Role of miR-191/425 Cluster in Tumorigenesis and Diagnosis of Gastric Cancer
Abstract
:1. Introduction
2. Results and Discussion
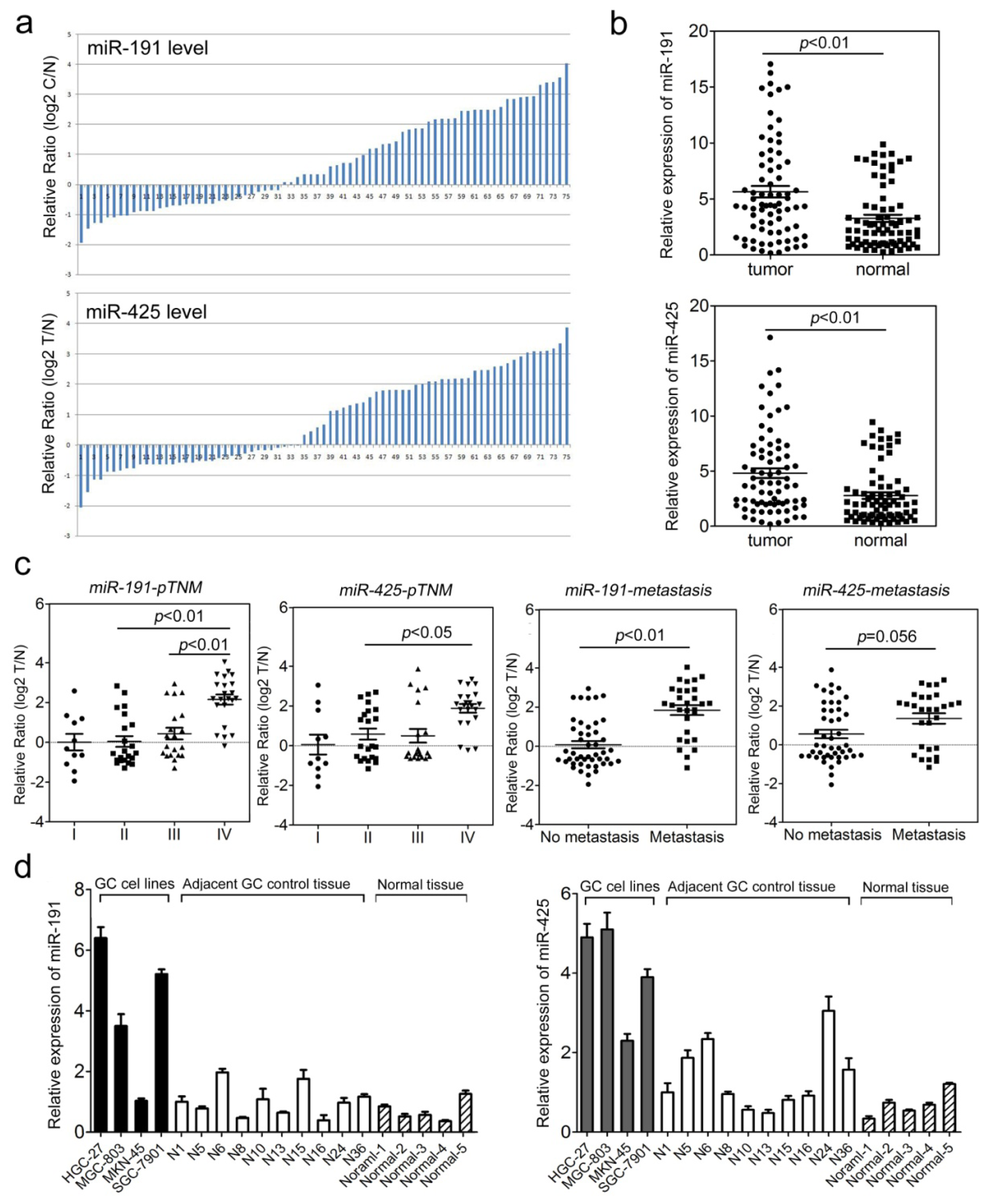
2.1. The miR-191/425 Cluster Is Highly Expressed in Gastric Cancer (GC)
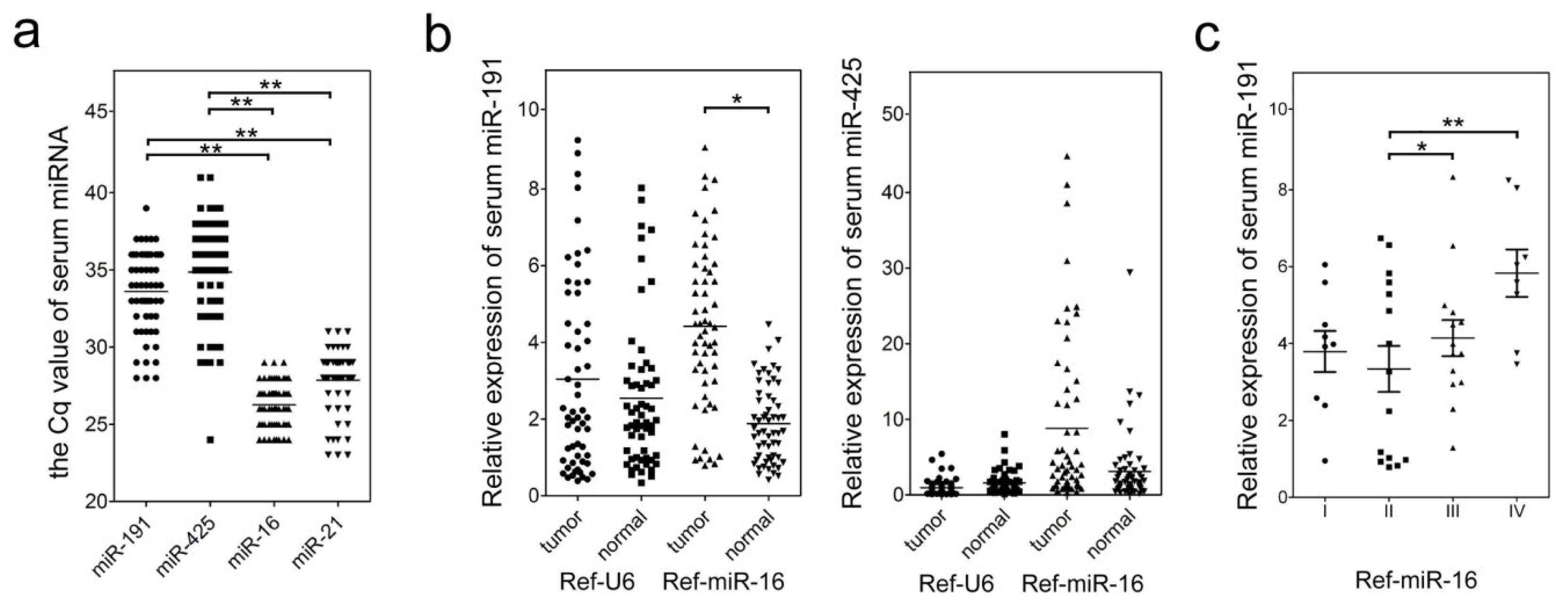
2.2. Clinical and Pathological Characteristics of miR-191/425 Levels in GC Serum
2.3. Evaluation of Serum miR-191/425 as a Potential GC Diagnostic Marker
2.4. Inhibition of miR-191 and miR-425 Repressed Cell Proliferation and Cell Cycle Progression of GC Cells
2.5. Inhibition of miR-191 and miR-1425 in GC Cells Inhibits Cell Migration and Invasion
2.6. Discussion
3. Experimental Section
3.1. Tissue Specimens
3.2. Serum Collection
3.3. Serum RNA Isolation and qRT-PCR
3.4. Cell Cultures and Transfection
3.5. Tissue RNA Isolation and qRT-PCR
3.6. Cell Proliferation and Cell Cycle Assay
3.7. Cell Migration and Invasion Assays
3.8. Statistical Analyses
4. Conclusions
Supplementary Information
| Item to Check | Importance | Check List | Where in the Manuscript; Additional Comment |
|---|---|---|---|
| Experimental Design | |||
| Definition of experimental and control groups | E | Yes | Materials and Methods: GC tissues and matched adjacent normal tissues; and in Table S1. |
| Number within each group | E | Yes | Materials and Methods: (1) 75 pairs of clinic GC tissues and matched adjacent normal tissue samples; (2) 57 GC patients and 58 healthy donor blood samples. |
| Assay carried out by core lab or investigator’s lab? | D | Yes | All assays were performed in investigator’s lab. |
| Acknowledgement of authors’ contributions | D | Yes | All mentioned authors met the authorship as defined by the journal. |
| SAMPLE | |||
| Description | E | Yes | Materials and Methods. |
| Volume/mass of sample processed | D | Not available | |
| Microdissection or macrodissection | E | Yes | Materials and Methods: macrodissection with histolological verification. |
| Processing procedure | E | Yes | Materials and Methods. |
| If frozen—how and how quickly?/If fixed—with what, how quickly? | E | Yes | Materials and Methods: all samples were immediately snap-frozen in liquid nitrogen, and stored at −196 °C in liquid nitrogen until RNA extraction. |
