Elucidating Pharmacological Mechanisms of Natural Medicines by Biclustering Analysis of the Gene Expression Profile: A Case Study on Curcumin and Si-Wu-Tang
Abstract
:1. Introduction
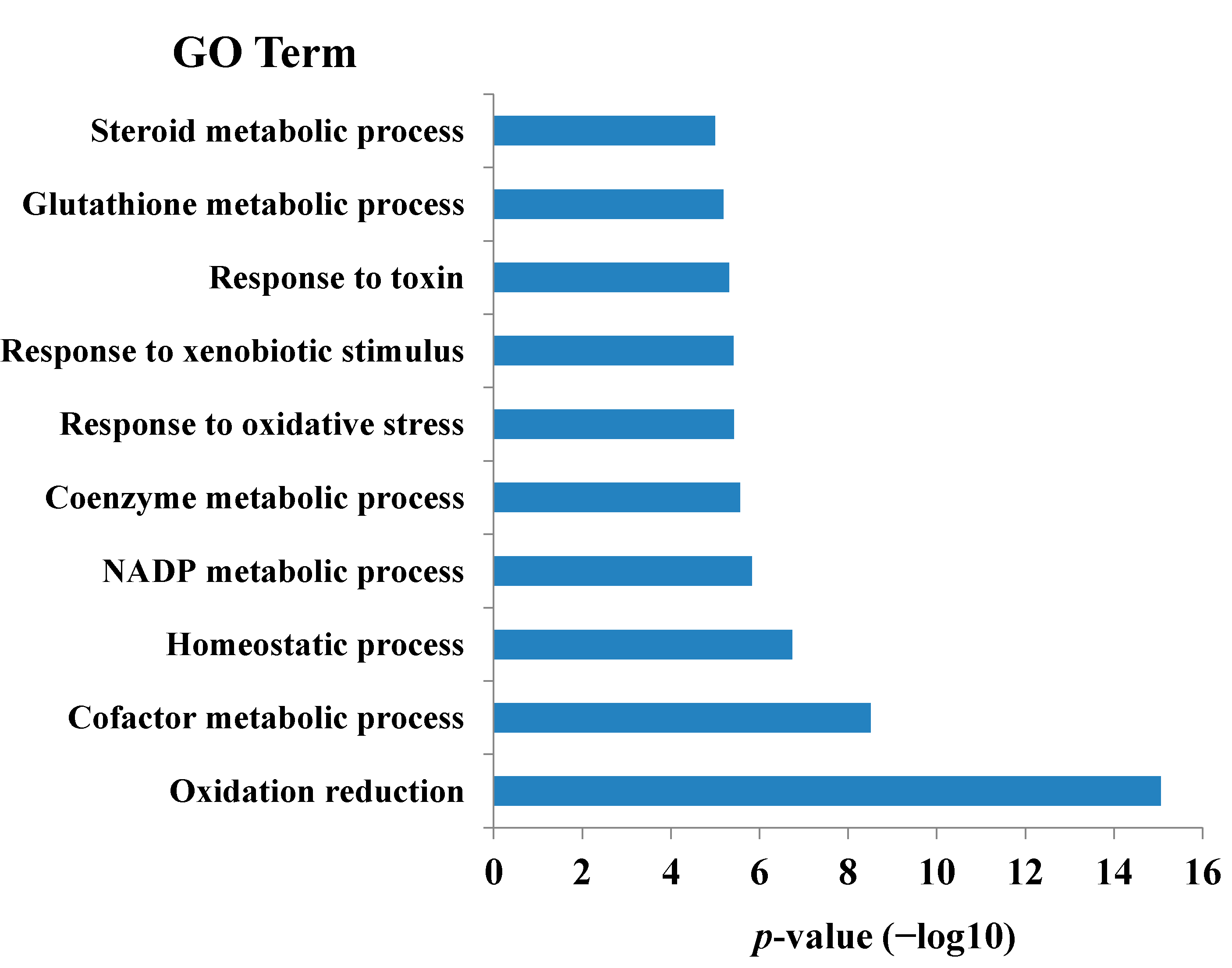
2. Results and Discussion



| Drugs | Tanimoto Coefficients | Therapeutic Uses |
|---|---|---|
| Esculin a | 0.48 | Anti-inflammatory |
| Clobetasol b | 0.42 | Anti-inflammatory Agents, Corticosteroids, Topical, Glucocorticoids |
| Iodixanol b | 0.41 | Contrast Media |
| Sulpiride b | 0.38 | Antidepressants, Antidepressive Agents, Second-generation, Antipsychotic Agents, Antipsychotics |
| Amantadine b | 0.38 | Analgesics, Non-narcotic, Antiparkinson Agents, Anti-viral Agents, Dopamine Agents |
| Oxolinic acid a | 0.38 | Anti-bacterial |
| Lomefloxacin b | 0.38 | Anti-infective Agents, Anti-infective Agents, Urinary, Antitubercular Agents, Photosensitizing Agents |
| Drugs | Tanimoto Coefficients | Therapeutic Uses |
|---|---|---|
| Clioquinol a | 0.41 | Anti-infective Agent |
| Carmustine b | 0.41 | Antineoplastic Agents, Antineoplastic Agents, Alkylating |
| Celastrol a | 0.40 | Anti-bacterial, Anti-proliferative |
| Spiperone a | 0.38 | Tranquilizer |
| Mometasone b | 0.38 | Anti-allergic Agents, Anti-inflammatory Agents |
| Lomustine b | 0.37 | Antineoplastic Agents, Antineoplastic Agents, Alkylating |
| Withaferin A b | 0.36 | Calcium Channel Blockers, Dihydropyridines, Tocolytic Agents, Vasodilator Agents |
| Nifedipine b | 0.36 | Vasodilator Agents |
| Diethylstilbestrol b | 0.36 | Antineoplastic Agents, Hormonal, Carcinogens, Estrogens, Non-steroidal |
| Chlorzoxazone b | 0.36 | Muscle Relaxants, Central |
3. Experimental Section
3.1. Data Preprocessing
3.2. Biclustering Analysis
3.3. Density Functional Theory Calculation
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Ji, H.F.; Li, X.J.; Zhang, H.Y. Natural products and drug discovery. EMBO Rep. 2009, 10, 194–200. [Google Scholar] [CrossRef] [PubMed]
- Quan, Y.; Wang, Z.Y.; Xiong, M.; Xiao, Z.T.; Zhang, H.Y. Dissecting traditional Chinese medicines by omics and bioinformatics. Nat. Prod. Commun. 2014, 9, 1391–1396. [Google Scholar]
- Cheng, H.M.; Li, C.C.; Chen, C.Y.; Lo, H.Y.; Cheng, W.Y.; Lee, C.H.; Yang, S.Z.; Wu, S.L.; Hsiang, C.Y.; Ho, T.Y. Application of bioactivity database of Chinese herbal medicine on the therapeutic prediction, drug development, and safety evaluation. J. Ethnopharmacol. 2010, 132, 429–437. [Google Scholar] [CrossRef] [PubMed]
- Wen, Z.; Wang, Z.; Wang, S.; Ravula, R.; Yang, L.; Xu, J.; Wang, C.; Zuo, Z.; Chow, M.S.; Shi, L.; et al. Discovery of molecular mechanisms of traditional Chinese medicinal formula Si-Wu-Tang using gene expression microarray and connectivity map. PLoS One 2011, 6, e18278. [Google Scholar]
