Functional and Structural Overview of G-Protein-Coupled Receptors Comprehensively Obtained from Genome Sequences
Abstract
: An understanding of the functional mechanisms of G-protein-coupled receptors (GPCRs) is very important for GPCR-related drug design. We have developed an integrated GPCR database (SEVENS http://sevens.cbrc.jp/) that includes 64,090 reliable GPCR genes comprehensively identified from 56 eukaryote genome sequences, and overviewed the sequences and structure spaces of the GPCRs. In vertebrates, the number of receptors for biological amines, peptides, etc. is conserved in most species, whereas the number of chemosensory receptors for odorant, pheromone, etc. significantly differs among species. The latter receptors tend to be single exon type or a few exon type and show a high ratio in the numbers of GPCRs, whereas some families, such as Class B and Class C receptors, have long lengths due to the presence of many exons. Statistical analyses of amino acid residues reveal that most of the conserved residues in Class A GPCRs are found in the cytoplasmic half regions of transmembrane (TM) helices, while residues characteristic to each subfamily found on the extracellular half regions. The 69 of Protein Data Bank (PDB) entries of complete or fragmentary structures could be mapped on the TM/loop regions of Class A GPCRs covering 14 subfamilies.1. Introduction
G-protein-coupled receptors (GPCRs) are membrane proteins characterized by seven transmembrane (TM) helices. They respond to various ligands such as, biological amines, peptides, hormones, and odorant substances, from the extracellular side of the cell. Such stimulations induce GPCRs to activate G-proteins and to transmit signals to the interior of the cell. GPCRs exist in most cells and the abnormal signal transduction of GPCRs is related to various serious conditions, such as allergy, heart trouble, cancer, high blood pressure, and inflammation, etc. As approximately 30% of the medicines distributed worldwide are designed to control this receptor system [1,2], many researchers in both academia and industry are trying to reveal the functional mechanisms of GPCRs. For a long time, experimental difficulties in both structure determination and gene expression process had prevented researchers from understanding the functional mechanisms of GPCRs. However, recently, a large amount of genome information from many species has become available and several new structures of GPCRs have been revealed. In this context, it is now possible to overview sequences and structure spaces to understand the general rule of GPCR function by using bioinformatics. For this purpose, we have identified GPCR genes from the genome sequences of various species and have stored available information from approximately 60 subfamilies in an integrated database (SEVENS http://sevens.cbrc.jp/). From the viewpoint of collecting GPCR sequences, many useful GPCR databases are already available: GPCRDB [3], IUPHAR (GPCR database) [4], GPCR-PD™ [5], ORDB [6], gpDB [7], Human-gpDB [8], and GLIDA [9], etc. GPCRDB is the most popular database and includes known GPCR sequences from UniProt and GENBANK. IUPHAR and GPCR-PD™ accumulate bibliographic information as well as sequence information. ORDB focuses on the olfactory receptor, a subfamily of GPCRs. Furthermore, unique data regarding the interaction of a GPCR with G-proteins, and with effectors are summarized in gpDB and Human-gpDB, whereas the relationship between GPCRs and their ligands are included in GLIDA. These databases are well-organized and useful for the analysis of known GPCR genes. However, to perform an overview of all GPCR genes, it is necessary to treat a comprehensive dataset that should include not only the expressed sequences but also the newly identified sequences that cannot be detected by in vivo experiments, although they definitely exist on the genome sequence. The SEVENS database is suitable for this purpose. In this review, we report the results of a survey of sequence and structure information of GPCRs using the SEVENS database.
2. Method: SEVENS Database
First, we identified GPCR genes from genome sequences using a computational gene discovery pipeline that is composed of the GPCR gene finding stage and the GPCR gene screening stage [10-12]. At the gene finding stage, the genomic regions to which exon sequences of known GPCRs showed significant match were selected by using TBLASTN and complete gene structures were constructed from exon regions by using the ALN program [13]. Then, we defined a gene as a pseudogene when a stop codon was found at the exon region or when there was a frame shift by insertion or deletion. The most accurate dataset (96.6% sensitivity and 99.4% specificity) was obtained by combining several outputs that were calculated by using the optimized threshold of the sequence similarity search by BLASTP, the GPCR specific Pfam domain assignments by HMMER [14], and the TM helix prediction by SOSUI [15] at the gene screening stage. Finally, screened sequences that showed hits to known GPCRs in UniProt ( http://www.uniprot.org/) with E-values < 10−30 against BLASTP search were categorized to the same subfamily of known GPCRs. Short protein sequences with less than 150 residues were eliminated.
