A Novel Combination of Withaferin A and Sulforaphane Inhibits Epigenetic Machinery, Cellular Viability and Induces Apoptosis of Breast Cancer Cells
Abstract
:1. Introduction
Sulforaphane (SFN) and Withaferin A (WA)
2. Results
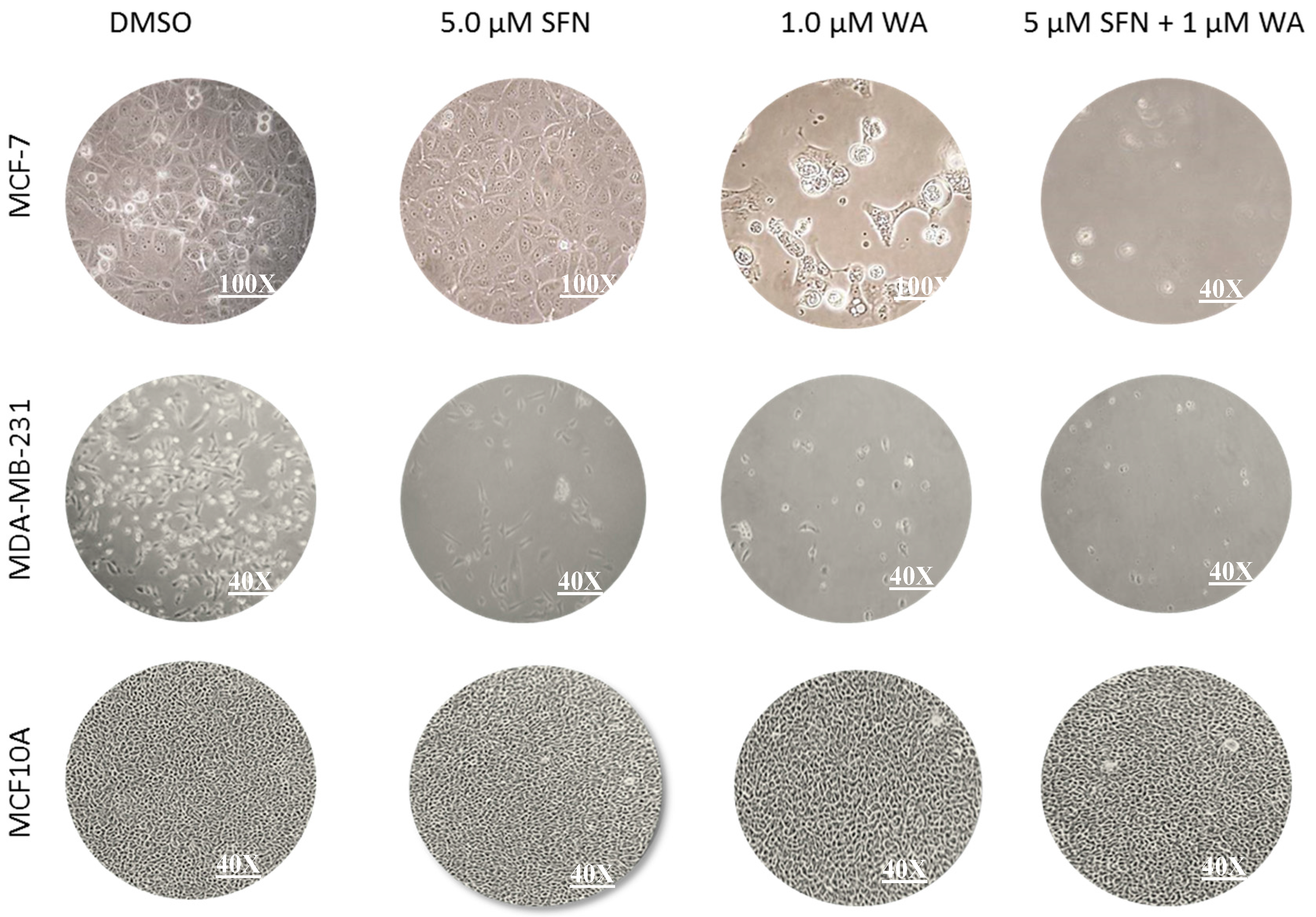
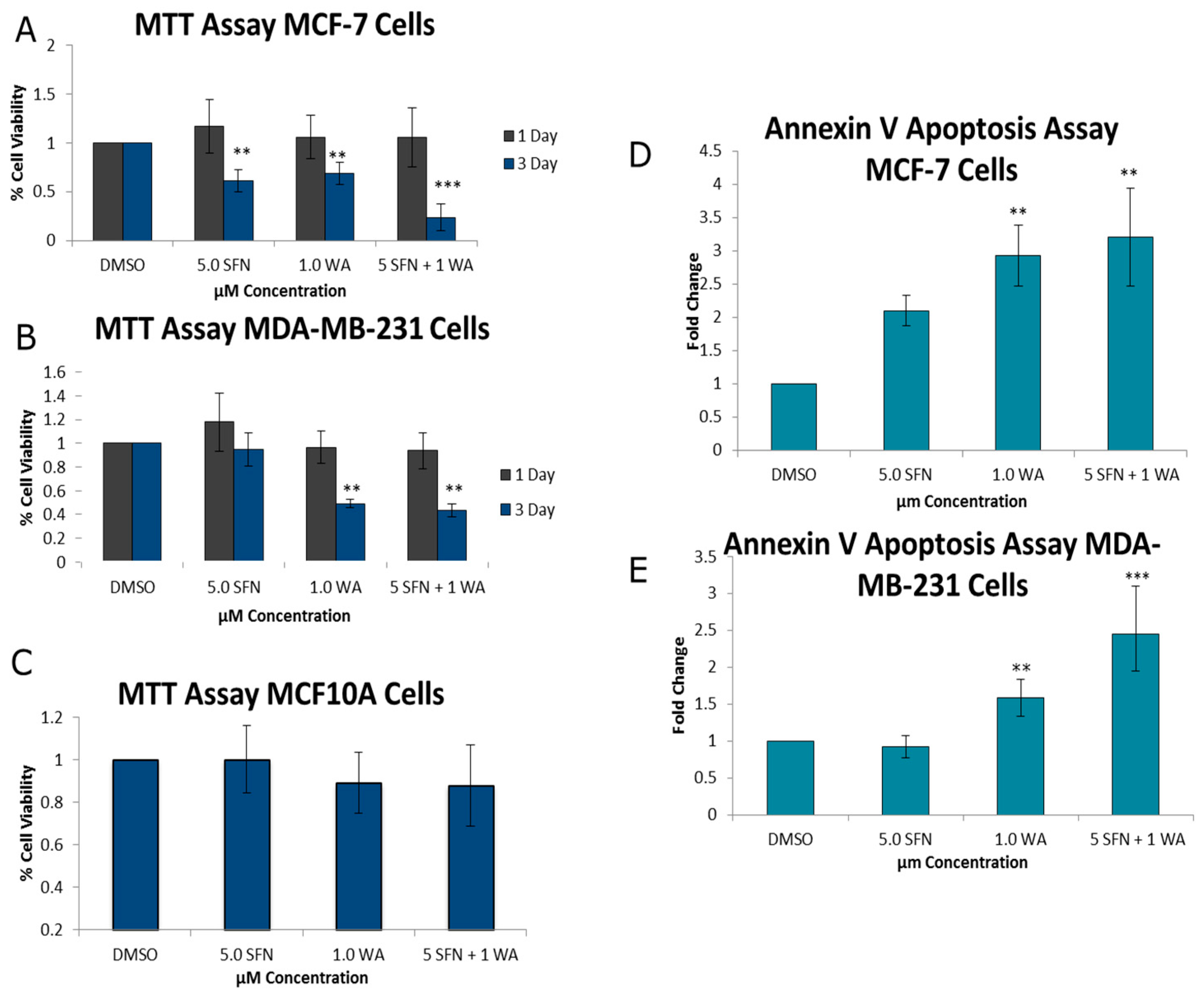
2.1. Combinatorial WA and SFN Promote Cell Death
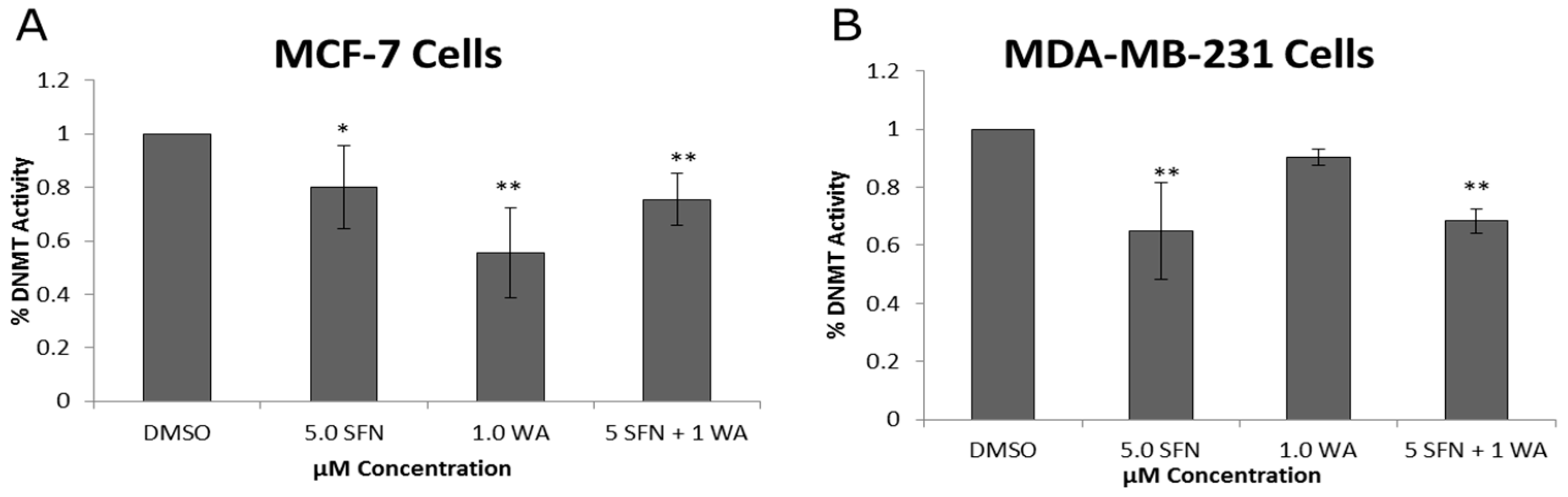
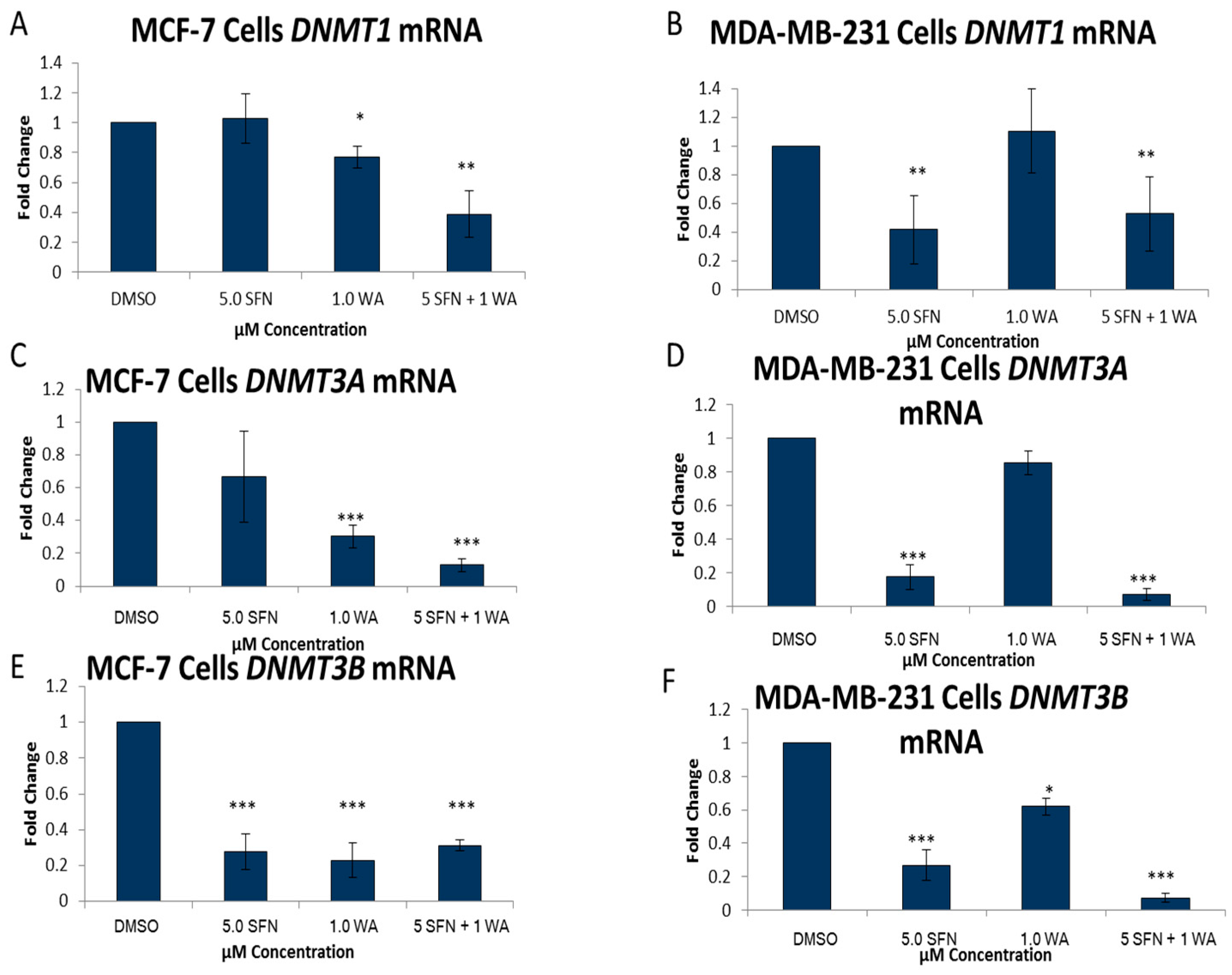
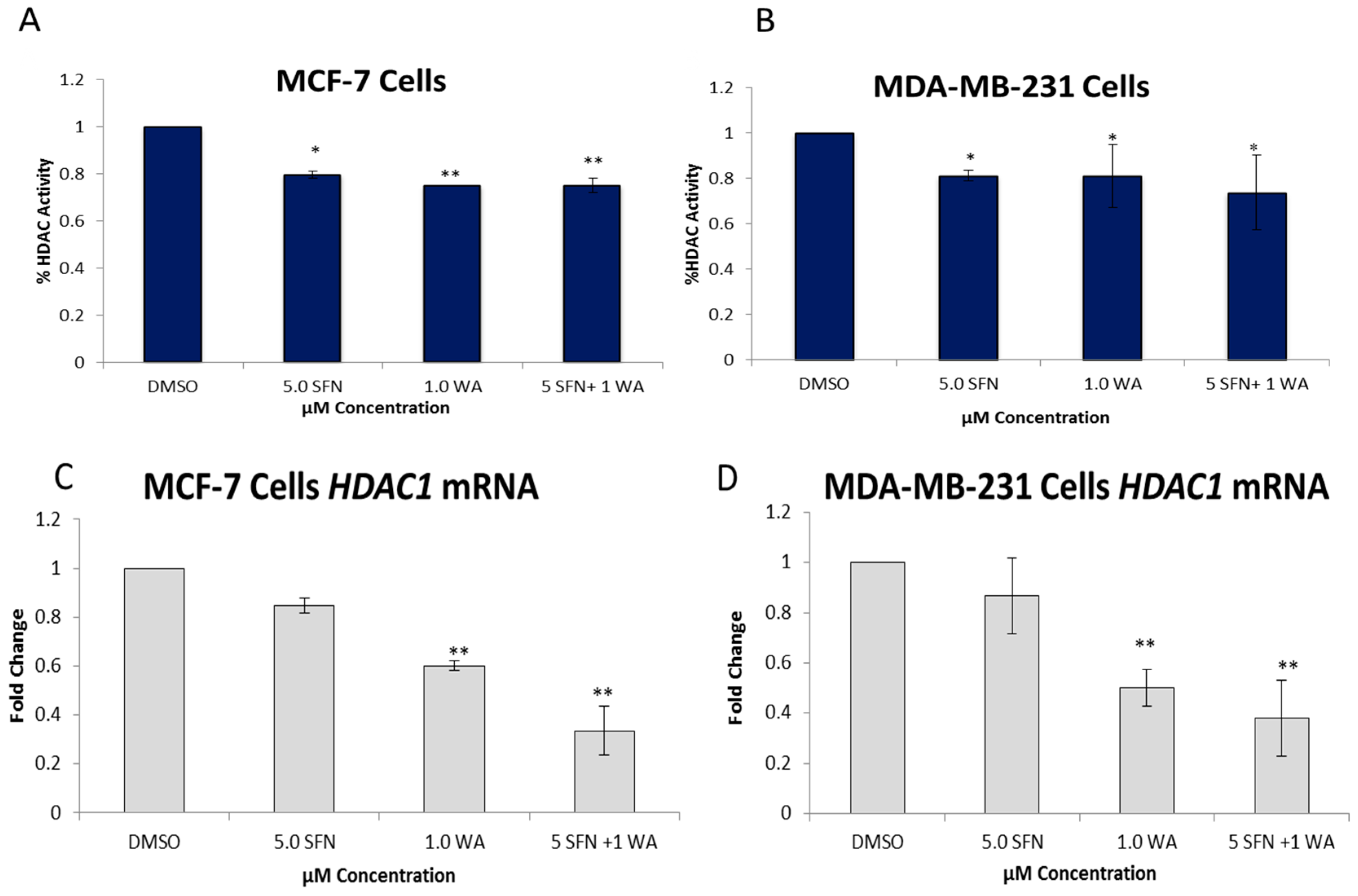
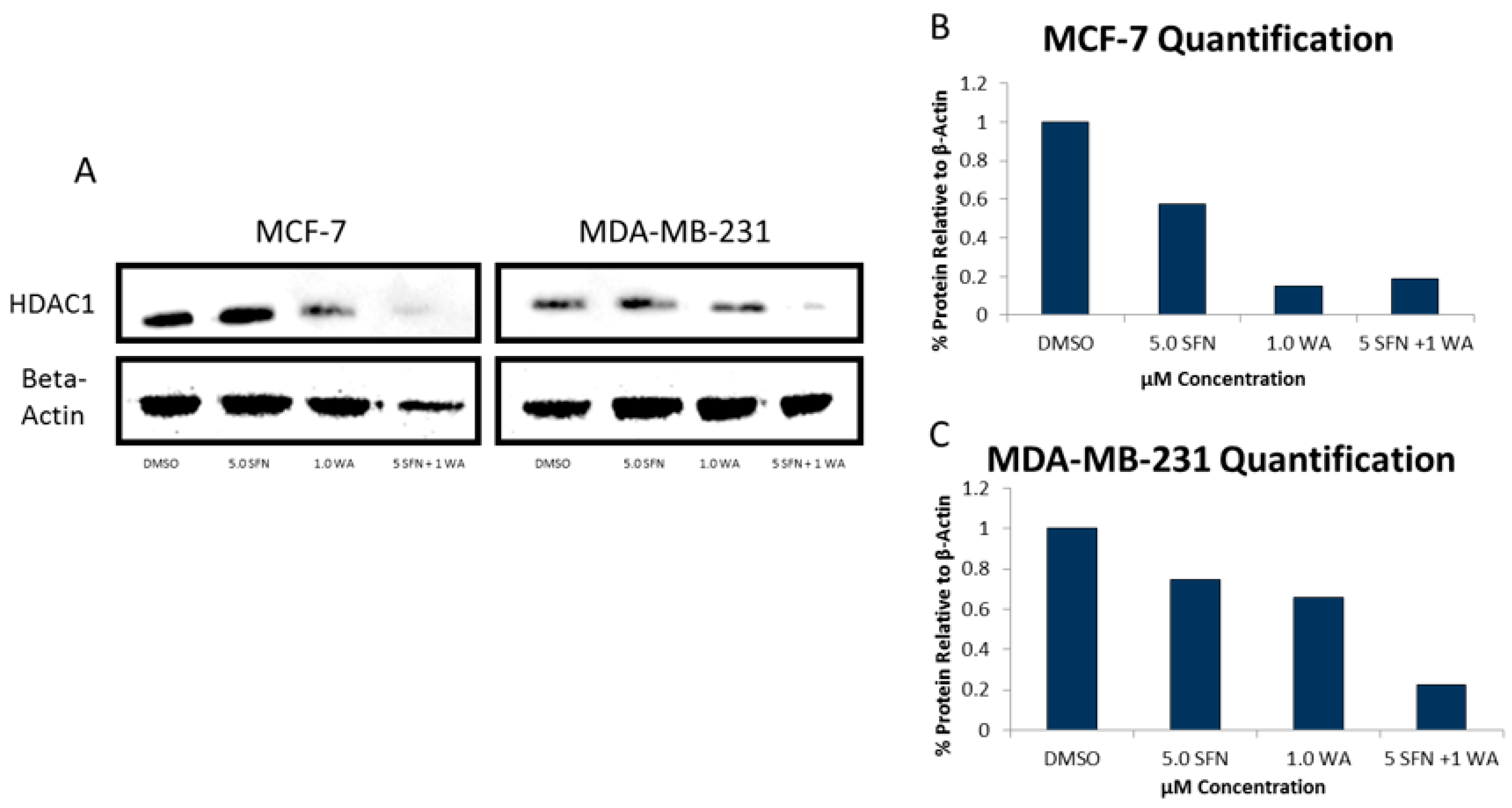
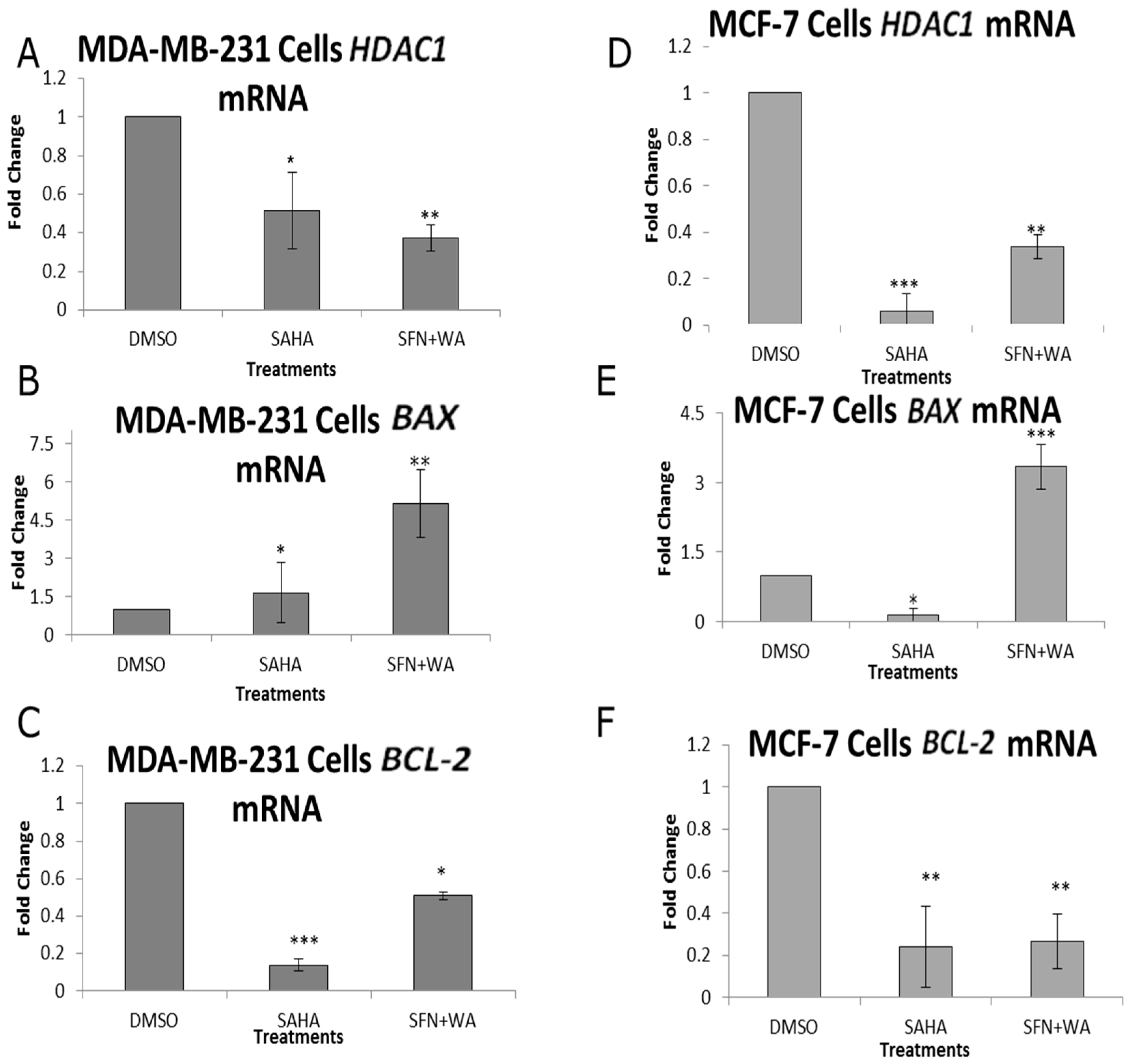
2.2. Combinatorial WA and SFN Administration Decreases HDAC Expression and Promotes Varying Changes in DNMT Expression
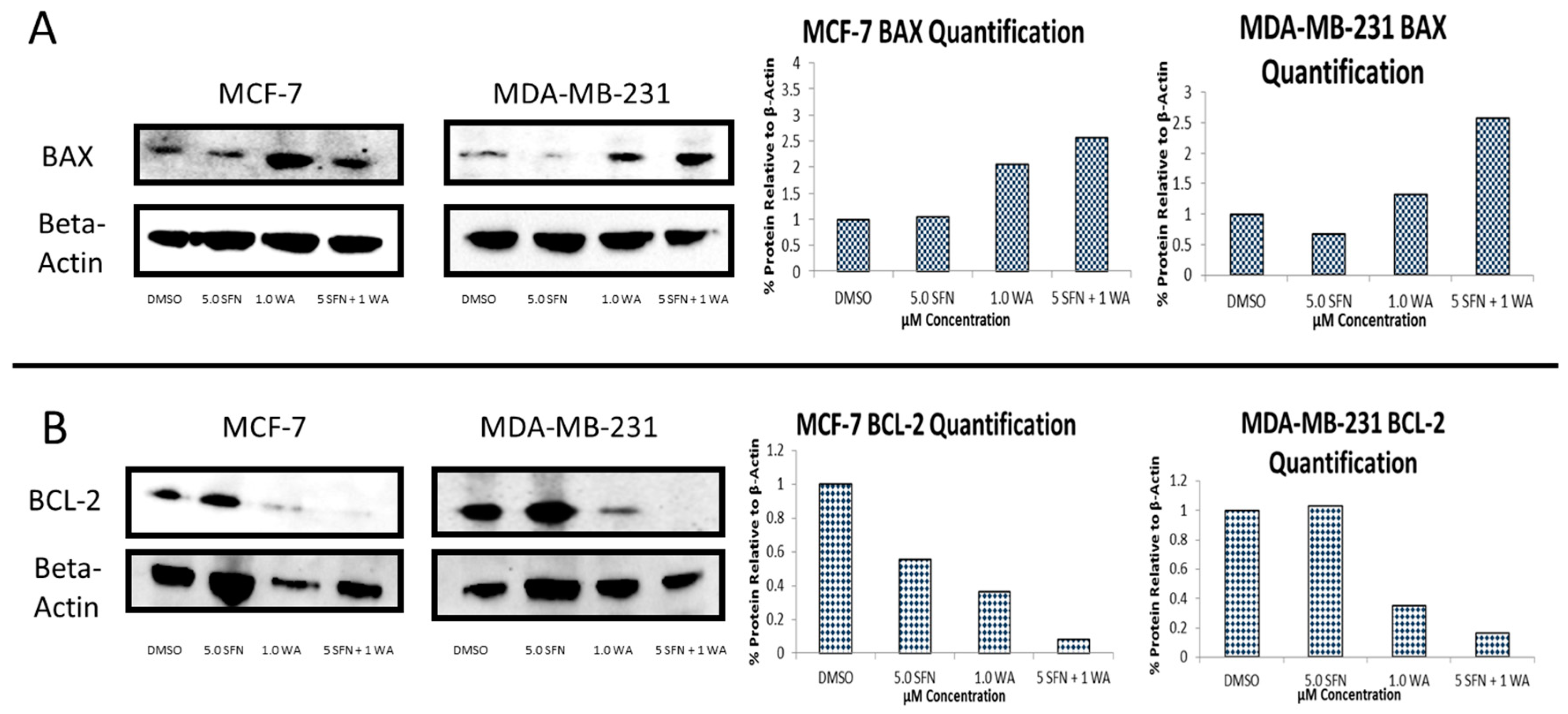
2.3. Combinatorial WA and SFN Induce Changes in BAX and BCL-2
3. Discussion
4. Materials and Methods
4.1. Cell Lines
4.2. Chemicals
4.3. Cell Culture and Treatment
4.4. Cell Density Assay
4.5. MTT Assay
4.6. RNA Isolation
4.7. Protein Extraction
4.8. Quantitative Real Time PCR (qRT-PCR)
4.9. Annexin V Apoptosis Assay FACS
4.10. Western Blot Analysis
4.11. DNMTs Activity Assay
4.12. HDACs Activity Assay
4.13. CompuSyn
4.14. Statistical Analysis
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Allis, C.D.; Jenuwein, T. The molecular hallmarks of epigenetic control. Nat. Rev. Genet. 2016, 17, 487–500. [Google Scholar] [CrossRef] [PubMed]
- Gryder, B.E.; Sodji, Q.H.; Oyelere, A.K. Targeted cancer therapy: Giving histone deacetylase inhibitors all they need to succeed. Future Med. Chem. 2012, 4, 505–524. [Google Scholar] [CrossRef] [PubMed]
