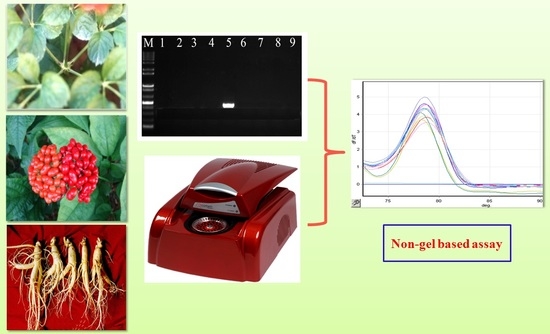
Development of EST Intron-Targeting SNP Markers for Panax ginseng and Their Application to Cultivar Authentication
Abstract
:1. Introduction
2. Results
2.1. PCR Amplification of Expressed Sequence Tags (EST)-Derived Introns
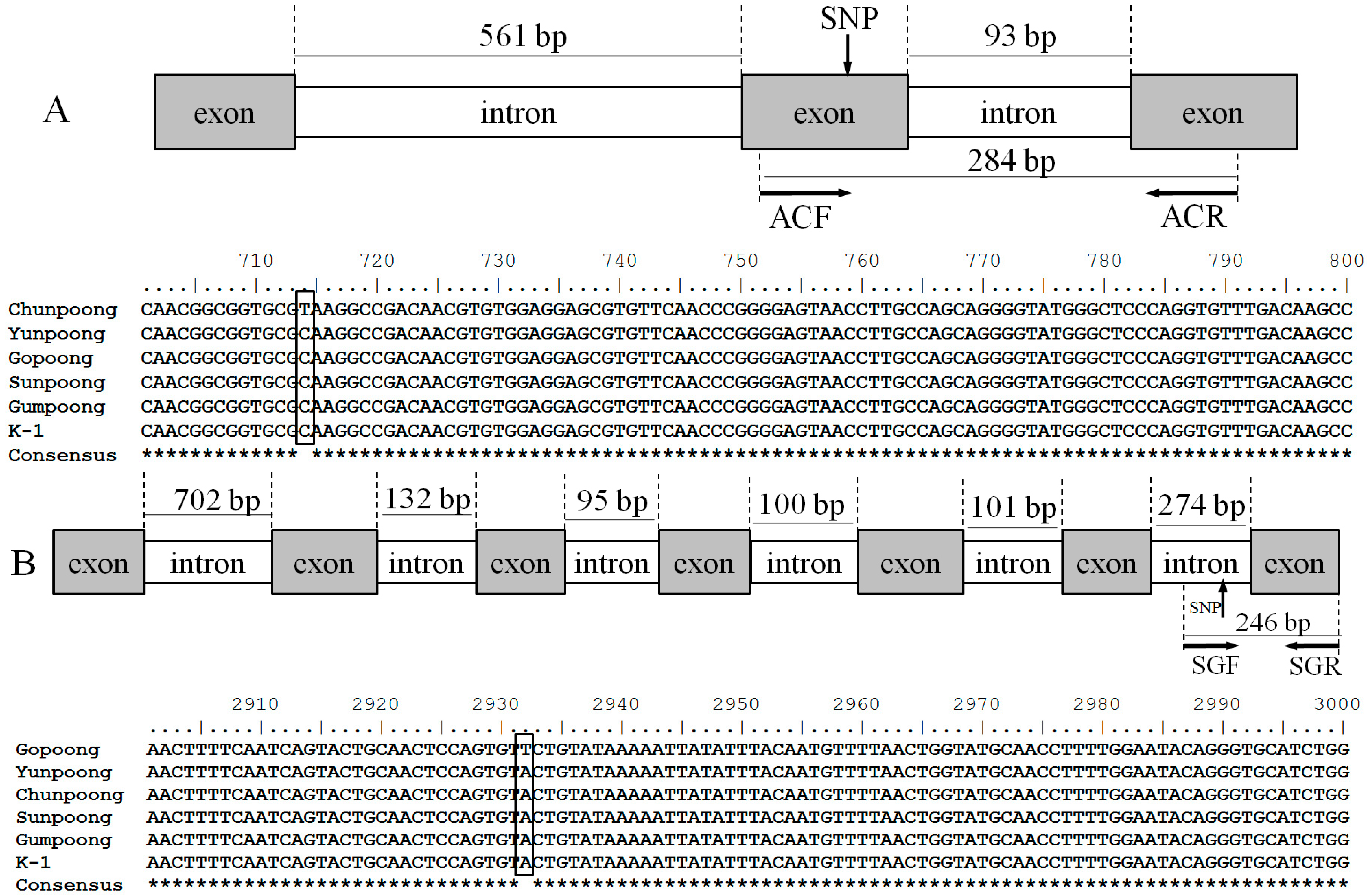
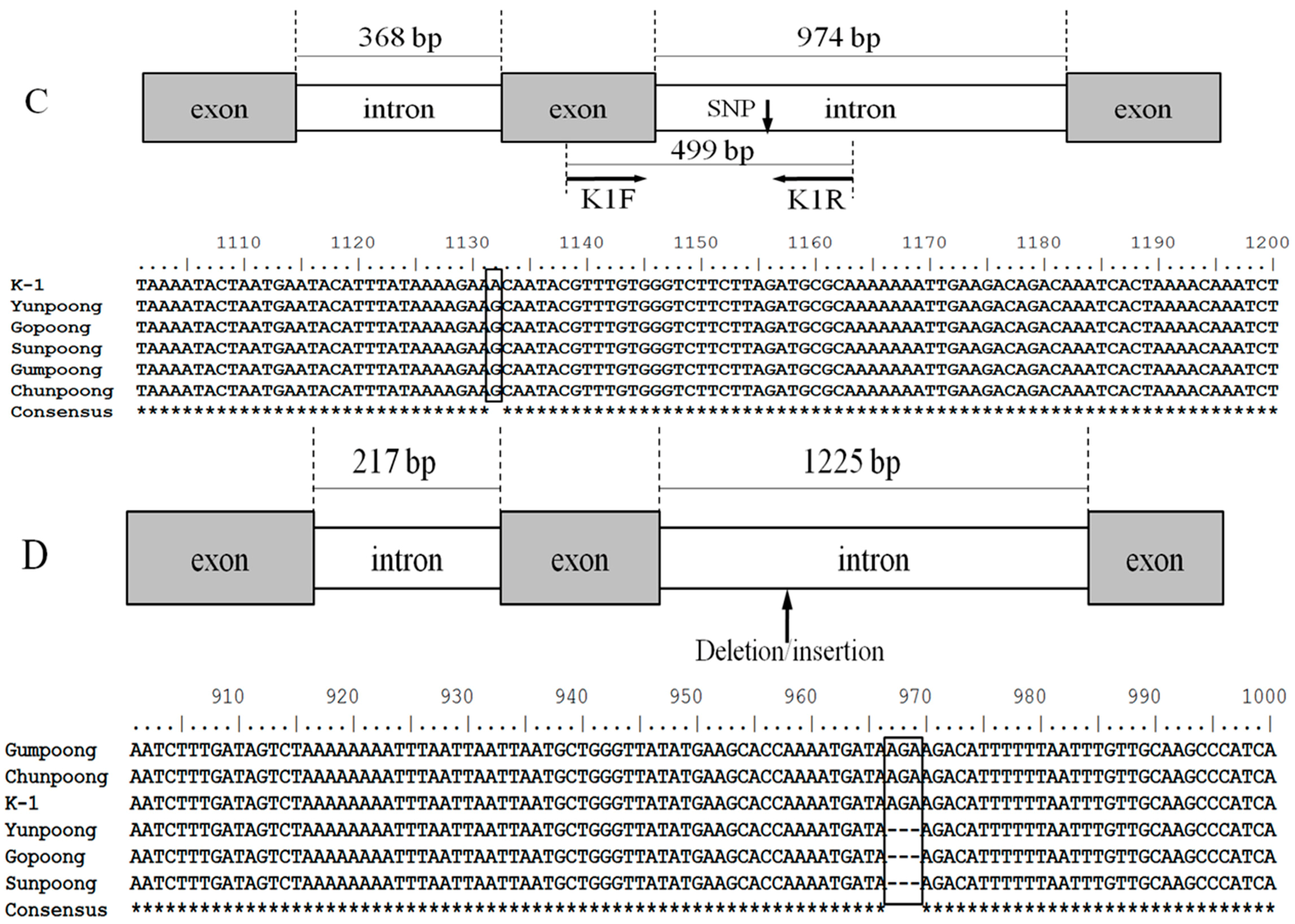
2.2. Sequence Analysis and Allele-Specific Primer Design
2.3. Allele-Specific PCR
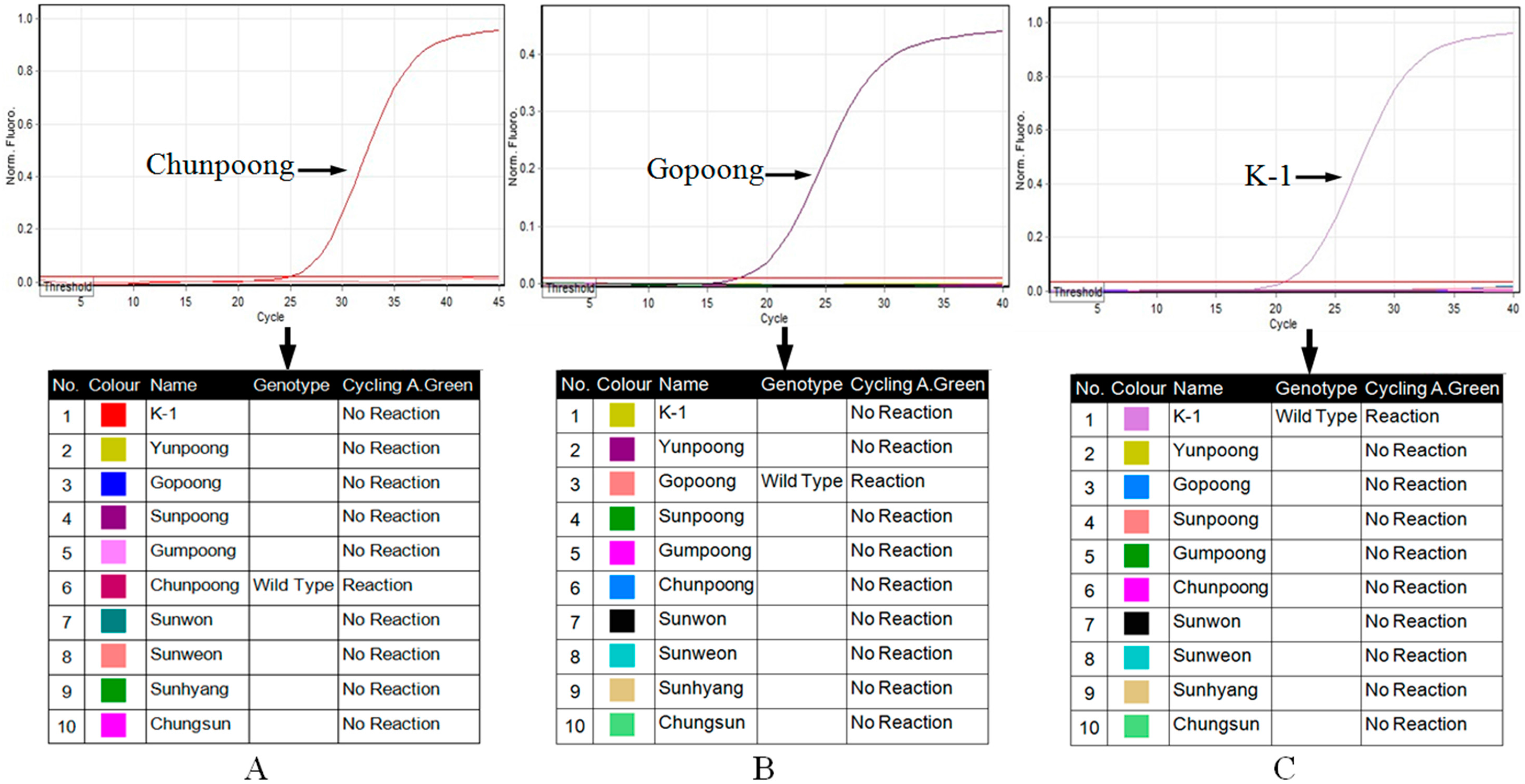
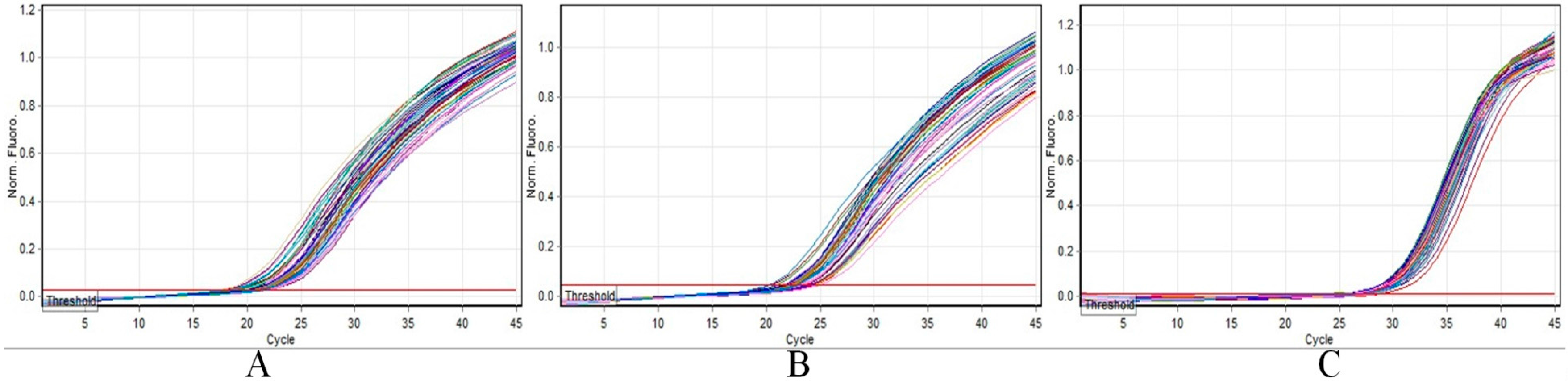
2.4. Real-Time Allele-Specific PCR Assay
3. Discussion
4. Materials and Methods
4.1. Plant Materials and DNA Isolation
4.2. Prediction of Introns and Intron-Flanking Primers Design
4.3. PCR Amplification and Sequencing
4.4. Sequence Analysis and Allele-Specific Primer Design
4.5. Allele-Specific PCR for Authentication of P. ginseng Cultivars
4.6. Real-Time Allele-Specific PCR Assay for Field Selection
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Choi, K.T. Botanical characteristics, pharmacological effects and medicinal components of Korean Panax ginseng C A Meyer. Acta Pharmacol. Sin. 2008, 29, 1109–1118. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.S.; Lee, J.H.; Ahn, I.O. Characteristics of new cultivars in Panax ginseng C.A. Meyer. In Proceedings of the Ginseng Society Conference, Seoul, Korea, November 2005; pp. 3–18.