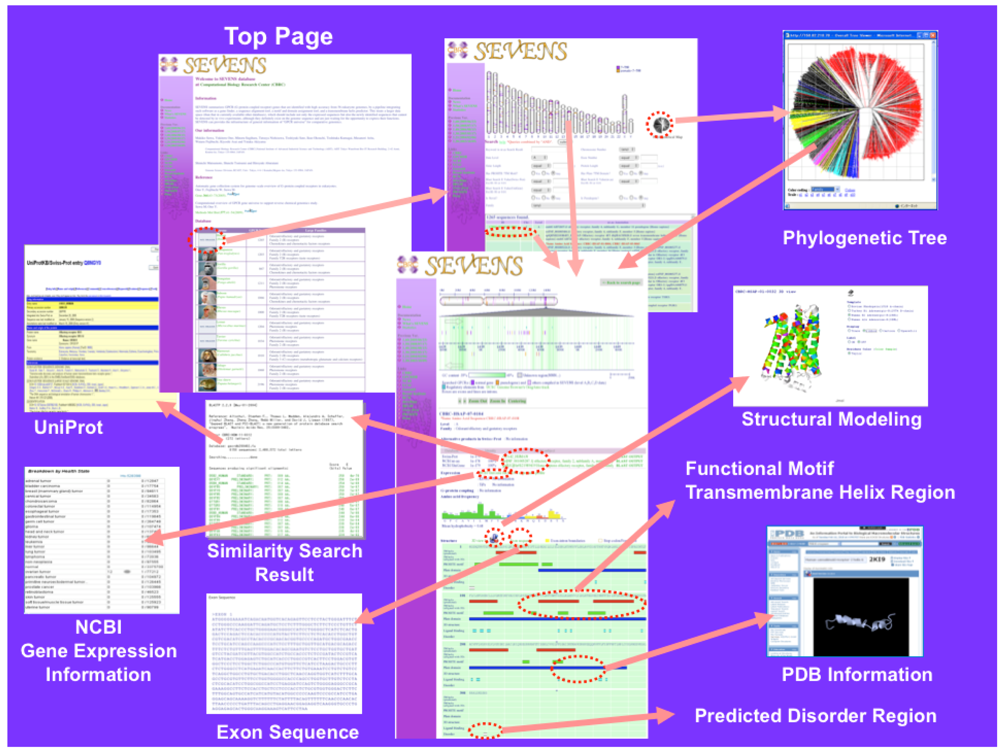
Currently, the SEVENS database (version 1.70) stores 64,090 genes from 56 eukaryotes and more than half (35,125 genes) of the total genes are olfactory receptor. It is an integrated database in which various kinds of functional and structural information of each GPCR gene are visually presented and organized in a hierarchical manner. Figure 1 shows the Web page of the SEVENS database. The top page shows a list of eukaryote genomes. When a species type is selected, the entry search page is shown. Users can access the gene annotation information page from the chromosomal map, the phylogenetic icon, or the search condition entry form.
In the gene annotation information page, the genetic coordinate of a selected GPCR is displayed on the chromosomal map together with information of known regulatory regions, GC contents, and other genes near the query. Furthermore, such information as sequence similarity search results, gene expression patterns, ligand binding, G-protein binding, amino acid sequence composition, predicted TM helix regions by SOSUI [15], functional motif regions, domain regions, predicted disorder regions by DISOPRED [16], exon-intron boundaries, pseudogenes, novel genes, and regions of known structures, is available.
Because several three-dimensional (3D) structures [17-25] had already been elucidated for the Class A (Family 1) type GPCRs, we could assign actual TM helix regions of this Class. The TM regions of the Class A GPCRs were determined based on the available crystal structures of bovine rhodopsin, adrenergic receptors, and adenosine receptors to which multiple sequence alignments of the selected sequences were assigned with the avoidance of gaps in the TM regions. Based on this information, the actual TM helix regions are displayed on the gene annotation information page, and a modeled structure of selected a Class A GPCR using the comparative modeling method is presented by Jmol viewer [26]. The sequence regions that show significant match to the large coverage region of the PDB sequence are represented by purple bars.
Using the aligned sequences we performed the statistical analysis of subfamily distribution, gene structure, amino acid residue conservation, and 3D structure information. For the analysis of subfamily distribution and gene structure, we used all sequences deposited in SEVENS database [10-12]. For the analysis of conserved residues in GPCRs, we retrieved the sequences of Class A GPCRs and selected 1,388 sequences from 49 vertebrate species in each subfamily, by comparing sequences in SEVENS with those in UniProt, with the criteria: Amino acid identity and sequence coverage rate are more than 80% and 90%, respectively. And finally, 1,272 sequences having complete lengths, which belong to 48 subfamilies of 31 vertebrate species, except olfactory/gustatory receptors, were considered for statistical analysis.