| Nucleic Acid Extraction | |||
| Procedure and/or instrumentation | E | Yes | Materials and Methods. |
| Details of DNase or RNAse treatment | E | Yes | No treatment. |
| Contamination assessment (DNA or RNA) | E | Yes | According to Chen et al. [33] miRNA measurements by the TaqMan assays are not affected by genomic DNA. |
| Nucleic acid quantification | E | Yes | The RNA was measured by Thermo Scientific™ Nanodrop™ 2000. |
| Reverse Transcription | |||
| Complete reaction conditions | E | Yes | Materials and Methods. |
| Cqs with and without RT | D | Yes | There were no Cqs (<40) in reactions without RT. |
| Storage conditions of cDNA | D | Yes | The cDNA was stored at −20 °C. |
| qPCR Target Information | Yes | ||
| Sequence accession number | E | Yes | Materials and Methods: Predesigned TaqMan MicroRNA Assays including primer set and TaqMan probe were purchased from Applied Biosystems. The miRBase Accession Numbers for miR-191 (MIMAT0000440) and miR-425 (MIMAT0003393). |
| Location of amplicon/Amplicon length/In silico specificity screen (BLAST, etc.)/Pseudogenes, retropseudogenes or other homologs?/Secondary structure analysis of amplicon | E | Yes | Materials and Methods: Use of miRNA specific TaqMan assays; specificity guaranteed by the manufacturer. |
| qPCR Oligonucleotides | |||
| Primer sequences/Probe sequences/Location and identity of any modifications | E | Yes | The manufacturer does not provide this information for miRNAs. |
| Manufacturer of oligonucleotides | D | Yes | Applied Biosystems as part of Life Technologies. |
| Purification method | D | Yes | Applied Biosystems does not provide information. |
| qPCR Protocol | |||
| Complete reaction conditions | E | Yes | Materials and Methods. |
| Reaction volume and amount of cDNA/DNA/Primer, (probe), Mg++ and dNTP concentrations/Polymerase identity and concentration/Buffer/kit identity and manufacturer | E | Yes | Use of miRNA specific TaqMan assays; specificity guaranteed by the manufacturer. |