- Reunova, G.D.; Kats, I.L.; Muzarok, T.I.; Zhuravlev Yu, N. Polymorphism of RAPD, ISSR and AFLP markers of the Panax ginseng C. A. Meyer (Araliaceae) genome. Genetika 2010, 46, 1057–1066. [Google Scholar] [CrossRef] [PubMed]
- Kim, O.; Bang, K.; In, D.; Lee, J.; Kim, Y.; Shin, Y.; Hyun, D.; Lee, S.; Cha, S.; Seong, N. Molecular authentication of ginseng cultivars by comparison of internal transcribed spacer and 5.8S rDNA sequences. Plant Biotechnol. Rep. 2007, 1, 163–167. [Google Scholar] [CrossRef]
- In, D.; Chung, J.; Kim, Y.; Bang, K.; Kim, O.-T.; Hyun, D.-Y.; Cha, S.-W.; Kim, T.-S.; Seong, N.-S. Genetic relationships of Panax species by RAPD and ISSR analyses. Korean J. Med. Crop Sci. 2005, 13, 249–253. [Google Scholar]
- Kim, N.H.; Choi, H.I.; Ahn, I.O.; Yang, T.J. EST-SSR marker sets for practical authentication of all nine registered ginseng cultivars in Korea. J. Ginseng Res. 2012, 36, 298–307. [Google Scholar] [CrossRef] [PubMed]
- Blair, M.W.; Cortes, A.J.; Penmetsa, R.V.; Farmer, A.; Carrasquilla-Garcia, N.; Cook, D.R. A high-throughput SNP marker system for parental polymorphism screening, and diversity analysis in common bean (Phaseolus vulgaris L.). Theor. Appl. Genet. 2013, 126, 535–548. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Zhu, Y.; Guo, X.; Sun, C.; Luo, H.; Song, J.; Li, Y.; Wang, L.; Qian, J.; Chen, S. Transcriptome analysis reveals ginsenosides biosynthetic genes, microRNAs and simple sequence repeats in Panax ginseng C.A. Meyer. BMC Genom. 2013, 14, 245. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Wang, H.T.; Kwon, W.S.; Kim, Y.J.; In, J.G.; Yang, D.C. A simple and rapid technique for the authentication of the ginseng cultivar, Yunpoong, using an SNP marker in a large sample of ginseng leaves. Gene 2011, 487, 75–79. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Sun, H.; Kwon, W.S.; Jin, H.; Yang, D.C. Molecular identification of the Korean ginseng cultivar “Chunpoong” using the mitochondrial nad7 intron 4 region. Mitochondrial DNA 2009, 20, 41–45. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Sun, H.; Kwon, W.S.; Jin, H.; Yang, D.C. A PCR-based SNP marker for specific authentication of Korean ginseng (Panax ginseng) cultivar “Chunpoong”. Mol. Biol. Rep. 2010, 37, 1053–1057. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Sun, H.; Kwon, W.S.; Jin, H.; Yang, D.C. A simplified method for identifying the Panax ginseng cultivar Gumpoong based on 26S rDNA. Planta Med. 2010, 76, 399–401. [Google Scholar] [CrossRef] [PubMed]
- Feltus, F.A.; Singh, H.P.; Lohithaswa, H.C.; Schulze, S.R.; Silva, T.D.; Paterson, A.H. A comparative genomics strategy for targeted discovery of single-nucleotide polymorphisms and conserved-noncoding sequences in orphan crops. Plant Physiol. 2006, 140, 1183–1191. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Chen, J.; Francis, D.M.; Shen, H.; Wu, T.; Yang, W. Discovery of intron polymorphisms in cultivated tomato using both tomato and Arabidopsis genomic information. Theor. Appl. Genet. 2010, 121, 1199–1207. [Google Scholar] [CrossRef] [PubMed]
