The Role of Circulating MicroRNA-126 (miR-126): A Novel Biomarker for Screening Prediabetes and Newly Diagnosed Type 2 Diabetes Mellitus
Abstract
:1. Introduction
2. Results and Discussion
2.1. Detection of Cel-miR-39 as a Negative Control
2.2. Baseline Characteristics and Serum miR-126 vs. Age/Gender
| Characteristics | Healthy Controls (138 *) | IGT/IFG (157) | DM (160) | P1 | P2 | P3 | ||
|---|---|---|---|---|---|---|---|---|
| Before Treatment | After Treatment | Before Treatment | After Treatment | |||||
| Sex (M:F) | 67/71 | 82/75 | 78/82 | 0.793 | ||||
| Age (year) | 46.7 ± 7.2 | 47.9 ± 7.8 | 50.2 ± 6.7 | 0.985 | ||||
| BMI (kg/m2) | 22.87 ± 0.32 | 23.10 ± 0.26 | 23.32 ± 0.31 | 0.314 | ||||
| Glu (mmol/L) | 4.54 ± 0.66 | 5.13 ± 0.62 | 4.64 ± 0.57 | 11.20 ± 2.65 | 8.14 ± 2.42 | <0.001 | <0.05 | <0.05 |
| HbAlc (%) | 4.69 ± 0.57 | 6.25 ± 0.67 | 5.62 ± 0.45 | 9.16 ± 1.64 | 7.02 ± 1.84 | <0.001 | <0.05 | <0.05 |
| CH | 3.87 ± 0.74 | 4.12 ± 0.86 | 4.02 ± 0.54 | 4.99 ± 1.52 | 4.68 ± 1.48 | 0.253 | 0.462 | 0.543 |
| TG | 1.43 ± 0.56 | 1.51 ± 0.43 | 1.48 ± 0.37 | 2.43 ± 3.3 | 2.23 ± 2.4 | 0.461 | 0.532 | 0.732 |
| HDL | 1.54 ± 0.22 | 1.53 ± 0.18 | 1.54 ± 0.16 | 1.49 ± 0.16 | 1.47 ± 0.20 | 0.375 | 0.422 | 0.653 |
| LDL | 2.35 ± 0.52 | 2.51 ± 0.47 | 2.50 ± 0.32 | 2.46 ± 0.38 | 2.44 ± 0.41 | 0.422 | 0.674 | 0.782 |
| Lp (a) | 14.53 ± 8.48 | 17.86 ± 7.52 | 16.64 ± 7.36 | 16.54 ± 7.23 | 17.04 ± 8.54 | 0.324 | 0.486 | 0.529 |
| miR-126 [lg 2(50–Ct)] | 8.58 ± 0.62 | 7.70 ± 0.41 | 7.99 ± 0.12 | 6.17 ± 0.57 | 6.63 ± 0.62 | <0.05 | 0.036 | 0.001 |