3. Results and Discussion
3.1. Annotated Subfamily Distribution
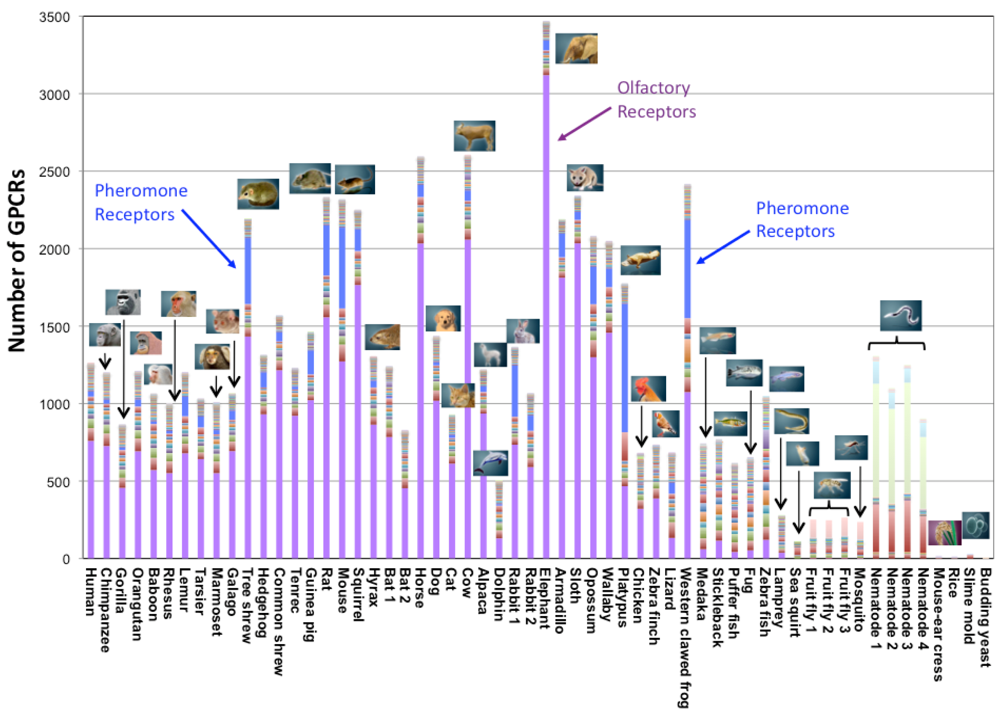
In eukaryotes, we identified a few GPCRs in yeasts, a dozen in plants, approximately 200 in insects, several hundreds in fishes and birds, and from several hundreds to several thousands in mammals. When we broke down those numbers into the subfamily classification (Figure 2), the number of subfamilies (represented by different colors in Figure 2) was around several in the lower species. It increased rapidly in the higher species after the fish, and became nearly 60 in mammals (e.g., 57 subfamilies in the case of human). Within those subfamilies, the number of receptors for biological amines, peptides, lipids, etc. is conserved in mammals, birds, and fishes. Currently, eight subfamilies are commonly found in all species, except for Yeasts and Plants. They are acetylcholine receptors, adrenaline receptors, dopamine receptors, serotonin receptors, Class B (Family 2) receptors, Class C (Family 3) receptors, Frizzled/Smoothened receptors, and glycoprotein receptors. The first four receptors affect nervous system control and the remaining four are related to the interaction between cells. (Genes of yeasts and plants are excluded from this analysis because they have a small number of GPCRs, making it difficult to apply comparative genome analysis.)
On the other hand, the number of receptors for chemical substances is distributed uniquely in different species. For example, the percentage of olfactory receptors (colored in purple in Figure 2) is approximately 70% in GPCR genes and shows much diversity in mammals. The percentage of olfactory receptors is approximately 60% in human and primates, and is extremely high (more than 80%) in elephant, armadillo, sloth, cow, horse, squirrel, etc. In contrast, one sea animal (dolphin) has a rather small number of olfactory receptors. Other types of olfactory receptors that are peculiar to insects (Drosophila odorant receptor) account for approximately half of all the genes of two kinds of insects (Drosophila and mosquito). It is interesting that the number of pheromone receptors (colored in blue in Figure 2) increases only in certain species: Tree shrew, rabbit, mouse, rat, platypus, and western clawed frog. More than 700 nematode chemosensory receptors are found in C. elegans. In addition, around 130 trace amine receptors are found, particularly in zebrafish.
Interestingly, such chemical substance receptors that are characteristic to each species show biased distribution on the genome sequences. Those receptors form large clusters in which several dozens of genes are found at high densities. For example, large clusters appear on human chromosome 11, mouse chromosome 1, dog chromosomes 18 and 21, and C. elegans chromosome 5, etc. In contrast, genes found in most of the other subfamilies are sparsely distributed on the genome sequences. Previous studies have also suggested that species-specific families form gene clusters on the genome sequences [27-32]. In general, it is suggested that gene duplication occurs on the genome sequence when the number of cluster members increases.
From the viewpoint of comprehensive collection of GPCRs, Fredriksson et al. [33] created a phylogenetic tree of approximately 800 known human GPCR sequences and proposed a new taxonomy, “GRAFS,” which consisted of five main groups. This was applied to 13 kinds of eukaryotes [34]. With regards to the comprehensive GPCR gene analysis of individual species, many publications, including those that deal with insect [35], plant [36], human and mouse [37,38], and human and dog [39], are available. The GPCR gene number collected by our pipeline is in good agreement with the previous studies, although it is somewhat larger [10-12].