- Sathiyamoorthy, S.; In, J.G.; Gayathri, S.; Kim, Y.J.; Yang, D.C. Generation and gene ontology based analysis of expressed sequence tags (EST) from a Panax ginseng C.A. Meyer roots. Mol. Biol. Rep. 2010, 37, 3465–3472. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Jin, G.; Zhao, X.; Zheng, Y.; Xu, Z.; Wu, W. PIP: A database of potential intron polymorphism markers. Bioinformatics 2007, 23, 2174–2177. [Google Scholar] [CrossRef] [PubMed]
- Sievers, F.; Higgins, D.G. Clustal omega. Curr. Protoc. Bioinform. 2014, 48, 1.25.1–1.25.33. [Google Scholar]

| EST No. | Intron-Flanking Primers (Forward (F) and Reverse (R), 5′→3′) | Annotation | Size of Target EST (bp) | Size of Amplicon (bp) |
|---|---|---|---|---|
| DC0_Contig 27 | F–CAATCAATCACCCACCTTTG | Metallothionein-like protein | 407 | ~1000 |
| R–TGACACAACAGGAAAGTCAAGG | ||||
| DC02017A01 | F–TATCTCGGCTTGAAGCGTCT | Cysteine proteinase | 574 | ~1950 |
| R–GTACCCCATGATCCAAATGC | ||||
| PG07020B06 | F–TACCCAGTTGTGAGCGAGGAGT | Ascorbate peroxidase | 406 | ~1300 |
| R–GGTCATTTCCCAGAGTAGCATTAG | ||||
| DC05020G06 | F–CTTGGCAAGTTCAGGAAGATG | Auxin repressed protein | 560 | 1215 |
| R–CAAACAGCAACGCTACTCGCA | ||||
| DC05003B01 | F–AGAACAAACGTGAAAGGCAT | Squalene epoxidase | 1751 | 3158 |
| R–GCCCTCGACTTATAGTCATACA | ||||
| DC0_contig 107 | F–GATAAAGGAGTTGATGTGTGAGC | Cytochrome P450 (CYP71A50U) | 1001 | 2353 |
| R–CAGGAAGGACAACACCCGAC | ||||