| Variables | Control | IGT/IFG | DM Patients | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Number | miR-126 Median (95% CI) | p Value | Number | miR-126 Median | p Value | Number | miR-126 Median | p Value | |
| Age (years) | |||||||||
| ≤50 | 68 | 8.79 (8.33–9.02) | 0.565 | 72 | 7.62 (7.36−7.94) | 0.209 | 78 | 6.32 (5.96−6.45) | 0.897 |
| >50 | 70 | 8.62 (8.40–8.95) | – | 85 | 7.96 (7.66−8.10) | – | 82 | 6.18 (5.78−6.51) | – |
| Gender | |||||||||
| Male | 73 | 8.60 (8.41–8.88) | 0.565 | 79 | 7.96 (7.62−8.09) | 0.337 | 77 | 7.81 (8.37−8.04) | 0.553 |
| Female | 65 | 8.74 (8.33–8.96) | – | 78 | 7.66 (7.41–7.98) | – | 83 | 6.33 (5.86–6.46) | – |
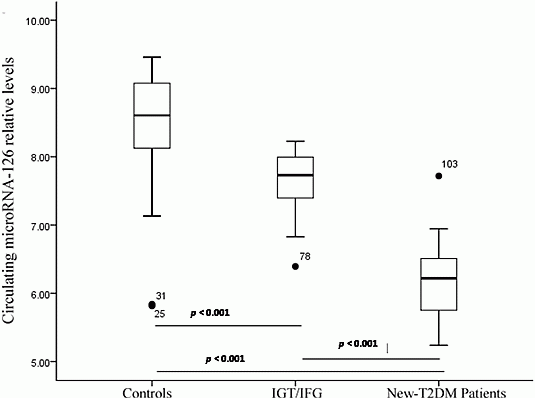
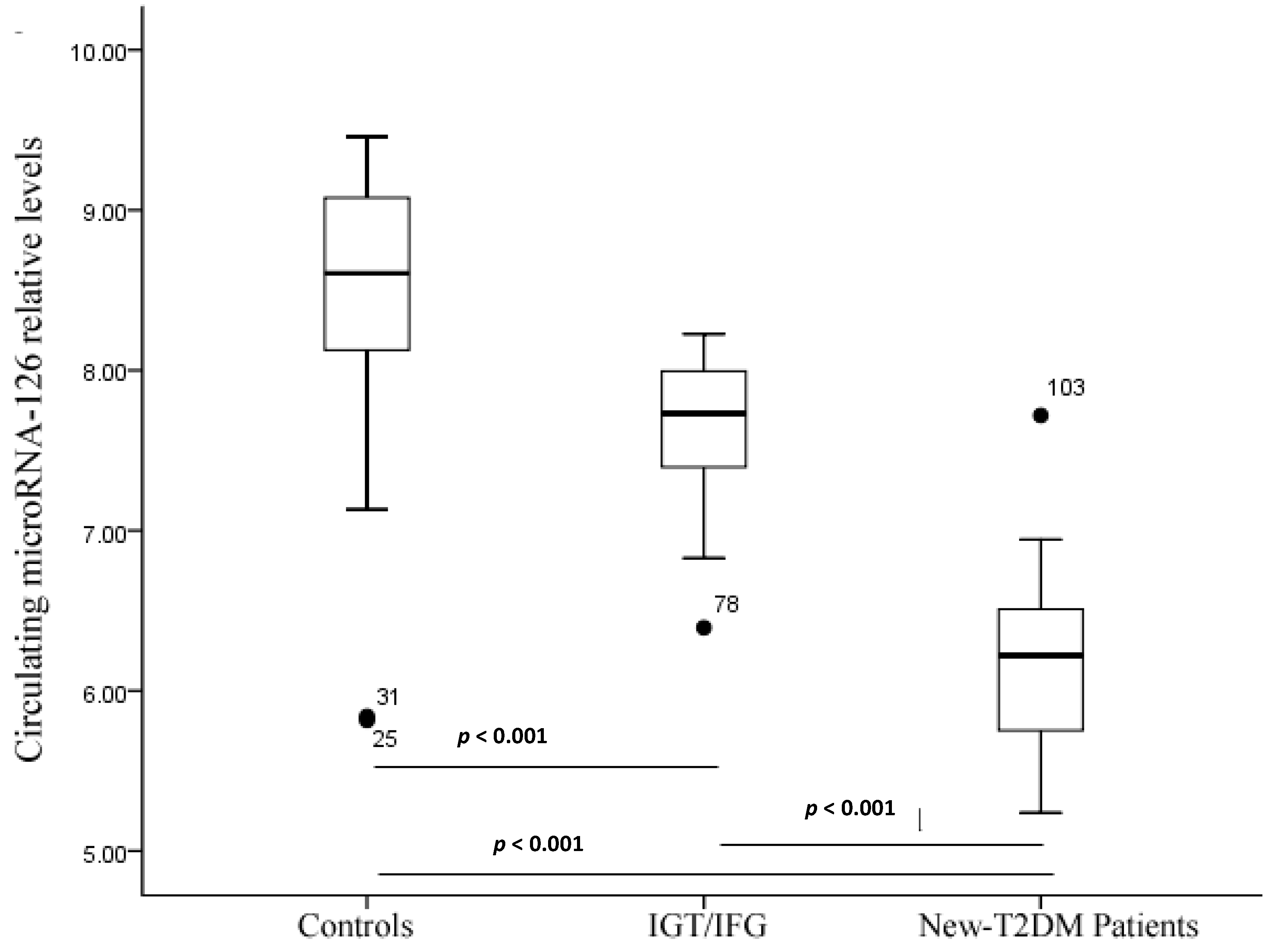
2.3. Serum miR-126 across Disease Conditions
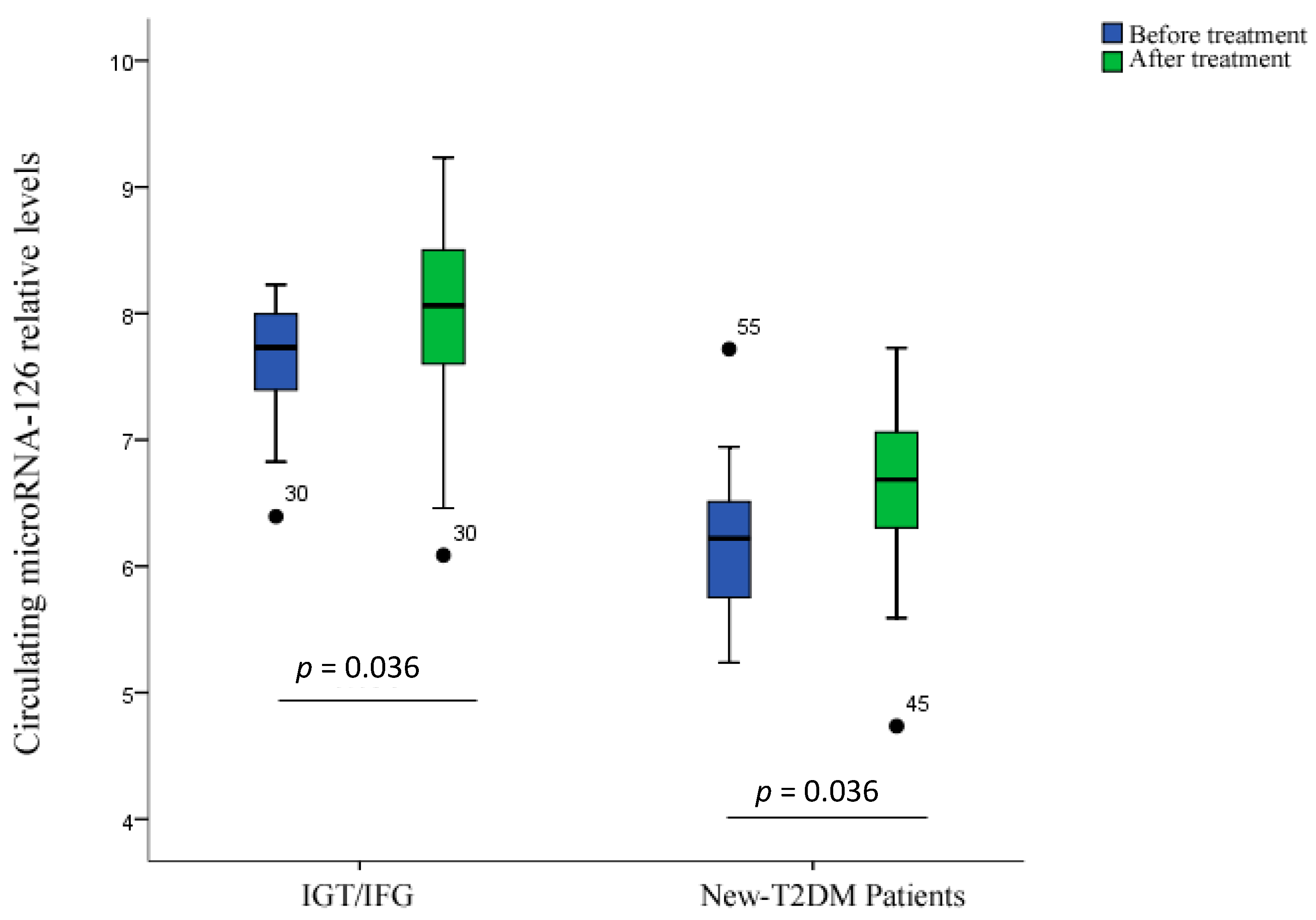
2.4. Treatment Response
2.5. Correlation Analysis
| Variable | Tertiles of Serum miR-126 Levels | p-Trend | ||
|---|---|---|---|---|
| 1 (Lowest) | 2 | 3 (Highest) | ||
| MiR-126 levels | ≤8.32 | 8.32–8.82 | ≥8.82 | |
| New-T2DM patients/Controls, n/n | 84/46 | 52/46 | 24/46 | <0.05 |
| Crude OR (95% CI) | 3.500 (1.901–6.445) | 1.615 (0.946–2.759) | 1 | <0.05 |
| Adjusted OR (95% CI), Model | 3.825 (2.336–8.547) | 1.834 (0.952–3.014) | 1 | <0.05 |
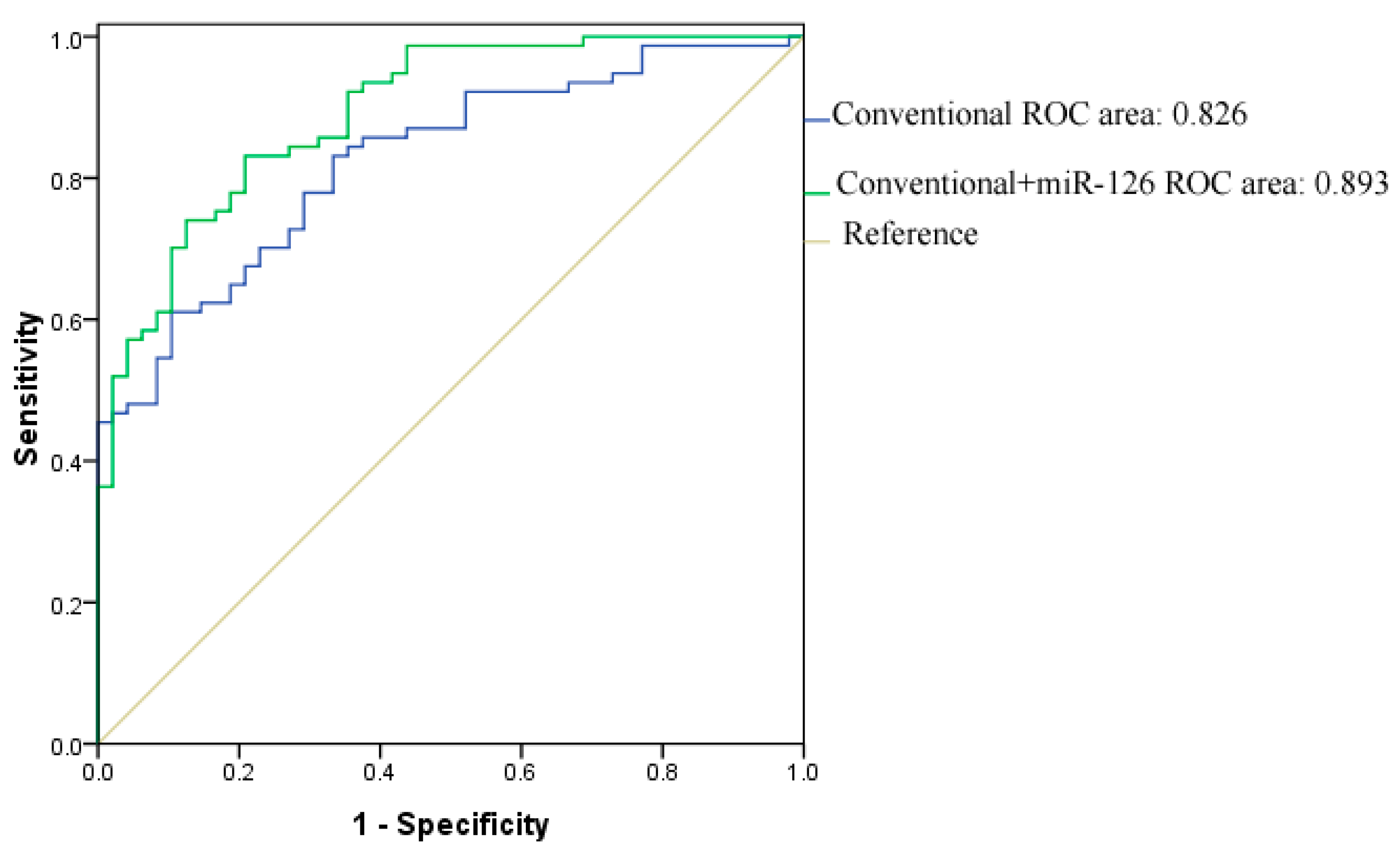
3. Experimental Section
3.1. Clinical Samples
3.2. Diagnosis
3.3. Reverse Transcription-Quantitative Polymerase Chain Reaction (RT-qPCR)
3.4. Statistical Analysis
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Ambros, V. MicroRNAs: Tiny regulators with great potential. Cell 2001, 107, 823–826. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Target recognition and regulatory functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef]