- Jung, Y.; Park, J.; Kim, T.Y.; Park, J.-H.; Jong, H.-S.; Im, S.-A.; Robertson, K.D.; Bang, Y.-J.; Kim, T.-Y. Potential advantages of DNA methyltransferase 1 (DNMT1)-targeted inhibition for cancer therapy. J. Mol. Med. 2007, 85, 1137–1148. [Google Scholar] [CrossRef] [PubMed]
- Bashtrykov, P.; Jeltsch, A. DNMT1-associated DNA methylation changes in cancer. Cell Cycle 2015, 14, 5. [Google Scholar] [CrossRef] [PubMed]
- Mann, B.S.; Johnson, J.R.; Cohen, M.H.; Justice, R.; Pazdur, R. FDA approval summary: Vorinostat for treatment of advanced primary cutaneous T-cell lymphoma. Oncologist 2007, 12, 1247–1252. [Google Scholar] [CrossRef] [PubMed]
- DeSantis, C.E.; Fedewa, S.A.; Goding Sauer, A.; Kramer, J.L.; Smith, R.A.; Jemal, A. Breast cancer statistics, 2015: Convergence of incidence rates between black and white women. CA Cancer J. Clin. 2016, 66, 31–42. [Google Scholar] [CrossRef] [PubMed]
- Esmaeili, M.A. Combination of siRNA-directed gene silencing with epigallocatechin-3-gallate (EGCG) reverses drug resistance in human breast cancer cells. J. Chem. Biol. 2016, 9, 41–52. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Meeran, S.M.; Patel, S.N.; Chen, H.; Hardy, T.M.; Tollefsbol, T.O. Epigenetic reactivation of estrogen receptor-α (ERα) by genistein enhances hormonal therapy sensitivity in ERα-negative breast cancer. Mol. Cancer 2013, 12, 9. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.M.; Tsai, C.C.; Hsu, Y.C. Sulforaphane, a dietary isothiocyanate, induces G2/M arrest in Cervical cancer cells through CyclinB1 downregulation and GADD45β/CDC2 association. Int. J. Mol. Sci. 2016, 17. [Google Scholar] [CrossRef] [PubMed]