| DC0_contig 37 | F–GCGGGATTATTGTCTTTACCT | Cytochrome P450 (CYP716A42) | 987 | 2431 |
| R–TTGGGATAAATCACACCGTT | ||||
| DC05033A07 | F–CGATGGAACCCTCGCTTTCTTC | Peroxiredoxin | 436 | ~880 |
| R–CTCCGAGCCTGAGACAGTGAAT | ||||
| DC02026E01 | F–CGAGTGGGAATTCAGAAGGAGAT | Leucine-rich repeat protein | 465 | ~1800 |
| R–CACACAAGTTGTTGCTTGAGACA | ||||
| PG07002C02 | F–GGCTTTGACATCGTTCGTTT | Nicotinamide adenine dinucleotide phosphate (NADP)-isocitrate dehydrogenase | 450 | ~3000 |
| R–CCAAAAGCATGTCTCCCAAT | ||||
| DC04035D04 | F–GCAAAGGAGCTGGTTTCATC | Glutaredoxin | 343 | ~1700 |
| R–CATGGTGCAATTAACCCACA | ||||
| Contig03320 | F–TTATCAGCTGCAATTCAAGC | Cycloartenol synthase | 404 | ~2200 |
| R–AATCTCCATTTTCCATTTGTG | ||||
| DC02009A02 | F–GGGTGTGTTCCATGTTGATAT | pathogenesis-related protein | 570 | ~1260 |
| R–ATATTACAATACTGGATTTATTATC | ||||
| DC05025C05 | F–GTGGGAATAAAGCACAGGAT | Epoxide hydrolase | 929 | ~1790 |
| R–TTGTTGATCTCATGGGGTCT | ||||
| DC04020A06 | F–GACTGATTCCGAAGAACATT | chalcone synthase | 530 | ~1900 |
| R–GAAAATAAAAAAGGCACGGAA |
| Primer Name | Nucleotide Sequence (5′→3′) | Artificial Mismatch | Amplicon Size (bp) |
|---|---|---|---|
| ACF | ATCTCCAACGGCGGTTCGT | G→T | 284 |
| ACR | GCTATCCTCTCCTCCTCAATG | ||
| SGF | AGTACTGCAACTCCAGCGTT | T→C | 246 |
| SGR | GCCCTCGACTTATAGTCATACA | ||
| K1F | GATAGAGAAACCATCAAAGCTG | T→A | 499 |
| K1R | AGAAGACCCACAAACGTAATGT |
| Ginseng Samples | Voucher | Location | Main Characteristics |
|---|---|---|---|