- Lin, W.C.; Tan, T.W. The role of gastric muscle relaxation in cytoprotection induced by San-Huang-Xie-Xin-Tang in rats. J. Ethnopharmacol. 1994, 44, 171–179. [Google Scholar] [CrossRef] [PubMed]
- Cheng, W.Y.; Wu, S.L.; Hsiang, C.Y.; Li, C.C.; Lai, T.Y.; Lo, H.Y.; Shen, W.S.; Lee, C.H.; Chen, J.C.; Wu, H.C.; et al. Relationship between San-Huang-Xie-Xin-Tang and its herbal components on the gene expression profiles in HepG2 cells. Am. J. Chin. Med. 2008, 36, 783–797. [Google Scholar]
- Suthram, S.; Dudley, J.T.; Chiang, A.P.; Chen, R.; Hastie, T.J.; Butte, A.J. Network-based elucidation of human disease similarities reveals common functional modules enriched for pluripotent drug targets. PLoS Comput. Biol. 2010, 6, e1000662. [Google Scholar] [CrossRef] [PubMed]
- Iskar, M.; Zeller, G.; Blattmann, P.; Campillos, M.; Kuhn, M.; Kaminska, K.H.; Runz, H.; Gavin, A.C.; Pepperkok, R.; van Noort, V.; et al. Characterization of drug-induced transcriptional modules: Towards drug repositioning and functional understanding. Mol. Syst. Biol. 2013, 9, 662. [Google Scholar]
- Xiong, M.; Li, B.; Zhu, Q.; Wang, Y.X.; Zhang, H.Y. Identification of transcription factors for drug-associated gene modules and biomedical implications. Bioinformatics 2013, 30, 305–309. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Xiong, M.; Zhang, H.Y. Elucidating polypharmacological mechanisms of polyphenols by gene module profile analysis. Int. J. Mol. Sci. 2014, 15, 11245–11254. [Google Scholar] [CrossRef] [PubMed]
- Prasad, S.; Gupta, S.C.; Tyagi, A.K.; Aggarwal, B.B. Curcumin, a component of golden spice: From bedside to bench and back. Biotechnol. Adv. 2014, 32, 1053–1064. [Google Scholar] [CrossRef] [PubMed]
- Strimpakos, A.S.; Sharma, R.A. Curcumin: Preventive and therapeutic properties in laboratory studies and clinical trials. Antioxid. Redox Signal. 2008, 10, 511–545. [Google Scholar] [CrossRef] [PubMed]
- Yeh, L.L.; Liu, J.Y.; Lin, K.S.; Liu, Y.S.; Chiou, J.M.; Liang, K.Y.; Tsai, T.F.; Wang, L.H.; Chen, C.T.; Huang, C.Y. A randomised placebo-controlled trial of a traditional Chinese herbal formula in the treatment of primary dysmenorrhoea. PLoS One 2007, 2, e719. [Google Scholar] [CrossRef] [PubMed]
- Meja, K.K.; Rajendrasozhan, S.; Adenuga, D.; Biswas, S.K.; Sundar, I.K.; Spooner, G.; Marwick, J.A.; Chakravarty, P.; Fletcher, D.; Whittaker, P.; et al. Curcumin restores corticosteroid function in monocytes exposed to oxidants by maintaining HDAC2. Am. J. Respir. Cell Mol. Biol. 2008, 39, 312–323. [Google Scholar]