- Ambros, V. The functions of animal microRNAs. Nature 2004, 431, 350–355. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Sempere, L.F.; Freemantle, S. Expression profiling of mammalian microRNAs uncovers a subset of brain-expressed microRNAs with possible roles in murine and human neuronal differentiation. Genome Biol. 2004, 5, R13. [Google Scholar] [CrossRef]
- Mitchell, P.S.; Parkin, R.K. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA 2008, 105, 10513–10518. [Google Scholar] [CrossRef]
- Zubakov, D.; Boersma, A.W.; Choi, Y.; van Kuijk, P.F.; Wiemer, E.A.; Kayser, M. MicroRNA markers for forensic body fluid identification obtained from microarray screening and quantitative RT-PCR confirmation. Int. J. Leg. Med. 2010, 124, 217–226. [Google Scholar] [CrossRef]
- Park, N.J.; Zhou, H.; Elashoff, D.; Henson, B.S.; Kastratovic, D.A.; Abemayor, E.; Wong, D.T. Salivary microRNA: Discovery, characterization, and clinical utility for oral cancer detection. Clin. Cancer 2009, 15, 5473–5477. [Google Scholar] [CrossRef]
- Jeyaseelan, K.; Lim, K.Y. MicroRNA expression in the blood and brain of rats subjected to transient focal ischemia by middle cerebral artery occlusion. Stroke 2008, 39, 959–966. [Google Scholar]
- Taylor, D.D.; Gercel-Taylor, C. MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol. Oncol. 2008, 110, 13–21. [Google Scholar] [CrossRef]