- Clarke, J.D.; Hsu, A.; Yu, Z.; Dashwood, R.H.; Ho, E. Differential effects of sulforaphane on histone deacetylases, cell cycle arrest and apoptosis in normal prostate cells versus hyperplastic and cancerous prostate cells. Mol. Nutr. Food Res. 2011, 55, 999–1009. [Google Scholar] [CrossRef] [PubMed]
- Sharma, C.; Sadrieh, L.; Priyani, A.; Ahmed, M.; Hassan, A.H.; Hussain, A. Anti-carcinogenic effects of sulforaphane in association with its apoptosis-inducing and anti-inflammatory properties in human cervical cancer cells. Cancer Epidemiol. 2011, 35, 272–278. [Google Scholar] [CrossRef] [PubMed]
- Su, Z.-Y.; Zhang, C.; Lee, J.H.; Shu, L.; Wu, T.-Y.; Khor, T.O.; Conney, A.H.; Lu, Y.-P.; Kong, A.-N.T. Requirement and epigenetics reprogramming of Nrf2 in suppression of tumor promoter TPA-induced mouse skin cell transformation by sulforaphane. Cancer Prev. Res. 2014, 7, 319–329. [Google Scholar] [CrossRef] [PubMed]
- Tomczyk, J.; Olejnik, A. Sulforaphane—A possible agent in prevention and therapy of cancer. Postepy Hig. Med. Dosw. 2010, 64, 590–603. [Google Scholar]
- Royston, K.J.; Tollefsbol, T.O. The epigenetic impact of cruciferous vegetables on cancer prevention. Curr. Pharmacol. Rep. 2015, 1, 46–51. [Google Scholar] [CrossRef] [PubMed]
- Ho, E.; Clarke, J.D.; Dashwood, R.H. Dietary sulforaphane, a histone deacetylase inhibitor for cancer prevention. J. Nutr. 2009, 139, 2393–2396. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Landen, C.N.; Li, Y.; Alvarez, R.D.; Tollefsbol, T.O. Epigallocatechin gallate and sulforaphane combination treatment induce apoptosis in paclitaxel-resistant ovarian cancer cells through hTERT and Bcl-2 down-regulation. Exp. Cell Res. 2013, 319, 697–706. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Landen, C.N.; Li, Y.; Alvarez, R.D.; Tollefsbol, T.O. Enhancement of cisplatin-mediated apoptosis in ovarian cancer cells through potentiating G2/M arrest and p21 upregulation by combinatorial epigallocatechin gallate and sulforaphane. J. Oncol. 2013, 2013, 872957. [Google Scholar] [CrossRef] [PubMed]
- Fong, M.Y.; Jin, S.; Rane, M.; Singh, R.K.; Gupta, R.; Kakar, S.S. Withaferin A synergizes the therapeutic effect of doxorubicin through ROS-mediated autophagy in ovarian cancer. PLoS ONE 2012, 7, e42265. [Google Scholar] [CrossRef] [PubMed]
- Hahm, E.-R.; Moura, M.B.; Kelley, E.E.; van Houten, B.; Shiva, S.; Singh, S.V. Withaferin A-induced apoptosis in human breast cancer cells is mediated by reactive oxygen species. PLoS ONE 2011, 6, e23354. [Google Scholar] [CrossRef] [PubMed]
- Hahm, E.R.; Singh, S.V. Withaferin A-induced apoptosis in human breast cancer cells is associated with suppression of inhibitor of apoptosis family protein expression. Cancer Lett. 2013, 334, 101–108. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.-H.; Hahm, E.-R.; Arlotti, J.A.; Samanta, S.K.; Moura, M.B.; Thorne, S.H.; Shuai, Y.; Anderson, C.J.; White, A.G.; Lokshin, A.; et al. Withaferin A inhibits in vivo growth of breast cancer cells accelerated by Notch2 knockdown. Breast Cancer Res. Treat. 2016, 157, 41–54. [Google Scholar] [CrossRef] [PubMed]