- Lamb, J.; Crawford, E.D.; Peck, D.; Modell, J.W.; Blat, I.C.; Wrobel, M.J.; Lerner, J.; Brunet, J.P.; Subramanian, A.; Ross, K.N.; et al. The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science 2006, 313, 1929–1935. [Google Scholar]
- Hochreiter, S.; Bodenhofer, U.; Heusel, M.; Mayr, A.; Mitterecker, A.; Kasim, A.; Khamiakova, T.; van Sanden, S.; Lin, D.; Talloen, W.; et al. FABIA: Factor analysis for bicluster acquisition. Bioinformatics 2010, 26, 1520–1527. [Google Scholar]
- Huang, da, W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [PubMed]
- Kaidery, N.A.; Banerjee, R.; Yang, L.; Smirnova, N.A.; Hushpulian, D.M.; Liby, K.T.; Williams, C.R.; Yamamoto, M.; Kensler, T.W.; Ratan, R.R.; et al. Targeting Nrf2-mediated gene transcription by extremely potent synthetic triterpenoids attenuate dopaminergic neurotoxicity in the MPTP mouse model of Parkinson’s disease. Antioxid. Redox Signal. 2013, 18, 139–157. [Google Scholar]
- Kensler, T.W.; Wakabayashi, N.; Biswal, S. Cell survival responses to environmental stresses via the Keap1-Nrf2-ARE pathway. Annu. Rev. Pharmacol. Toxicol. 2006, 47, 89–116. [Google Scholar] [CrossRef]
- Sun, Y.M.; Xiao, Z.T.; Zhang, H.Y. Structure-activity relationships of tanshinones in activating Nrf2. A DFT study and implications for multifunctional antioxidant discovery. Nat. Prod. Commun. 2014, 9, 453–454. [Google Scholar]
- Abiko, Y.; Miura, T.; Phuc, B.H.; Shinkai, Y.; Kumagai, Y. Participation of covalent modification of Keap1 in the activation of Nrf2 by tert-butylbenzoquinone, an electrophilic metabolite of butylated hydroxyanisole. Toxicol. Appl. Pharmacol. 2011, 255, 32–39. [Google Scholar] [CrossRef] [PubMed]
- Tao, S.; Zheng, Y.; Lau, A.; Jaramillo, M.C.; Chau, B.T.; Lantz, R.C.; Wong, P.K.; Wondrak, G.T.; Zhang, D.D. Tanshinone I activates the Nrf2-dependent antioxidant response and protects against As(III)-induced lung inflammation in vitro and in vivo. Antioxid. Redox Signal. 2013, 19, 1647–1661. [Google Scholar] [CrossRef] [PubMed]
- Sharma, R.A.; Euden, S.A.; Platton, S.L.; Cooke, D.N.; Shafayat, A.; Hewitt, H.R.; Marczylo, T.H.; Morgan, B.; Hemingway, D.; Plummer, S.M.; et al. Phase I clinical trial of oral curcumin: Biomarkers of systemic activity and compliance. Clin. Cancer Res. 2004, 10, 6847–6854. [Google Scholar]
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. B. 1995, 57, 289–300. [Google Scholar]
- Wishart, D.S. DrugBank and its relevance to pharmacogenomics. Pharmacogenomics 2008, 9, 1155–1162. [Google Scholar] [CrossRef] [PubMed]