3.2. Gene Structure
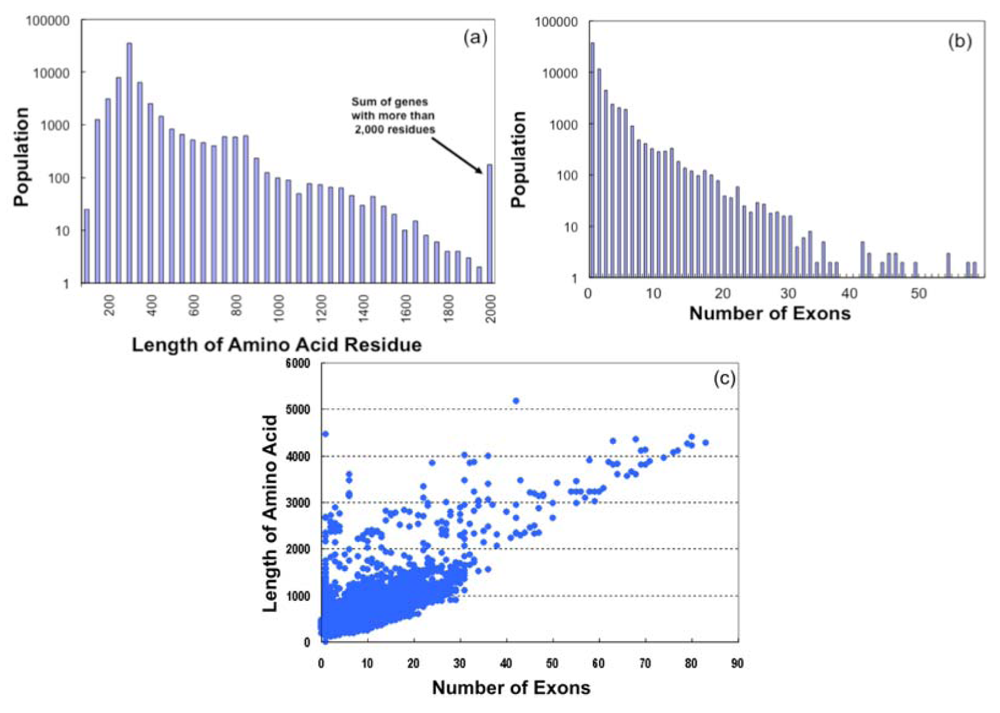
On examination of the amino acid lengths of GPCR sequences (Figure 3a), it was found that most of the GPCR sequences contain 250 ∼ 400 residues (367 residues on average), which are mainly the lengths of olfactory receptor sequences, while such receptors as Class B and Class C types have more than 1,000 residues. The longest receptor sequence was found in Class C, which has 5,179 amino acid residues.
Based on the exon-intron mapping information stored in the SEVENS database, we obtained the number of exons for each gene. Figure 3b suggests that most GPCR genes in 64,090 sequences are of the single exon type (37,459, ca. 58%) or a few exon type which are consisted of mostly olfactory receptors. In contrast, the number of genes with more than 10 exons is 2,465, with a Class C receptor gene having the largest exon number (Nmax = 83). The large number of exons in most of such genes is due to the extracellular loop regions of Class B and Class C receptors. Figure 3c shows the amino acid length (L) plotted as a function of the number of exons (N). Large clusters are found in the region where L < 2000 and N < 30, and a linear relationship (L ∼50*N) is observed where N > 30.
From the gene mapping information, we observed that exons have long dispersion genomic regions of 2,000,000 nt at most. It is natural that GPCR genes with a large number of exons require long genomic regions, although there is no clear correlation between the number of exons and the length of the dispersion region of each exon. We found a gene with 14 exons requiring the longest dispersion region (1,982,894 nt) and this region is longer than that of the gene with the largest exon number (Nmax = 83).
Based on our method of gene identification, the genomic coordinate of some GPCRs is the product of prediction and is not confirmed by experiments. However, the results of statistical analysis seem to be similar to that of previous work that dealt with all of the genes [40-42]. In contrast, GPCR-specific features are observed in our analyses, i.e., the content of single-exon-type sequences mainly related to olfactory receptors is high.
3.3. Residue Conservation
To understand the functional mechanism of GPCRs, it is important to study the amino acid residues conserved in GPCR sequences. Conserved residues at the extracellular side affect ligand-binding selectivity, whereas those at the cytoplasmic side affect G-protein coupling selectivity. Many articles have reported the positions of conserved residues [43-49] and the mutation experiments of key residues that have significant influence on the ligand binding or G-protein coupling selectivity [43,45-47,49].
For statistical analysis of conserved residues, we retrieved the sequences of Class A GPCRs deposited in the SEVENS database [10-12], because it is the major Class that accounts for approximately 70% of all GPCRs and several full-length 3D structures for this families have already been revealed [17-25]. The 1,272 sequences, having complete lengths, from 48 subfamilies of 31 vertebrate species, except olfactory/gustatory receptors, were considered for statistical analysis (See “Method section” for detail). Here, strongly conserved residues in the 1,272 sequences were observed in each TM helix region: Asn (100%, Asn55: residue with the corresponding position of bovine rhodopsin) in TM1; Asp (97%, Asp83) in TM2; Cys (90%, Cys110), Ser (82%, Ser127), Asp (68%, Glu134), Arg (98%, Arg135), and Tyr (74%, Tyr136) of DRY motif in TM3; Trp (97%, Trp161) and Pro (63%, Pro170) in TM4; Pro (82%, Pro215) and Tyr (88%, Tyr223) in TM5; Phe (84%, Phe261), Cys (75%, Cys264), Trp (78%, Trp265) and Pro (100%, Pro267) of CWxP motif in TM6; and Asn (80%, Asn302), Pro (95%, Pro303), and Tyr (92%, Tyr306) of NPxxY motif in TM7. Most of these conserved residues are concentrated on the cytoplasmic side of TM helices while residues characteristic to each subfamily were found on the extracellular side. It is suggested that DRY motif area form hydrogen bonding networks between surrounding TM helices [48] and is important for G-protein coupling selectivity [49]. This motif area is almost fully occupied by a major combination of Asp-Arg-Tyr (57%) or Glu-Arg-Tyr (10%).
Statistics of the 1,272 vertebrate sequences show that the length of the third cytoplasmic loop (CL3) varies considerably (average: 34), ranging from 1 (chemokine receptor) to 230 (acetylcholine receptor). A similar tendency was observed in the N- and C-terminal loops. It is interesting that almost all loops of these N- and C-terminal loops and CL3 are predicted to have disorder regions by using DISOPRED program [16]. In contrast, the other loops have nearly the same lengths: 7 (the first Extracellular Loop (EL) 1), 24 (EL2), 13 (EL3), 7 (the Cytoplasmic Loop (CL) 1), and 12 (CL2) with several residues dispersion at most, suggesting that the structures around TM 1, 2, 3, and 4 are similar.