- Mirza, S.; Sharma, G.; Parshad, R.; Gupta, S.D.; Pandya, P.; Ralhan, R. Expression of DNA methyltransferases in breast cancer patients and to analyze the effect of natural compounds on DNA methyltransferases and associated proteins. J. Breast Cancer 2013, 16, 23–31. [Google Scholar] [CrossRef] [PubMed]
- Mohan, R.; Hammers, H.; Bargagna-mohan, P.; Zhan, X.; Herbstritt, C.; Ruiz, A.; Zhang, L.; Hanson, A.; Conner, B.; Rougas, J.; et al. Withaferin A is a potent inhibitor of angiogenesis. Angiogenesis 2004, 7, 115–122. [Google Scholar] [CrossRef] [PubMed]
- Chou, T.C. Preclinical versus clinical drug combination studies. Leuk. Lymphoma 2008, 49, 2059–2080. [Google Scholar] [CrossRef] [PubMed]
- Szarc vel Szic, K.; Op de Beeck, K.; Ratman, D.; Wouters, A.; Beck, I.M.; Declerck, K.; Heyninck, K.; Fransen, E.; Bracke, M.; de Bosscher, K.; et al. Pharmacological levels of Withaferin A (Withania somnifera) trigger clinically relevant anticancer effects specific to triple negative breast cancer cells. PLoS ONE 2014, 9, e87850. [Google Scholar] [CrossRef] [PubMed]
- Greenbaum, D.; Colangelo, C.; Williams, K.; Gerstein, M. Comparing protein abundance and mRNA expression levels on a genomic scale. Genome Biol. 2003, 4, 117. [Google Scholar] [CrossRef] [PubMed]
- Maier, T.; Guell, M.; Serrano, L. Correlation of mRNA and protein in complex biological samples. FEBS Lett. 2009, 583, 3966–3973. [Google Scholar] [CrossRef] [PubMed]
- Pathania, R.; Ramachandran, S.; Elangovan, S.; Padia, R.; Yang, P.; Cinghu, S.; Veeranan-Karmegam, R.; Arjunan, P.; Gnana-Prakasam, J.P.; Sadanand, F.; et al. DNMT1 is essential for mammary and cancer stem cell maintenance and tumorigenesis. Nat. Commun. 2015, 6, 6910. [Google Scholar] [CrossRef] [PubMed]