| Chunpoong | GB001 | Kochang, Korea | Good root shape |
| Yunpoong | GB002 | Kochang, Korea | High yield |
| Gopoong | GB003 | Chuncheon, Korea | High saponin content |
| Sunpoong | GB004 | Kochang, Korea | Early germination |
| Gumpoong | GB005 | Kochang, Korea | High seed yield and yellow berry |
| K-1 | GB010 | Buyeo, Korea | Good root shape and well developed root hairs |
| Sunwon | GBD048 | Daejeon, Korea | High yield |
| Sunweon | GBD043 | Daejeon, Korea | Wrinkled leaf edges |
| Sunhyang | GBD058 | Daejeon, Korea | High aroma constituents |
| Chungsun | GBD073 | Daejeon, Korea | Long stem and early germination |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, H.; Li, G.; Kwon, W.-S.; Yang, D.-C. Development of EST Intron-Targeting SNP Markers for Panax ginseng and Their Application to Cultivar Authentication. Int. J. Mol. Sci. 2016, 17, 884. https://doi.org/10.3390/ijms17060884
Wang H, Li G, Kwon W-S, Yang D-C. Development of EST Intron-Targeting SNP Markers for Panax ginseng and Their Application to Cultivar Authentication. International Journal of Molecular Sciences. 2016; 17(6):884. https://doi.org/10.3390/ijms17060884
Chicago/Turabian StyleWang, Hongtao, Guisheng Li, Woo-Saeng Kwon, and Deok-Chun Yang. 2016. "Development of EST Intron-Targeting SNP Markers for Panax ginseng and Their Application to Cultivar Authentication" International Journal of Molecular Sciences 17, no. 6: 884. https://doi.org/10.3390/ijms17060884