3.4. Structure Space
Because GPCRs exist in the membrane environment, crystallization is difficult and the number of 3D structures available is very small. However, in the past two to three years, the number of 3D structures of GPCRs has increased rapidly. β1- and β2-adrenergic receptors [18-20], A2A adenosine receptors [21], and squid rhodopsin [22,23] were crystallized and their 3D structures completely solved. Furthermore, in 2010, the structures of CXCR4 chemokine [24] and D3 dopamine receptors [25] were successfully determined. This situation depends greatly on recent technological developments [50], such as (1) a method to express large amounts of functioning receptors by introducing genes into insect cell membrane; (2) technology for crystallization promotion by CL3 modification due to insertion of a globular protein or binding to an antibody protein; (3) crystallization technology in the environment that is very similar to that of native membrane; and (4) data extraction technology from a fine crystal with a very bright X-ray beam-line having less than 10 μm width.
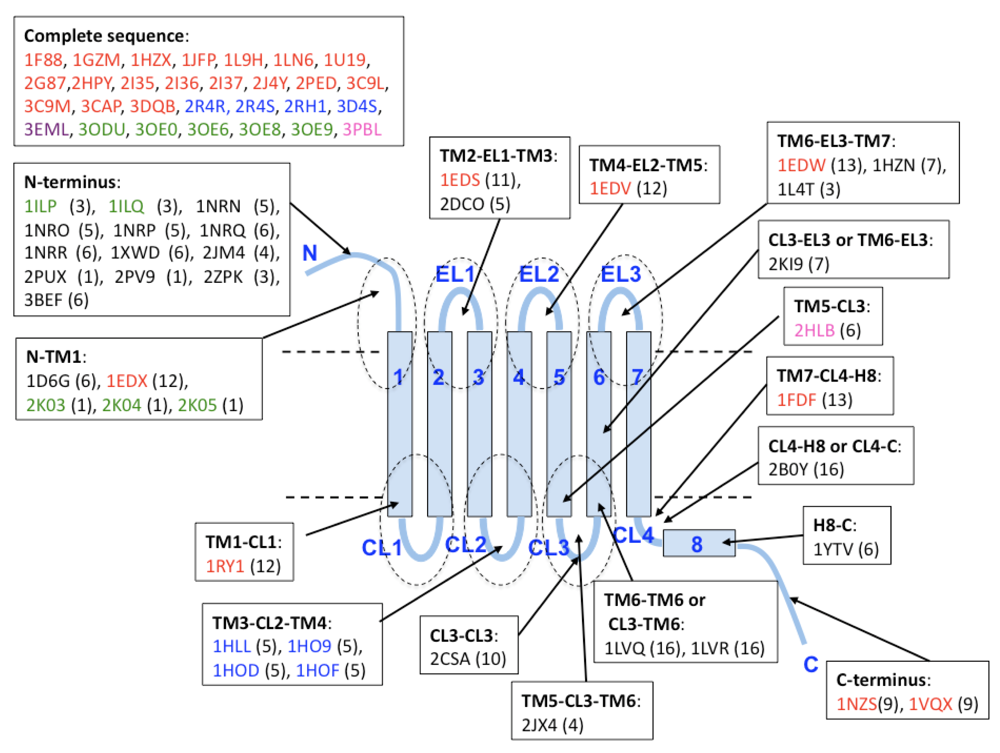
Beside the PDB entries of complete structures (29 structures of five subfamilies), there are many fragmentary structures related to GPCRs. For Class A GPCRs, we can compute the statistics of the fragmentary structures mapped on the TM helix and loop regions as the actual boundary of the two regions can be obtained by alignment to known structures [17-25].
These positions are schematically shown in Figure 4. For fragmentary structure mapping, we used the criterion of more than 80% similarity between whole sequences in PDB and those in the SEVENS database (see Methods section) and therefore, one PDB structure matches several sequences in SEVENS. The 69 PDB entries in Figure 4 are the products of X-ray crystallography (38 structures), NMR measurement (30 structures) or electron microscopy (1 structure), although the coordinates predicted by comparative modeling are excluded. We observed 22 fragmentary structure entries on the N- and C-terminal loop regions (18 in N-terminus and 4 in C-terminus) and 18 in the other loops, which included few residues of the edge of TM regions. The resolution of those structures varies from 1.2 Å (the most accurate resolution) to 4 Å. The structure of bovine rhodopsin shows the largest number (26 entries), because it can be determined in many experimental conditions. These coordinates cover 14 subfamilies (muscarinic receptor, acetylcholine receptor, adenosine/adenine nucleotide receptors, adrenergic receptor, cannabinoid receptor, chemokine/chemotactic factor receptors, cholecystokinin/gastrin receptor, dopamine receptor, glycoprotein hormone receptor, lysolipid receptor, opsins, proteinase-activated receptor, relaxin receptor, and vasopressin receptor).
Meanwhile, most of the revealed structures for Class B and Class C GPCRs are the products from long N- and C-terminal loops. It should be carefully examined why some of the domains of non GPCR proteins (globular proteins), such as tubulin-like domain, SH3-like domain, etc., showed a significant match with long extracellular loops of GPCR families (bile acid receptor, trace amine receptor, somatostatin and urotensin receptors, olfactory and gustatory receptors, melanocortin receptor, oxytocin receptor, and UNKNOWN types). Given the current situation that 3D structures are still rare for GPCRs, this is important information, even though they are only fragmentary structures.