| Exact chemical constitution of the buffer | D | Yes | The manufacturer does not provide this information. |
| Additives (SYBR Green I, DMSO, etc.) | E | Yes | No additional additives. |
| Manufacturer of plates/tubes and catalog number | D | Yes | The MicroAmp® Optical 384-Well Reaction Plates (4343370) were purchased from Applied Biosystems. |
| Complete thermocycling parameters | E | Yes | Use of miRNA specific TaqMan assays; specificity guaranteed by the manufacturer. |
| Reaction setup (manual/robotic) | D | Yes | Manual setup. |
| Manufacturer of qPCR instrument | E | Yes | 7900HT Fast Real-Time PCR system (Applied Biosystems). |
| qPCR Validation | |||
| Evidence of optimisation (from gradients) | D | Not available | |
| Specificity (gel, sequence, melt, or digest) | E | Yes | Specificity guaranteed by the manufacturer of the TaqMan assays. |
| Calibration curves with slope and y-intercept | E | Not available | |
| DATA ANALYSIS | |||
| qPCR analysis program (source, version) | E | Yes | Materials and Methods: Relative expression of the target miRNAs was calculated using the ΔΔCq method. |
| Results of NTCs | E | Yes | NTC did not result in any amplification (data not shown). |
| Justification of number and choice of reference genes | E | Yes | Materials and Methods: U6snRNA and miR-16 were used as the endogenous control for data normalization, respectively. |
| Description of normalisation method | E | Yes | Materials and Methods. |
| Number and stage (RT or qPCR) of technical replicates | E | Yes | Materials and Methods: All PCR reactions including no-template controls were run in triplicate. |
| Repeatability (intra-assay and inter-assay variation, %CV variation) | E | Yes | Materials and Methods. |
| Statistical methods for result significance | E | Yes | Materials and Methods. |
| Software (source, version) | E | Yes | Materials and Methods. |
| Cq or raw data submission using RDML | D | Not available | |
Acknowledgments
Conflicts of Interest
- Author ContributionsJ.Y. and Z-B.L. conceived the project, W-Z.P. and R.M. collected the tissue and serum samples, performed qPCR and statistical analyses, F.W. performed cell experiments.