- Tanaka, M.; Oikawa, K. Down-regulation of miR-92 in human plasma is a novel marker for acute leukemia patients. PLoS One 2009, 4, e5532. [Google Scholar] [CrossRef]
- Laterza, O.F.; Lim, L. Plasma microRNAs as sensitive and specific biomarkers of tissue injury. Clin. Chem. 2009, 55, 1977–1983. [Google Scholar] [CrossRef]
- Wang, K.; Zhang, S. Circulating microRNAs, potential biomarkers for drug-induced liver injury. Proc. Natl. Acad. Sci. USA 2009, 106, 4402–4407. [Google Scholar] [CrossRef]
- Ji, X.; Takahashi, R. Plasma miR-208 as a biomarker of myocardial injury. Clin. Chem. 2009, 55, 1944–1949. [Google Scholar] [CrossRef]
- Wang, G.K.; Zhu, J.Q. Circulating microRNA: A novel potential biomarker for early diagnosis of acute myocardial infarction in humans. Eur. Heart J. 2010, 31, 659–666. [Google Scholar] [CrossRef]
- Fichtlscherer, S.; de Rosa, S. Circulating microRNAs in patients with coronary artery disease. Circ. Res. 2010, 107, 677–684. [Google Scholar] [CrossRef]
- Tijsen, A.J.; Creemers, E.E. MiR423-5p as a circulating biomarker for heart failure. Circ. Res. 2010, 106, 1035–1039. [Google Scholar] [CrossRef]
- Igaz, I.; Szőnyi, M.; Varga, P.; Topa, L. Potential relevance of microRNAs in the diagnostics of inflammatory bowel diseases. Orv. Hetil. 2014, 155, 487–491. [Google Scholar] [CrossRef]
- Küçükali, C.I.; Kürtüncü, M.; Coban, A.; Cebi, M.; Tüzün, E. Epigenetics of multiple sclerosis: An updated review. Neuromol. Med. 2014, in press. [Google Scholar]
- Nathan, D.M. Long-term complications of diabetes mellitus. N. Engl. J. Med. 1993, 328, 1676–1685. [Google Scholar] [CrossRef]