4. Conclusions
In this review, we overviewed the sequences and structure spaces of GPCRs in the integrated SEVENS database from the viewpoint of family distribution, gene structure, amino acid residue conservation, and 3D structure. As the dataset stored in SEVENS has been identified accurately and comprehensively from complete genome sequences, the available information and the results of analyses are expected to reveal the GPCR proteome and the evolutional diversity of GPCR functions.
It is necessary to take into account the recent achievements in the research on the 3D structures of GPCRs. For a long time, bovine rhodopsin is the only GPCR whose 3D structure has been solved and it has been used as a template for the comparative modeling of GPCRs analyzed in drug discovery researches. However, from 2007 to 2010, the GPCR structures of different families [17-25] were determined by X-ray crystallography and conventional research methods are expected to change rapidly.
From the new 3D structures, the diversities in the structures of the ligand binding sites and the G-protein binding sites have become clear and should be considered in the case of studying different families. Thus, it is necessary to determine the 3D structures of all major GPCR families. However, such high throughput structure determination is impossible because the producing functional receptors in the membrane and crystallization of receptors are bottlenecks. Therefore, structural information must be obtained through a new approach that does not directly depend on 3D structure determination. It is important to extract and overview information that reflects the available 3D structures for each family at the sequence level by bioinformatics, and the SEVENS database has been constructed to serve this purpose.

Acknowledgements
This work was supported by a Grant-in-Aid for Scientific Research (B) (Kakenhi 20300104) from the Japanese Ministry of Education, Culture, Sports, Science and Technology. We thank Wataru Fujibuchi for useful discussions.
References
- Vauquelin, G.; Mentzer, B.G. Protein-Coupled Receptors; John Wiley & Sons, Ltd.: West Sussex, UK, 2007. [Google Scholar]
- Rosenbaum, D.M.; Rasmaussen, S.G.F.; Kobilka, B.K. The structure and function of G-protein-coupled receptors. Nature 2009, 459, 356–363. [Google Scholar]
- Horn, F.; Vriend, G.; Cohen, F.E. Collecting and harvesting biological data: the GPCRDB and NucleaRDB information systems. Nucl. Acid. Res. 2001, 29, 346–349. [Google Scholar]
- Harmar, A.J.; Hills, R.A.; Rosser, E.M.; Jones, M.; Buneman, O.P.; Dunbar, D.R.; Greenhill, S.D.; Hale, V.A.; Sharman, J.L.; Bonner, T.I.; et al. IUPHAR-DB: The IU-PHAR database of G protein coupled receptors and ion channels. Nucl. Acid. Res. 2009, 37, D680–D685. [Google Scholar]
- Hodges, P.E.; Carrico, P.M.; Hogan, J.D.; O'Neill, K.E.; Owen, J.J.; Mangan, M.; Davis, B.P.; Brooks, J.E.; Carrels, J.I. Annotating the human proteome: The human proteome survey database (HumanPSD™) and an in-depth target database for G protein-coupled receptors (GPCR-PD™) from incyte genomics. Nucl. Acid. Res. 2002, 30, 137–141. [Google Scholar]
- Crasto, C.; Marenco, L.; Miller, P.; Shepherd, G. Olfactory receptor database: A metadata-driven automated population from sources of gene and protein sequences. Nucl. Acid. Res. 2002, 30, 354–360. [Google Scholar]
- Theodoropoulou, M.C.; Bagos, P.G.; Spyropoulos, I.C.; Hamodrakas, S.J. gpDB: A database of GPCRs, G-proteins, effectors and their interactions. Bioinformatics 2008, 24, 1471–1472. [Google Scholar]
- Satagopam, V.P.; Theodoropoulou, M.C.; Stampolakis, C.K.; Pavlopoulos, G.A.; Papandreou, N.C.; Bagos, P.G.; Schneider, R.; Hamodrakas, S.J. GPCRs, G-proteins, effectors and their interactions: human-gpDB, a database employing visualization tools and data integration techniques. Database (Oxford) 2010, 2010, baq019. [Google Scholar]
- Okuno, Y.; Tamon, A.; Yabuuchi, H.; Niijima, S.; Minowa, Y.; Tonomura, K.; Kunimoto, R.; Feng, C. GLIDA: GPCR-ligand database for chemical genomics drug discovery-database and tools update. Nucl. Acid. Res. 2008, 36, D907–D912. [Google Scholar]
- Suwa, M.; Ono, Y. Computational overview of GPCR gene Universe to support reverse chemical genomics study. In Reverse Chemical Genetics, Methods in Molecular Biology, 1st ed.; Koga, H., Ed.; Springer: Tokyo, Japan, 2009; Volume 577, pp. 41–54. [Google Scholar]