- Senese, S.; Zaragoza, K.; Minardi, S.; Muradore, I.; Ronzoni, S.; Passafaro, A.; Bernard, L.; Draetta, G.F.; Alcalay, M.; Seiser, C.; et al. Role for histone deacetylase 1 in human tumor cell proliferation. Mol. Cell Biol. 2007, 27, 4784–4795. [Google Scholar] [CrossRef] [PubMed]
- Hahm, E.R.; Lee, J.; Huang, Y.; Singh, S.V. Withaferin a suppresses estrogen receptor-α expression in human breast cancer cells. Mol. Carcinog. 2011, 50, 614–624. [Google Scholar] [CrossRef] [PubMed]
- Meeran, S.M.; Patel, S.N.; Li, Y.; Shukla, S.; Tollefsbol, T.O. Bioactive dietary supplements reactivate ER expression in ER-negative breast cancer cells by active chromatin modifications. PLoS ONE 2012, 7, e37748. [Google Scholar] [CrossRef] [PubMed]
- Liang, Y.; Yan, C.; Schor, N.F. Apoptosis in the absence of caspase 3. Oncogene 2001, 20, 6570–6578. [Google Scholar] [CrossRef] [PubMed]
- Um, H.J.; Min, K.J.; Kim, D.E.; Kwon, T.K. Withaferin A inhibits JAK/STAT3 signaling and induces apoptosis of human renal carcinoma Caki cells. Biochem. Biophys. Res. Commun. 2012, 427, 24–29. [Google Scholar] [CrossRef] [PubMed]
- Abbas, A.; Hall, J.A.; Patterson, W.L.; Ho, E.; Hsu, A.; Al-Mulla, F.; Georgel, P.T. Sulforaphane modulates telomerase activity via epigenetic regulation in prostate cancer cell lines. Biochem. Cell Biol. 2016, 94, 71–81. [Google Scholar] [CrossRef] [PubMed]