- Frankel, D.S.; Meigs, J.B. Von Willebrand factor, type 2 diabetes mellitus, and risk of cardiovascular disease: The Framingham offspring study. Circulation 2008, 118, 2533–2539. [Google Scholar] [CrossRef]
- Van den Oever, I.A. Endothelial dysfunction, infiammation, and apoptosis in diabetes mellitus. Mediat. Inflamm. 2010, 2010, 792393. [Google Scholar]
- Zampetaki, A.; Kiechl, S. Plasma microRNA profiling reveals loss of endothelial miR-126 and other microRNAs in type 2 diabetes. Circ. Res. 2010, 107, 810–817. [Google Scholar] [CrossRef]
- Fish, J.E.; Santoro, M.M. miR-126 regulates angiogenic signaling and vascular integrity. Dev. Cell 2008, 15, 272–284. [Google Scholar] [CrossRef]
- Wang, S.; Aurora, A.B. The endothelial-specific microRNA miR-126 governs vascular integrity and angiogenesis. Dev. Cell 2008, 15, 261–271. [Google Scholar]
- Zernecke, A.; Bidzhekov, K. Delivery of microRNA-126 by apoptotic bodies induces CXCL12-dependent vascular protection. Sci. Signal. 2009, 2, ra81. [Google Scholar]
- Fichtlscherer, S.; Zeiher, A.M. Circulating micrornas: Biomarkers or mediators of cardiovascular diseases? Arterioscler. Thromb. Vasc. Biol. 2011, 31, 2383–2390. [Google Scholar] [CrossRef]
- Asaga, S.; Kuo, C. Hoon1 direct serum assay for microRNA-21 concentrations in early and advanced breast cancer. Clin. Chem. 2011, 57, 84–91. [Google Scholar] [CrossRef]
- Murata, K.; Furu, M.; Yoshitomi, H.; Ishikawa, M.; Shibuya, H.; Hashimoto, M.; Imura, Y.; Fujii, T.; Ito, H.; Mimori, T.; et al. Comprehensive microRNA analysis identifies miR-24 and miR-125a-5p as plasma biomarkers for rheumatoid arthritis. PLoS One 2013, 8, e69118. [Google Scholar]