- Ono, Y.; Fujibuchi, W.; Suwa, M. Automatic gene collection system for genome-scale overview of G-protein coupled receptors in eukaryotes. Gene 2005, 364, 63–73. [Google Scholar]
- Suwa, M.; Ono, Y. A bioinformatics strategy to produce a cyclically developing project structure. –Comprehensive functional analysis of the drug design target genes. Synthesiology 2010, 3, 1–12. [Google Scholar]
- Gotoh, O. Homology-based gene structure prediction: Simsplified matching algorithm using a translated codon (tron) and improved accuracy by allowing for long gaps. Bioinformatics 2000, 16, 190–202. [Google Scholar]
- Bateman, A.; Birney, E.; Durbin, R.; Eddy, S.R.; Howe, K.L.; Sonnhammer, E.L. The Pfam protein families database. Nucl. Acid. Res. 2000, 28, 263–266. [Google Scholar]
- Hirokawa, T.; Boon-Chieng, S.; Mitaku, S. SOSUI: Classification and secondary structure prediction system for membrane proteins. Bioinformatics 1998, 14, 378–379. [Google Scholar]
- Ward, J.J; McGuffin, L.J.; Bryson, K.; Buxton, B.F.; Janes, T.J. The DISOPRED server for the prediction of protein disorder. Bioinformatics 2004, 20, 2138–2139. [Google Scholar]
- Palczewski, K.; Kumasaka, T.; Hori, T.; Behnke, C.A.; Motoshima, H.; Fox, B.A.; Le Trong, I.; Teller, D.C.; Okada, T.; Stenkamp, R.E.; Yamamoto, M.; Miyano, M. Crystal structure of rhodopsin: A G protein-coupled receptor. Science 2000, 289, 739–745. [Google Scholar]
- Cherezov, V.; Rosenbaum, D.M.; Hanson, M.A.; Rasmussen, S.G.; Thian, F.S.; Kobilka, T.S.; Choi, H.J.; Kuhn, P.; Weis, W.I.; Kobilka, B.K.; et al. High-resolution crystal structure of an engineered human β2-adrenergic G protein-coupled receptor. Science 2007, 318, 1258–1265. [Google Scholar]
- Rasmussen, S.G.; Choi, H.J.; Rosenbaum, D.M.; Kobilka, T.S.; Thian, F.S.; Edwards, P.C.; Burghammer, M.; Ratnala, V.R.; Sanishvili, R.; Fischetti, R.F.; et al. Crystal structure of the human β2-adrenergic G-protein-coupled receptor. Nature 2007, 450, 383–387. [Google Scholar]
- Warne, A.; Serrano-Vega, M.J.; Baker, J.G.; Moukhametzianov, R.; Edwards, P.C.; Henderson, R.; Leslie, A.G.W.; Tate, C.G.; Schertler, G.F.X. Structure of the β1-adrenergic G protein-coupled receptor. Nature 2008, 454, 486–491. [Google Scholar]
- Jaakola, V.P.; Griffith, M.T.; Hanson, M.A.; Cherezov, V.; Chien, E.Y.; Lane, J.R.; Ijzerman, A.P.; Stevens, R.C. The 2.6 Å crystal structure of a human A2A adenosine receptor bound to antagonist. Science 2008, 322, 1211–1217. [Google Scholar]
- Murakami, M.; Kouyama, T. Crystal structure of squid rhodopsin. Nature 2008, 453, 363–368. [Google Scholar]
- Shimamura, T.; Hiraki, K.; Takahashi, N.; Hori, T.; Ago, H.; Masuda, K.; Takao, K.; Ishiguro, M.; Miyano, M. Crystal structure of squid rhodopsin with intracellularly extended cytoplasmic region. J. Biol. Chem. 2008, 283, 17753–17756. [Google Scholar]
- Wu, B.; Chien, E.Y.; Mol, C.D.; Fenalti, G.; Liu, W.; Katritch, V.; Abagyan, R.; Brooun, A.; Wells, P.; Bi, F.C.; et al. Structures of the CXCR4 chemokine GPCR with small-molecule and cyclic peptide antagonists. Science 2010, 330, 1066–1071. [Google Scholar]
- Ellen, Y.T.C.; Liu, W.; Zhao, Q.; Katritch, V.; Han, G.W.; Hanson, M.A.; Shi, L.; Newman, A.H.; Javitch, J.A.; Cherezov, V.; Stevens, R.C. Structure of the human dopamine D3 Rreceptor in complex with a D2/D3 selective agonist. Science 2010, 330, 1091–1095. [Google Scholar]
- Hanson, R.M. Jmol—A paradigm shift in crystallographic visualization. J. Appl. Crystal. 2010, 43, 1250–1260. [Google Scholar]
- Glusman, G.; Yanai, I.; Rubin, I.; Lancet, D. The complete Human Olfactory Subgenome. Genome Res. 2001, 11, 685–702. [Google Scholar]
- Yong, J.M.; Trask, B.J. The sence of smell: Genomics of vertebrate odorant receptors. Hum. Mol. Genomics 2002, 11, 1153–1160. [Google Scholar]
- Zozulya, S.; Echeverri, F.; Nguyan, T. The human olfactory receptor repertoire. Genome Biol. 2001, 2, 18:1–18:12. [Google Scholar]
- Niimura, Y.; Nei, M. Evolutionary dynamics of olfactory and other chemosensory receptor genes in vertebrates. J. Hum. Genet. 2006, 51, 505–517. [Google Scholar]
- Niimura, Y.; Nei, M. Comparative evolutionary analysis of olfactory receptor gene clusters between humans and mice. Gene 2005, 346, 13–21. [Google Scholar]