- Meeran, S.M.; Patel, S.N.; Tollefsbol, T.O. Sulforaphane causes epigenetic repression of hTERT expression in human breast cancer cell lines. PLoS ONE 2010, 5, e11457. [Google Scholar] [CrossRef] [PubMed]
- Nagalingam, A.; Kuppusamy, P.; Singh, S.V.; Sharma, D.; Saxena, N.K. Mechanistic elucidation of the antitumor properties of withaferin a in breast cancer. Cancer Res. 2014, 74, 2617–2629. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Ren, J.; Chen, H.B.; Wang, Y.; Liu, Q.; Zhang, R.; Jiang, S.W.; Li, J. Cytostatic and apoptotic effects of DNMT and HDAC inhibitors in endometrial cancer cells. Curr. Pharm. Des. 2014, 20, 1881–1887. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Samadi, A.K.; Roby, K.F.; Timmermann, B.; Cohen, M.S. Inhibition of cell growth and induction of apoptosis in ovarian carcinoma cell lines CaOV3 and SKOV3 by natural withanolide Withaferin A. Gynecol. Oncol. 2012, 124, 606–612. [Google Scholar] [CrossRef] [PubMed]
- Kala, R.; Shah, H.N.; Martin, S.L.; Tollefsbol, T.O. Epigenetic-based combinatorial resveratrol and pterostilbene alters DNA damage response by affecting SIRT1 and DNMT enzyme expression, including SIRT1-dependent gamma-H2AX and telomerase regulation in triple-negative breast cancer. BMC Cancer 2015, 15, 672. [Google Scholar] [CrossRef] [PubMed]


| Cell Lines | SFN Dose | WA Dose | Average CI |
|---|---|---|---|
| MCF-7 | 5.0 µM | 1.0 µM | 0.715503 |
| MDA-MB-231 | 5.0 µM | 1.0 µM | 1.000525 |
| Forawrd Primer Sequences | Reverse Primer Sequences |
|---|---|
| DNMT1 Sense: 5′-AACCTTCACCTAGCCCCAG-3′ | DNMT1 Anti-sense: 5′-CTCATCCGATTTGGCTCTTCA-3′ |
| DNMT3A Sense: 5′-TATTGATGAGCGCACAAGGC-3′ | DNMT3A Anti-sense: 5′-GGGTGTTCCAGGGTAACATTGAG-3′ |
