Functional Analysis of the Dioxin Response Elements (DREs) of the Murine CYP1A1 Gene Promoter: Beyond the Core DRE Sequence
Abstract
:1. Introduction
2. Results
2.1. DREs Conserved in Mouse, Human and Rat
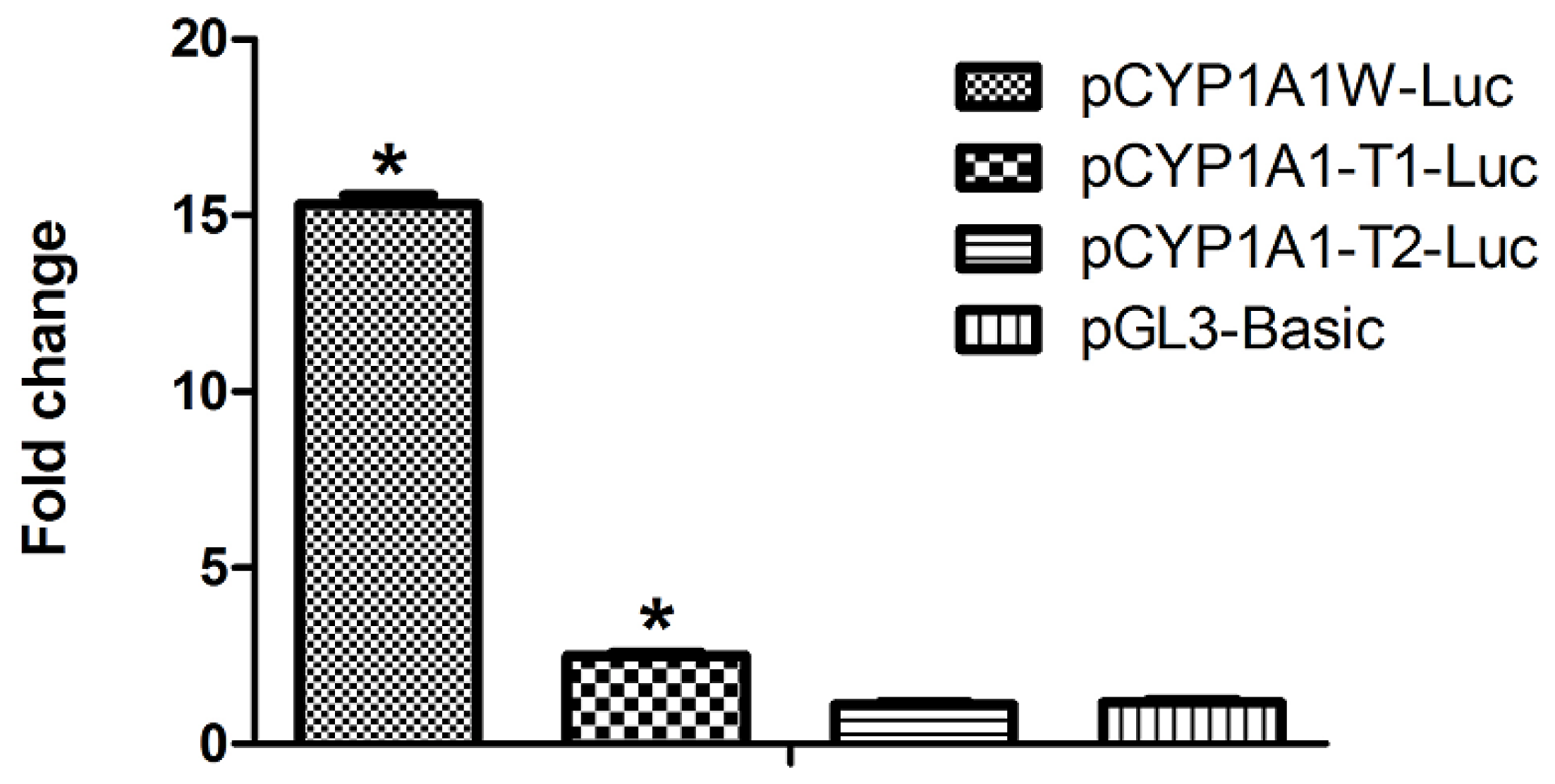
2.2. Single DRE at Position −488 Is Enough to Activate Transcription in Response to Dioxin
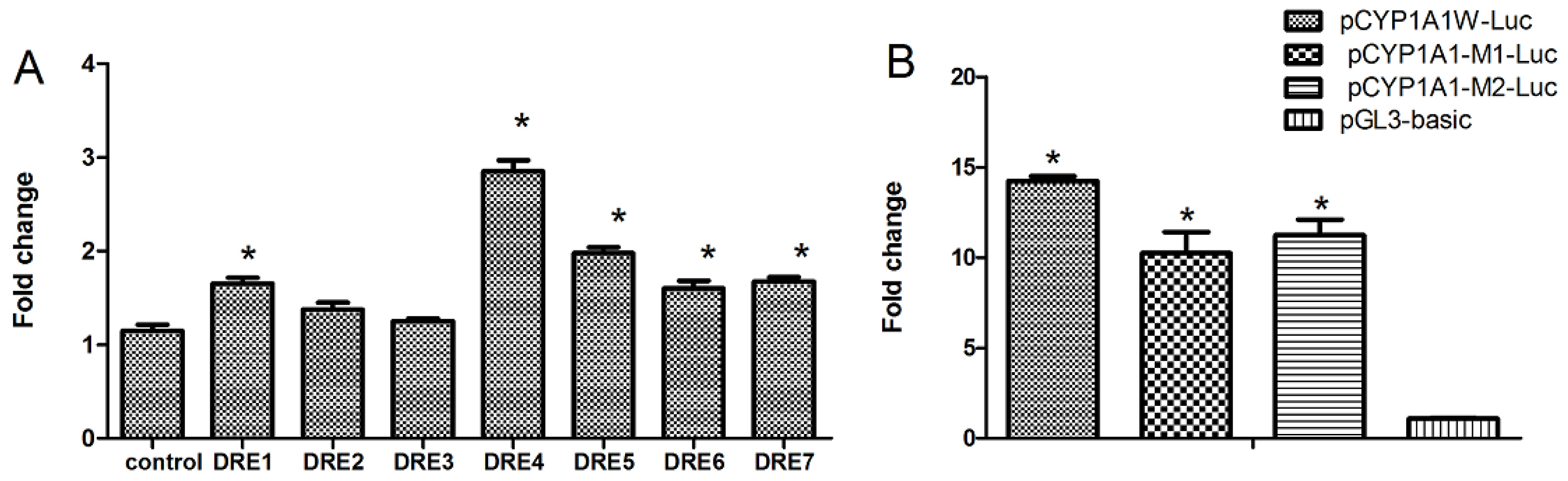
2.3. Seven DREs Have Different Transcriptional Efficiency Induced by TCDD
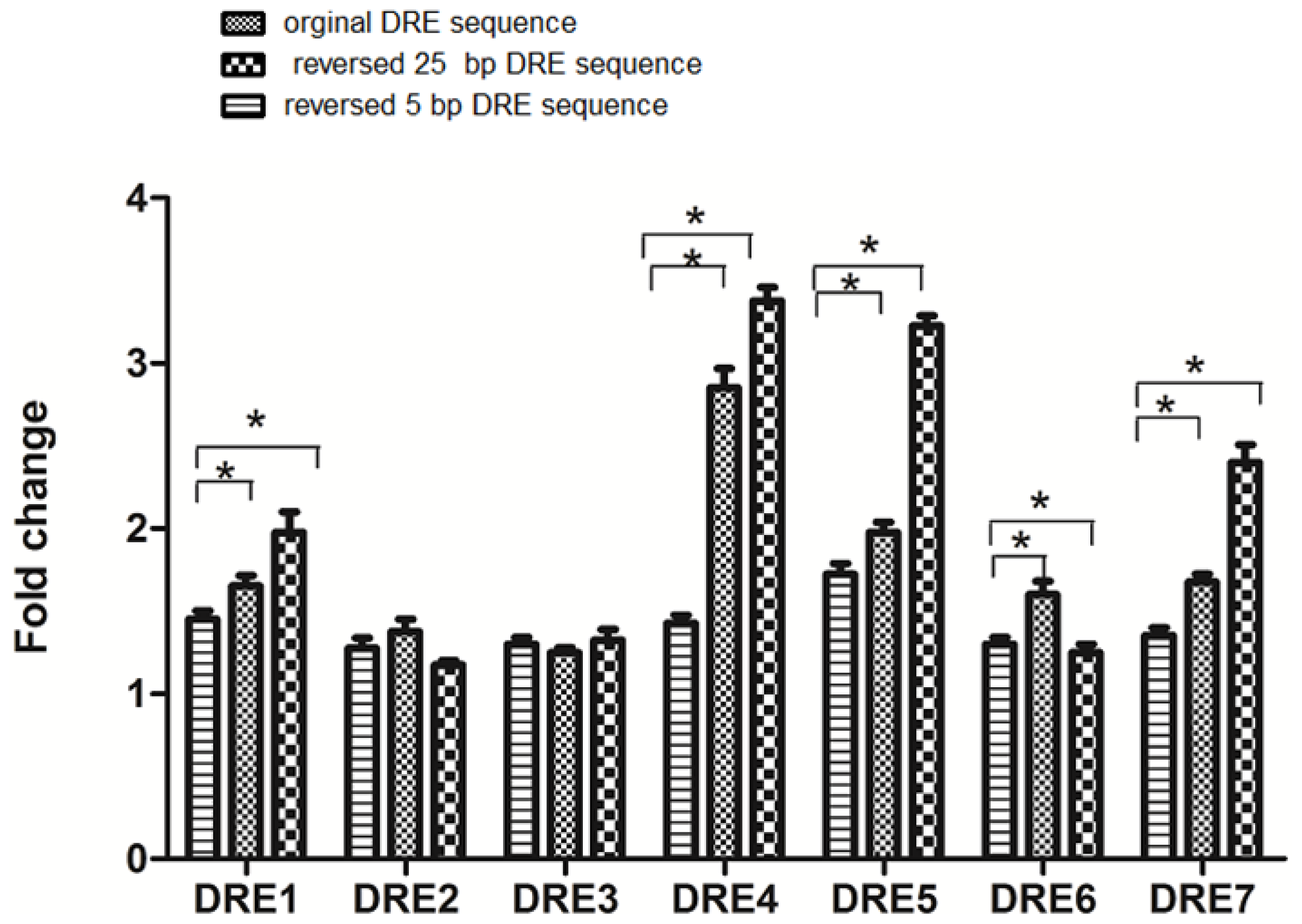
2.4. The DRE Direction and Its Adjacent Sequences Can Affect the Efficiency of Transcription in Response to TCDD
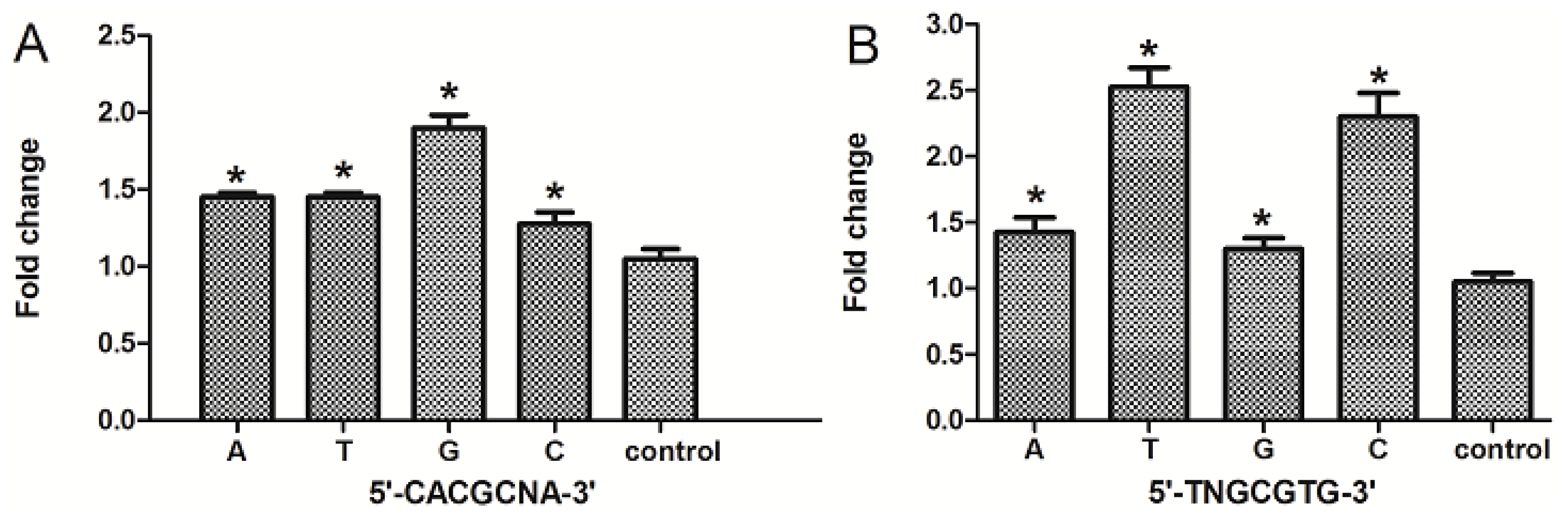
2.5. The Nucleotide “N” in the Core DRE (5′-TNGCGTG-3′) Sequence Influenced the Transcription Triggered by TCDD
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. Cell Culture and Chemical Treatment
4.3. Plasmid Construction
4.4. Transient Transfection
4.5. Luciferase Assay
4.6. Statistics
5. Conclusions
Supplementary Information
ijms-15-06475-s001.pdfAcknowledgments
Conflicts of Interest
- Author ContributionsConceived of and designed the experiments: S.L., X.P., W.Z., Q.X. and B.Z. Performed the experiments: S.L. and X.P. Analyzed the data: S.L., X.P., W.Z., Q.X., and B.Z. Contributed reagents/materials/analysis tools: S.L., X.P., W.Z., Q.X., and B.Z. Wrote the paper: S.L. and B.Z.
References
- Schmidt, J.V.; Su, G.H.; Reddy, J.K.; Simon, M.C.; Bradfield, C.A. Characterization of a murine Ahr null allele: Involvement of the Ah receptor in hepatic growth and development. Proc. Natl. Acad. Sci. USA 1996, 93, 6731–6736. [Google Scholar]