- Glusman, G.; Sosinsky, A.; Ben-Asher, E.; Avidan, N.; Sonkin, D.; Bahar, A.; Clifon, S.; Roe, B.; Ferras, C.; Demalle, J.; et al. Sequence, strructure, and evoluation of a complete human olfactory receptor gene cluster. Genome Res. 2000, 63, 2276–2245. [Google Scholar]
- Fredriksson, R.; Lagerström; Lundin, L. G.; Schiöth, H.B. The G-protein-coupled receptors in the human genome form five main families. Phylogenetic analysis, paralogon groups, and fingerprints. Mol. Pharmacol. 2003, 63, 1256–1272. [Google Scholar]
- Fredriksson, R.; Schiöth, H.B. The repertoire of G-protein-coupled receptors in fully sequenced genomes. Mol. Pharmacol. 2005, 67, 1414–1425. [Google Scholar]
- Hill, C.A.; Fox, A.N.; Pitts, R.J.; Kent, L.B.; Tan, P.L.; Chrystal, M.A.; Cravchik, A.; Collins, F.H.; Robertson, H.M. G protein-coupled receptors in Anopheles gambiae. Science 2002, 298, 176–178. [Google Scholar]
- Josefsson, L.G. Evidence for kinship between diverse G-protein coupled receptors. Gene 1999, 239, 333–340. [Google Scholar]
- Vassilatis, D.K.; Hohmann, J.G.; Zeng, H.; Li, F.; Ranchalis, J.E.; Mortrud, M.T.; Brown, A.; Rodriguez, S.S.; Weller, J.R.; Wright, A.C.; et al. The G protein-coupled receptor repertoires of human and mouse. Proc. Natl. Acad. Sci. USA 2003, 100, 4903–4908. [Google Scholar]
- Thora, K.B.; David, E.G.; Sofia, H.H.; Helena, K.; Fredriksson, R.; Helgi, B. S. Comprehensive and phylogenetic analysis of the G protein-coupled receptors in human and mouse. Genomics 2006, 88, 263–273. [Google Scholar]
- Haitina, T; Fredriksson, R.; Foord, S.M.; Schioth, H.; Gloriam, D.E. The G protein-coupled receptor subset of the dog genome is more similar to that in humans than rodents. BMC Genomes 2009, 10, 24. [Google Scholar]
- International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 2001, 409, 860–921. [Google Scholar]
- Ivashenko, A.T.; Khailenko, V.A.; Atambaeva, Sh.A. Variations of the length of exons and introns in human genome genes. Mol. Genet. 2009, 45, 16–22. [Google Scholar]
- Kaplunovsky, A.; Khailenko, V.; Bolshy, A.; Atambyeva, Sh.; Ivashenko, A. Statistics of exon lengths in animal, plants, fungi, and protists. World Acad. Sci. Engeneer. Technol. 2009, 52, 17–22. [Google Scholar]
- Nygaard, R.; Frimurer, T.M.; Holst, B.; Rosenkilde, M.M.; Schwartz, T.W. Ligand binding and micro-switched in 7TM receptor structures. Trends Pharmacol. Sci. 2009, 30, 249–259. [Google Scholar]
- Mizadegan, T.; Benkö, G.; Filipek, S.; Palczewski, K. Sequence analysis of G-Protein-coupled-receptors: similarities to rhodopsin. Biochemistry 2003, 42, 2759–2767. [Google Scholar]
- Karmik, S.S.; Gogonea, C.; Patil, S.; Saad, Y.; Takezako, T. Activation of G-protein-coupled receptors: a common molecular mechanism. Trends Endocrinol. Metab. 2003, 14, 431–437. [Google Scholar]
- Madabushi, S.; Gross, A.K.; Philippi, A.; Meng, E.C.; Wensel, T.G.; Lichtarge, O. Evolutionary trace of G protein-coupled receptors reveals clusters of residues that determine global and class-specific functions. J. Biol. Chem. 2004, 279, 8126–8132. [Google Scholar]
- Muramatsu, T.; Suwa, M. Statistical analysis and prediction of functional residues effective for GPCR-G-protein coupling selectivity. Protein Eng. 2006, 19, 277–283. [Google Scholar]
- Park, J.H.; Scheerer, P.; Hofmann, K.P.; Choe, H.W.; Ernst, O.P. Crystal structure of the ligand-free G-protein-coupled receptor opsin. Nature 2008, 454, 185–187. [Google Scholar]
- Wess, J. Molecular basis of receptor/G-protein-coupling selectivity. Pharmacol. Ther. 1998, 80, 231–264. [Google Scholar]
- Kobilka, B.; Schertler, F.X. New G-protein-coupled receptor crystal structures; insights and limitations. Trends Pharm. Sci. 2008, 29, 79–83. [Google Scholar]
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Suwa, M.; Sugihara, M.; Ono, Y. Functional and Structural Overview of G-Protein-Coupled Receptors Comprehensively Obtained from Genome Sequences. Pharmaceuticals 2011, 4, 652-664. https://doi.org/10.3390/ph4040652
Suwa M, Sugihara M, Ono Y. Functional and Structural Overview of G-Protein-Coupled Receptors Comprehensively Obtained from Genome Sequences. Pharmaceuticals. 2011; 4(4):652-664. https://doi.org/10.3390/ph4040652
Chicago/Turabian StyleSuwa, Makiko, Minoru Sugihara, and Yukiteru Ono. 2011. "Functional and Structural Overview of G-Protein-Coupled Receptors Comprehensively Obtained from Genome Sequences" Pharmaceuticals 4, no. 4: 652-664. https://doi.org/10.3390/ph4040652