References
- Bertuccio, P.; Chatenoud, L.; Levi, F.; Praud, D.; Ferlay, J.; Negri, E.; Malvezzi, M.; la Vecchia, C. Recent patterns in gastric cancer: A global overview. Int. J. Cancer 2005, 125, 666–673. [Google Scholar]
- Jemal, A.; Bray, F.; Center, M.M.; Ferlay, J.; Ward, E.; Forman, D. Global cancer statistics. CA Cancer J. Clin. 2011, 61, 69–90. [Google Scholar]
- Melton, S.D.; Genta, R.M.; Souza, R.F. Biomarkers and molecular diagnosis of gastrointestinal and pancreatic neoplasms. Nat. Rev. Gastroenterol. Hepatol. 2010, 7, 620–628. [Google Scholar]
- Sethi, S.; Ali, S.; Philip, P.A.; Sarkar, F.H. Clinical advances in molecular biomarkers for cancer diagnosis and therapy. Int. J. Mol. Sci. 2013, 14, 14771–14784. [Google Scholar]
- Parkin, D.M.; Pisani, P.; Ferlay, J. Estimates of the worldwide incidence of 25 major cancers in 1990. Int. J. Cancer 1999, 80, 827–841. [Google Scholar]
- Macfarlane, L.A.; Murphy, P.R. microRNA: Biogenesis function and role in cancer. Curr. Genomics 2010, 11, 537–561. [Google Scholar]
- Ketting, R.F. microRNA biogenesis and function: An overview. Adv. Exp. Med. Boil. 2011, 700, 1–14. [Google Scholar]
- Treiber, T.; Treiber, N.; Meister, G. Regulation of microRNA biogenesis and function. Thromb. Haemost. 2012, 107, 605–610. [Google Scholar]
- Guo, J.; Miao, Y.; Xiao, B.; Huan, R.; Jiang, Z.; Meng, D.; Wang, Y. Differential expression of microRNA species in human gastric cancer versus non-tumorous tissues. J. Gastroenterol. Hepatol. 2009, 24, 652–657. [Google Scholar]
- Ueda, T.; Volinia, S.; Okumura, H.; Shimizu, M.; Taccioli, C.; Rossi, S.; Alder, H.; Liu, C.G.; Oue, N.; Yasui, W.; et al. Relation between microRNA expression and progression and prognosis of gastric cancer: A microRNA expression analysis. Lancet Oncol. 2010, 11, 136–146. [Google Scholar]
- Kim, C.H.; Kim, H.K.; Rettig, R.L.; Kim, J.; Lee, E.T.; Aprelikova, O.; Choi, I.J.; Munroe, D.J.; Green, J.E. miRNA signature associated with outcome of gastric cancer patients following chemotherapy. BMC Med. Genomics 2011, 4, 79. [Google Scholar]
- Zhao, X.; Dou, W.; He, L.; Liang, S.; Tie, J.; Liu, C.; Li, T.; Lu, Y.; Mo, P.; Shi, Y.; et al. MicroRNA-7 functions as an anti-metastatic microRNA in gastric cancer by targeting insulin-like growth factor-1 receptor. Oncogene 2013, 32, 1363–1372. [Google Scholar]
- Zhang, Z.; Liu, S.; Shi, R.; Zhao, G. miR-27 promotes human gastric cancer cell metastasis by inducing epithelial-to-mesenchymal transition. Cancer Genet. 2011, 204, 486–491. [Google Scholar]
- Xia, L.; Zhang, D.; Du, R.; Pan, Y.; Zhao, L.; Sun, S.; Hong, L.; Liu, J.; Fan, D. miR-15b and miR-16 modulate multidrug resistance by targeting BCL2 in human gastric cancer cells. Int. J. Cancer 2008, 123, 372–379. [Google Scholar]