- Hankinson, O. The aryl hydrocarbon receptor complex. Annu. Rev. Pharmacol. Toxicol 1995, 35, 307–340. [Google Scholar]
- Safe, S. Polychlorinated biphenyls (PCBs), dibenzo-p-dioxins (PCDDs), dibenzofurans (PCDFs), and related compounds: Environmental and mechanistic considerations which support the development of toxic equivalency factors (TEFs). Crit. Rev. Toxicol 1990, 21, 51–88. [Google Scholar]
- Whitlock, J.P., Jr. Induction of cytochrome P4501A1. Annu. Rev. Pharmacol. Toxicol 1999, 39, 103–125. [Google Scholar]
- Senft, A.P.; Daltion, T.P.; Nebert, D.W.; Genter, M.B.; Hutchinson, R.J.; Schertzer, H.G. Dioxin increases reactive oxygen production in mouse liver mitochondria. Toxicol. Appl. Pharmacol 2002, 178, 15–21. [Google Scholar]
- Shertzer, H.G.; Nebert, D.W.; Puga, A.; Ary, M.; Sonntag, D.; Dixon, K.; Robinson, L.J.; Cianciolo, E.; Dalton, T.P. Dioxin causes a sustained oxidative stress response in the mouse. Biochem. Biophys. Res. Commun 1998, 253, 44–48. [Google Scholar]
- Phelan, D.; Winter, G.M.; Rogers, W.J.; Lam, J.C.; Denison, M.S. Activation of the Ah receptor signal transduction pathway by bilirubin and biliverdin. Arch. Biochem. Biophys 1998, 357, 155–163. [Google Scholar]
- Denison, M.S.; Pandini, A.; Nagy, S.R.; Baldwin, E.P.; Bonati, L. Ligand binding and activation of the Ah receptor. Chem. Biol. Interact 2002, 141, 3–24. [Google Scholar]
- Machala, M.; Vondrácek, J.; Bláha, L.; Ciganek, M.; Neca, J.V. Aryl hydrocarbon receptor-mediated activity of mutagenic polycyclic aromatic hydrocarbons determined using in vitro reporter gene assay. Mutat. Res 2001, 497, 49–62. [Google Scholar]