- Gilad, S.; Meiri, E.; Yogev, Y.; Benjamin, S.; Lebanony, D.; Yerushalmi, N.; Benjamin, H.; Kushnir, M.; Cholakh, H.; Melamed, N. Serum microRNAs are promising novel biomarkers. PLoS One 2008, 3, e3148. [Google Scholar] [CrossRef]
- Wang, G.; Chan, E.S.; Kwan, B.C.; Li, P.K.; Yip, S.K.; Szeto, C.C.; Ng, C.F. Expression of microRNAs in the urine of patients with bladder cancer. Clin. Genitourin. Cancer 2012, 10, 106–113. [Google Scholar] [CrossRef]
- Prevention of Diabetes mellitus. Report of a WHO study group. World Health Organ. Tech. Rep. Ser. 1994, 844, 1–100.
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Liu, Y.; Gao, G.; Yang, C.; Zhou, K.; Shen, B.; Liang, H.; Jiang, X. The Role of Circulating MicroRNA-126 (miR-126): A Novel Biomarker for Screening Prediabetes and Newly Diagnosed Type 2 Diabetes Mellitus. Int. J. Mol. Sci. 2014, 15, 10567-10577. https://doi.org/10.3390/ijms150610567
Liu Y, Gao G, Yang C, Zhou K, Shen B, Liang H, Jiang X. The Role of Circulating MicroRNA-126 (miR-126): A Novel Biomarker for Screening Prediabetes and Newly Diagnosed Type 2 Diabetes Mellitus. International Journal of Molecular Sciences. 2014; 15(6):10567-10577. https://doi.org/10.3390/ijms150610567
Chicago/Turabian StyleLiu, Yang, Guangqiang Gao, Chun Yang, Kun Zhou, Baozhong Shen, Hongyan Liang, and Xiaofeng Jiang. 2014. "The Role of Circulating MicroRNA-126 (miR-126): A Novel Biomarker for Screening Prediabetes and Newly Diagnosed Type 2 Diabetes Mellitus" International Journal of Molecular Sciences 15, no. 6: 10567-10577. https://doi.org/10.3390/ijms150610567