylation of the iron superoxide dismutase from the psychrophilic eubacterium Pseudoalteromonas haloplanktis. Biochim. Biophys. Acta 2008, 1784, 816–826. [Google Scholar] [CrossRef]
- Castellano, I.; Cecere, F.; de Vendittis, A.; Cotugno, R.; Chambery, A.; Di Maro, A.; Michniewicz, A.; Parlato, G.; Masullo, M.; Avvedimento, E.V.; et al. Rat mitochondrial manganese superoxide dismutase: Amino acid positions involved in covalent modifications, activity, and heat stability. Biopolymers 2009, 91, 1215–1226. [Google Scholar] [CrossRef]
- Russo, G.L.; Russo, M. Ovothiol induces cell death in different human transformed cell lines. 2014; Unpublished work. [Google Scholar]
- Zhang, X.; Chen, L.X.; Ouyang, L.; Cheng, Y.; Liu, B. Plant natural compounds: Targeting pathways of autophagy as anti-cancer therapeutic agents. Cell Prolif. 2012, 45, 466–476. [Google Scholar] [CrossRef]
- Zhou, M.; Wang, R. Small-molecule regulators of autophagy and their potential therapeutic applications. Chem. Med. Chem. 2013, 8, 694–707. [Google Scholar] [CrossRef]
- Orlikova, B.; Legrand, N.; Panning, J.; Dicato, M.; Diederich, M. Anti-inflammatory and anticancer drugs from nature. Cancer Treat. Res. 2014, 159, 123–143. [Google Scholar] [CrossRef]
- Lozy, F.; Karantza, V. Autophagy and cancer cell metabolism. Semin. Cell Dev. Biol. 2012, 23, 395–401. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Russo, G.L.; Russo, M.; Castellano, I.; Napolitano, A.; Palumbo, A. Ovothiol Isolated from Sea Urchin Oocytes Induces Autophagy in the Hep-G2 Cell Line. Mar. Drugs 2014, 12, 4069-4085. https://doi.org/10.3390/md12074069
Russo GL, Russo M, Castellano I, Napolitano A, Palumbo A. Ovothiol Isolated from Sea Urchin Oocytes Induces Autophagy in the Hep-G2 Cell Line. Marine Drugs. 2014; 12(7):4069-4085. https://doi.org/10.3390/md12074069
Chicago/Turabian StyleRusso, Gian Luigi, Maria Russo, Immacolata Castellano, Alessandra Napolitano, and Anna Palumbo. 2014. "Ovothiol Isolated from Sea Urchin Oocytes Induces Autophagy in the Hep-G2 Cell Line" Marine Drugs 12, no. 7: 4069-4085. https://doi.org/10.3390/md12074069