- Ma, Q.; Whitlock, J.P., Jr. A novel cytoplasmic protein that interacts with the Ah receptor, contains tetratricopeptide repeat motifs, and augments the transcriptional response to 2,3,7,8-tetrachlorodibenzo-p-dioxin. J. Biol. Chem 1997, 272, 8878–8884. [Google Scholar]
- Meyer, B.K.; Pray-Grant, M.G.; Vanden Heuvel, J.P.; Perdew, G.H. Hepatitis B virus X-associated protein 2 is a subunit of the unliganded aryl hydrocarbon receptor core complex and exhibits transcriptional enhancer activity. Mol. Cell. Biol 1998, 18, 978–988. [Google Scholar]
- Hord, N.G.; Perdew, G.H. Physicochemical and immunocytochemical analysis of the aryl hydrocarbon receptor nuclear translocator: Characterization of two monoclonal antibodies to the aryl hydrocarbon receptor nuclear translocator. Mol. Pharmacol 1994, 46, 618–626. [Google Scholar]
- Pollenz, R.S.; Sattler, C.A.; Poland, A. The aryl hydrocarbon receptor and aryl hydrocarbon receptor nuclear translocator protein show distinct subcellular localizations in Hepa 1c1c7 cells by immunofluorescence microscopy. Mol. Pharmacol 1994, 45, 428–438. [Google Scholar]
- Probst, M.R.; Reisz-Porszasz, S.; Agbunag, R.V.; Ong, M.S.; Hankinson, O. Role of the aryl hydrocarbon receptor nuclear translocator protein in aryl hydrocarbon (dioxin) receptor action. Mol. Pharmacol 1993, 44, 511–518. [Google Scholar]
- Denison, M.S.; Fisher, J.M.; Whitlock, J.P., Jr. Protein–DNA interactions at recognition sites for the dioxin-Ah receptor complex. J. Biol. Chem 1989, 264, 16478–16482. [Google Scholar]