| DNMT3B Sense: 5′-TGGTACATGGCTTTTCGATAGGA-3′ | DNMT3B Anti-sense: 5′-GGCAAGTTCTCCGAGGTCTCTG-3′ |
| HDAC1 Sense: 5′-CTGTCCGGTATTTGATGGCT-3′ | HDAC1 Anti-sense: 5′-CACGAACTCCACACACTTGG-3′ |
| BAX Sense: 5′-TGG AGCTGCAGAGGATGATTG-3′ | BAX Anti-sense: 5′-GAAGTTGCCGTCAGAAAACATG-3′ |
| BCL-2 Sense: 5′-CATGCTGGGGCCGTACAG-3′ | BCL-2 Anti-sense: 5′-GAACCGGCACCTGCACAC-3′ |
| GAPDH Sense: 5′-TGCACCACCAACTGCTTAGC-3′ | GAPDH Anti-Sense: 5′-GGCATGGACTGTGGTCATGAG-3′ |
| Caspase 3 Sense: 5′-TTAATAAAGGTATCCATGGAGAACACT-3′ | Caspase 3 Anti-Sense: 5′-TTAGTGATAAAAA TAGAGTTCTTTTGTGAG-3′ |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Royston, K.J.; Udayakumar, N.; Lewis, K.; Tollefsbol, T.O. A Novel Combination of Withaferin A and Sulforaphane Inhibits Epigenetic Machinery, Cellular Viability and Induces Apoptosis of Breast Cancer Cells. Int. J. Mol. Sci. 2017, 18, 1092. https://doi.org/10.3390/ijms18051092
Royston KJ, Udayakumar N, Lewis K, Tollefsbol TO. A Novel Combination of Withaferin A and Sulforaphane Inhibits Epigenetic Machinery, Cellular Viability and Induces Apoptosis of Breast Cancer Cells. International Journal of Molecular Sciences. 2017; 18(5):1092. https://doi.org/10.3390/ijms18051092
Chicago/Turabian StyleRoyston, Kendra J., Neha Udayakumar, Kayla Lewis, and Trygve O. Tollefsbol. 2017. "A Novel Combination of Withaferin A and Sulforaphane Inhibits Epigenetic Machinery, Cellular Viability and Induces Apoptosis of Breast Cancer Cells" International Journal of Molecular Sciences 18, no. 5: 1092. https://doi.org/10.3390/ijms18051092