- Tsujiura, M.; Ichikawa, D.; Komatsu, S.; Shiozaki, A.; Takeshita, H.; Kosuga, T.; Konishi, H.; Morimura, R.; Deguchi, K.; Fujiwara, H.; et al. Circulating microRNAs in plasma of patients with gastric cancers. Br. J. Cancer 2010, 102, 1174–1179. [Google Scholar]
- Moldovan, L.; Batte, K.; Wang, Y.; Wisler, J.; Piper, M. Analyzing the circulating microRNAs in exosomes/extracellular vesicles from serum or plasma by qRT-PCR. Methods Mol. Biol. 2013, 1024, 129–145. [Google Scholar]
- Li, B.S.; Zhao, Y.L.; Guo, G.; Li, W.; Zhu, E.D.; Luo, X.; Mao, X.H.; Zou, Q.M.; Yu, P.W.; Zuo, Q.F.; et al. Plasma microRNAs miR-223 miR-21 and miR-218 as novel potential biomarkers for gastric cancer detection. PLoS One 2012, 7, e41629. [Google Scholar]
- Liu, H.; Zhu, L.; Liu, B.; Yang, L.; Meng, X.; Zhang, W.; Ma, Y.; Xiao, H. Genome-wide microRNA profiles identify miR-378 as a serum biomarker for early detection of gastric cancer. Cancer Lett. 2012, 316, 196–203. [Google Scholar]
- Li, C.; Li, J.F.; Cai, Q.; Qiu, Q.Q.; Yan, M.; Liu, B.Y.; Zhu, Z.G. MiRNA-199a-3p: A potential circulating diagnostic biomarker for early gastric cancer. J. Surg. Oncol. 2013, 108, 89–92. [Google Scholar]
- Cai, H.; Yuan, Y.; Hao, Y.F.; Guo, T.K.; Wei, X.; Zhang, Y.M. Plasma microRNAs serve as novel potential biomarkers for early detection of gastric cancer. Med. Oncol. 2013, 30, 452. [Google Scholar]
- Valladares-Ayerbes, M.; Reboredo, M.; Medina-Villaamil, V.; Iglesias-Diaz, P.; Lorenzo-Patino, M.J.; Haz, M.; Santamarina, I.; Blanco, M.; Fernandez-Tajes, J.; Quindos, M.; et al. Circulating miR-200c as a diagnostic and prognostic biomarker for gastric cancer. J. Transl. Med. 2012, 10, 186. [Google Scholar]
- Elyakim, E.; Sitbon, E.; Faerman, A.; Tabak, S.; Montia, E.; Belanis, L.; Dov, A.; Marcusson, E.G.; Bennett, C.F.; Chajut, A.; et al. hsa-miR-191 is a candidate oncogene target for hepatocellular carcinoma therapy. Cancer Res. 2010, 70, 8077–8087. [Google Scholar]
- Shen, J.; DiCioccio, R.; Odunsi, K.; Lele, S.B.; Zhao, H. Novel genetic variants in miR-191 gene and familial ovarian cancer. BMC Cancer 2010, 10, 47. [Google Scholar]
- Di Leva, G.; Piovan, C.; Gasparini, P.; Ngankeu, A.; Taccioli, C.; Briskin, D.; Cheung, D.G.; Bolon, B.; Anderlucci, L.; Alder, H.; et al. Estrogen mediated-activation of miR-191/425 cluster modulates tumorigenicity of breast cancer cells depending on estrogen receptor status. PLoS Genet. 2013, 9, e1003311. [Google Scholar]
- Wynendaele, J.; Bohnke, A.; Leucci, E.; Nielsen, S.J.; Lambertz, I.; Hammer, S.; Sbrzesny, N.; Kubitza, D.; Wolf, A.; Gradhand, E.; et al. An illegitimate microRNA target site within the 3′ UTR of MDM4 affects ovarian cancer progression and chemosensitivity. Cancer Res. 2010, 70, 9641–9649. [Google Scholar]