- Dong, L.; Ma, Q.; Whitlock, J.P., Jr. DNA binding by the heterodimeric Ah receptor: Relationship to dioxin-induced CYP1A1 transcription in vivo. J. Biol. Chem. 1996, 271, 7942–8948. [Google Scholar]
- Bank, P.A.; Yao, E.F.; Phelps, C.L.; Harper, P.A.; Denison, M.S. Species–specific binding of transformed Ah receptor to a dioxin responsive transcriptional enhancer. Eur. J. Pharmacol 1992, 228, 85–94. [Google Scholar]
- Ma, Q. Induction of CYP1A1—The AhR/DRE paradigm: Transcription, receptor regulation, and expanding biological roles. Curr. Drug Metab 2001, 2, 149–164. [Google Scholar]
- Nukaya, M.; Bradfield, C.A. Conserved genomic structure of the Cyp1a1 and Cyp1a2 loci and their dioxin responsive elements cluster. Biochem. Pharmacol 2009, 77, 654–659. [Google Scholar]
- Shen, E.S.; Whitlock, J.P., Jr. The potential role of DNA methylation in the response to 2,3,7,8-tetrachlorodibenzo-p-dioxin. J. Biol. Chem 1989, 264, 17754–17758. [Google Scholar]
- Henry, E.C.; Rucci, G.; Gasiewicz, T.A. Characterization of multiple forms of the Ah receptor: Comparison of species and tissues. Biochemistry 1989, 28, 6430–6440. [Google Scholar]
- Israel, D.I.; Whitlock, J.P., Jr. Regulation of cytochrome P1-450 gene transcription by 2,3,7,8-tetrachlorodibenzo-p-dioxin in wild type and variant mouse hepatoma cells. J. Biol. Chem 1984, 259, 5400–5402. [Google Scholar]
- Jones, K.W.; Whitlock, J.P., Jr. Functional analysis of the transcriptional promoter for the CYP1A1 gene. Mol. Cell. Biol 1990, 10, 5098–5105. [Google Scholar]