- Seiler, K.P.; George, G.A.; Happ, M.P.; Bodycombe, N.E.; Carrinski, H.A.; Norton, S.; Brudz, S.; Sullivan, J.P.; Muhlich, J.; Serrano, M.; et al. ChemBank: A small-molecule screening and cheminformatics resource database. Nucleic Acids Res. 2008, 36, D351–D319. [Google Scholar]
- Liang, Q.D.; Gao, Y.; Tan, H.L.; Guo, P.; Li, Y.F.; Zhou, Z.; Tan, W.; Ma, Z.C.; Ma, B.P.; Wang, S.Q. Effects of four Si-Wu-Tang’s constituents and their combination on irradiated mice. Biol. Pharm. Bull. 2006, 29, 1378–1382. [Google Scholar] [CrossRef] [PubMed]
- NCBI. Available online: http://www.ncbi.nlm.nih.gov/ (accessed on 1 May 2013).
- Lee, C.; Yang, W.; Parr, R.G. Development of the Colle-Salvetti correlation energy formula into a functional of the electron density. Phys. Rev. B Condens. Matter 1988, 37, 785–789. [Google Scholar] [CrossRef] [PubMed]
- Miertuš, S.; Scrocco, E.; Tomasi, J. Electrostatic interaction of a solute with a continuum. A direct utilization of ab initio molecular potentials for the prevision of solvent effects. Chem. Phys. 1981, 55, 117–129. [Google Scholar]
- Scott, A.P.; Radom, L. Harmonic vibrational frequencies: an evaluation of Hartree Fock, Møller-Plesset, quadratic configuration interaction, density functional theory and semiempirical scale factors. J. Phys. Chem. 1996, 100, 16502–16513. [Google Scholar] [CrossRef]
- Rienstra-Kiracofe, J.C.; Tschumper, G.S.; Schaefer, H.F. Atomic and molecular electron affinities: Photoelectron experiments and theoretical computations. Chem. Rev. 2002, 102, 231–282. [Google Scholar] [CrossRef] [PubMed]
- Frisch, M.J.; Trucks, G.W.; Schlegel, H.B.; Scuseria, G.E.; Robb, M.A.; Cheeseman, J.R.; Zakrzewski, V.G.; Montgomery, J.A., Jr.; Stratmann, R.E.; Burant, J.C.; et al. GAUSSIAN 03, Revision B.04; Gaussian Inc.: Wallingford, CT, USA, 2004.
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Quan, Y.; Li, B.; Sun, Y.-M.; Zhang, H.-Y. Elucidating Pharmacological Mechanisms of Natural Medicines by Biclustering Analysis of the Gene Expression Profile: A Case Study on Curcumin and Si-Wu-Tang. Int. J. Mol. Sci. 2015, 16, 510-520. https://doi.org/10.3390/ijms16010510
Quan Y, Li B, Sun Y-M, Zhang H-Y. Elucidating Pharmacological Mechanisms of Natural Medicines by Biclustering Analysis of the Gene Expression Profile: A Case Study on Curcumin and Si-Wu-Tang. International Journal of Molecular Sciences. 2015; 16(1):510-520. https://doi.org/10.3390/ijms16010510
Chicago/Turabian StyleQuan, Yuan, Bin Li, You-Min Sun, and Hong-Yu Zhang. 2015. "Elucidating Pharmacological Mechanisms of Natural Medicines by Biclustering Analysis of the Gene Expression Profile: A Case Study on Curcumin and Si-Wu-Tang" International Journal of Molecular Sciences 16, no. 1: 510-520. https://doi.org/10.3390/ijms16010510