- Colamaio, M.; Borbone, E.; Russo, L.; Bianco, M.; Federico, A.; Califano, D.; Chiappetta, G.; Pallante, P.; Troncone, G.; Battista, S.; et al. miR-191 down-regulation plays a role in thyroid follicular tumors through CDK6 targeting. J. Clin. Endocrinol. Metab. 2011, 96, E1915–E1924. [Google Scholar]
- Wang, B.; Zhang, Q. The expression and clinical significance of circulating microRNA-21 in serum of five solid tumors. J. Cancer Res. Clin. Oncol. 2012, 138, 1659–1666. [Google Scholar]
- Song, J.; Bai, Z.; Han, W.; Zhang, J.; Meng, H.; Bi, J.; Ma, X.; Han, S.; Zhang, Z. Identification of suitable reference genes for qPCR analysis of serum microRNA in gastric cancer patients. Dig. Dis. Sci. 2012, 57, 897–904. [Google Scholar]
- Hanash, S.M.; Baik, C.S.; Kallioniemi, O. Emerging molecular biomarkers—Blood-based strategies to detect and monitor cancer. Nat. Rev. Clin. Oncol. 2011, 8, 142–150. [Google Scholar]
- Lawrie, C.H.; Gal, S.; Dunlop, H.M.; Pushkaran, B.; Liggins, A.P.; Pulford, K.; Banham, A.H.; Pezzella, F.; Boultwood, J.; Wainscoat, J.S.; et al. Detection of elevated levels of tumour-associated microRNAs in serum of patients with diffuse large B-cell lymphoma. Br. J. Haematol. 2008, 141, 672–675. [Google Scholar]
- Song, M.Y.; Pan, K.F.; Su, H.J.; Zhang, L.; Ma, J.L.; Li, J.Y.; Yuasa, Y.; Kang, D.; Kim, Y.S.; You, W.C. Identification of serum microRNAs as novel non-invasive biomarkers for early detection of gastric cancer. PLoS One 2012, 7, e33608. [Google Scholar]
- Bustin, S.A.; Benes, V.; Garson, J.A.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; et al. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 2009, 55, 611–622. [Google Scholar]
- Chen, C.; Ridzon, D.A.; Broomer, A.J.; Zhou, Z.; Lee, D.H.; Nguyen, J.T.; Barbisin, M.; Xu, N.L.; Mahuvakar, V.R.; Andersen, M.R.; et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res. 2005, 33, e179. [Google Scholar]



© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Peng, W.-Z.; Ma, R.; Wang, F.; Yu, J.; Liu, Z.-B. Role of miR-191/425 Cluster in Tumorigenesis and Diagnosis of Gastric Cancer. Int. J. Mol. Sci. 2014, 15, 4031-4048. https://doi.org/10.3390/ijms15034031
Peng W-Z, Ma R, Wang F, Yu J, Liu Z-B. Role of miR-191/425 Cluster in Tumorigenesis and Diagnosis of Gastric Cancer. International Journal of Molecular Sciences. 2014; 15(3):4031-4048. https://doi.org/10.3390/ijms15034031
Chicago/Turabian StylePeng, Wei-Zhao, Ren Ma, Fang Wang, Jia Yu, and Zhi-Bin Liu. 2014. "Role of miR-191/425 Cluster in Tumorigenesis and Diagnosis of Gastric Cancer" International Journal of Molecular Sciences 15, no. 3: 4031-4048. https://doi.org/10.3390/ijms15034031