- Jones, P.B.; Durrin, L.K.; Fisher, J.M.; Whitlock, J.P. Control of gene expression by 2,3,7,8-tetrachlorodibenzo-p-dioxin: Multiple dioxin-responsive domains 5′-ward of the cytochrome P1-450 gene. J. Biol. Chem 1986, 261, 6647–6650. [Google Scholar]
- Jones, P.B.; Durrin, L.K.; Galeazzi, D.R.; Whitlock, J.P., Jr. Control of cytochrome P1-450 gene expression: Analysis of a dioxin-responsive enhancer system. Proc. Natl. Acad. Sci. USA 1986, 83, 2802–2806. [Google Scholar]
- Denison, M.S.; Fisher, J.M.; Whitlock, J.P., Jr. Inducible, receptor-dependent protein–DNA interactions at a dioxin-responsive transcriptional enhancer. Proc. Natl. Acad. Sci. USA 1988, 85, 2528–2532. [Google Scholar]
- Fisher, J.M.; Wu, L.; Denison, M.S.; Whitlock, J.P., Jr. Organization and function of a dioxin-responsive enhancer. J. Biol. Chem 1990, 265, 9676–9681. [Google Scholar]
- Lusska, A.; Shen, E.; Whitlock, J.P., Jr. Protein–DNA interactions at a dioxin-responsive enhancer: Analysis of six bona fide DNA-binding sites for the liganded Ah receptor. J. Biol. Chem 1993, 268, 6575–6580. [Google Scholar]
- Kress, S.; Reichert, J.; Schwarz, M. Functional analysis of the human cytochrome P4501A1 (CYP1A1) gene enhancer. Eur. J. Biochem 1998, 258, 803–812. [Google Scholar]
- Galijatovic, A.; Beaton, D.; Nguyen, N.; Chen, S.; Bonzo, J.; Johnson, R.; Maeda, S.; Karin, M.; Guengerich, F.P.; Tukey, R.H. The human CYP1A1 gene is regulated in a developmental and tissue-specific fashion in transgenic mice. J. Biol. Chem 2004, 279, 23969–23976. [Google Scholar]
- Yao, E.F.; Denison, M.S. DNA sequence determinants for binding of transformed Ah receptor to a dioxin-responsive enhancer. Biochemistry 1992, 31, 5060–5067. [Google Scholar]
- Ko, H.P.; Okino, S.T.; Ma, Q.; Whitlock, J.P., Jr. Dioxin-induced CYP1A1 transcription in vivo: The aromatic hydrocarbon receptor mediates transactivation, enhancer-promoter communication, and changes in chromatin structure. Mol. Cell Biol 1996, 16, 430–436. [Google Scholar]
- Shen, E.S.; Whitlock, J.P., Jr. Protein–DNA interactions at a dioxin-responsive enhancer: Mutational analysis of the DNA-binding site for the liganded Ah receptor. J. Biol. Chem 1992, 267, 6815–6819. [Google Scholar]

© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Li, S.; Pei, X.; Zhang, W.; Xie, H.Q.; Zhao, B. Functional Analysis of the Dioxin Response Elements (DREs) of the Murine CYP1A1 Gene Promoter: Beyond the Core DRE Sequence. Int. J. Mol. Sci. 2014, 15, 6475-6487. https://doi.org/10.3390/ijms15046475
Li S, Pei X, Zhang W, Xie HQ, Zhao B. Functional Analysis of the Dioxin Response Elements (DREs) of the Murine CYP1A1 Gene Promoter: Beyond the Core DRE Sequence. International Journal of Molecular Sciences. 2014; 15(4):6475-6487. https://doi.org/10.3390/ijms15046475
Chicago/Turabian StyleLi, Shuaizhang, Xinhui Pei, Wen Zhang, Heidi Qunhui Xie, and Bin Zhao. 2014. "Functional Analysis of the Dioxin Response Elements (DREs) of the Murine CYP1A1 Gene Promoter: Beyond the Core DRE Sequence" International Journal of Molecular Sciences 15, no. 4: 6475-6487. https://doi.org/10.3390/ijms15046475



