Ectopic Expression of BraYAB1-702, a Member of YABBY Gene Family in Chinese Cabbage, Causes Leaf Curling, Inhibition of Development of Shoot Apical Meristem and Flowering Stage Delaying in Arabidopsis thaliana
Abstract
:1. Introduction
2. Results and Discussion
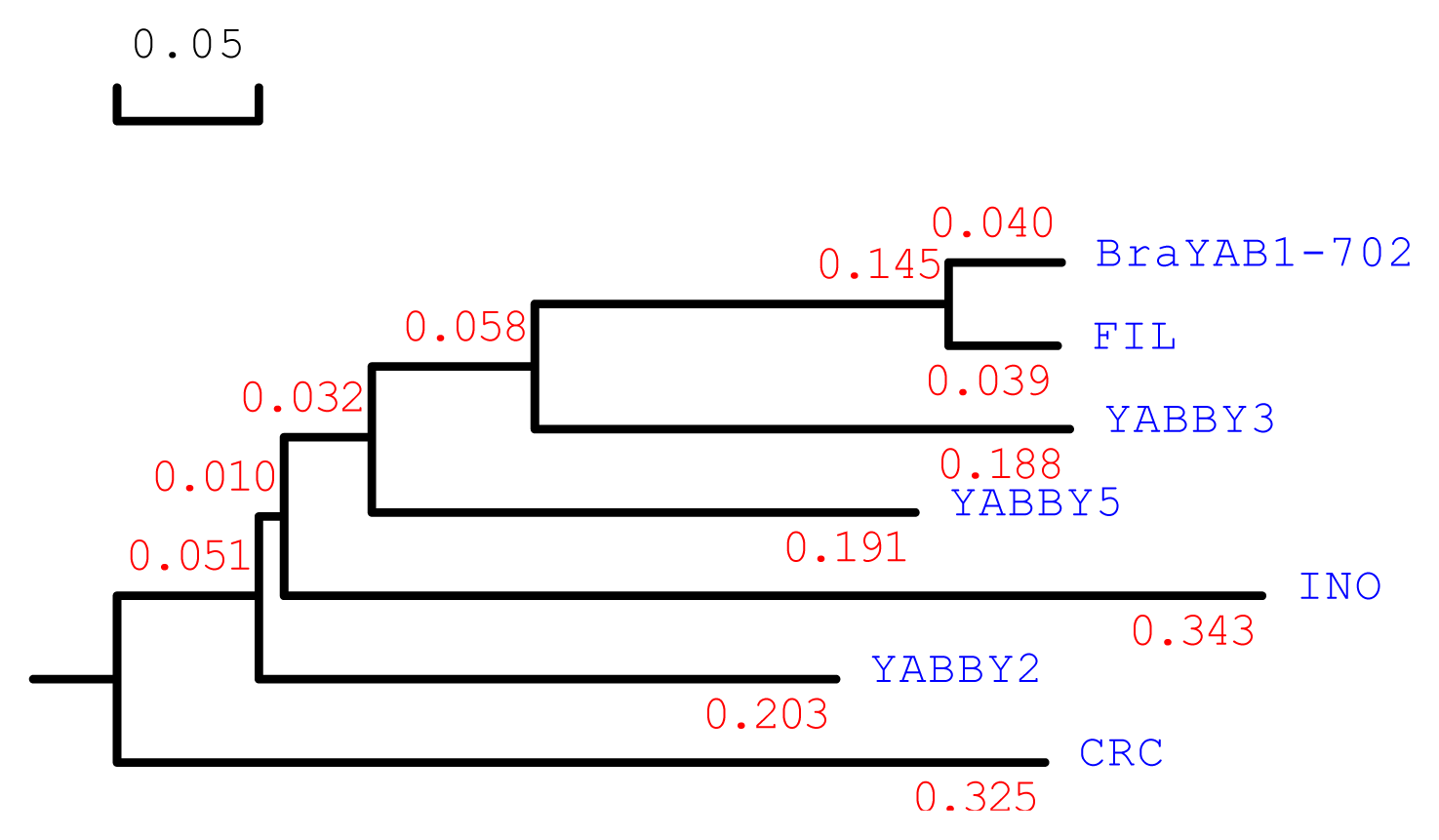
2.1. Cloning and Sequence Analysis of BraYAB1-702 Gene
2.2. Analysis of Promoter Structure of BraYAB1-702 Gene from Chinese Cabbage and FIL Gene from Arabidopsis thaliana
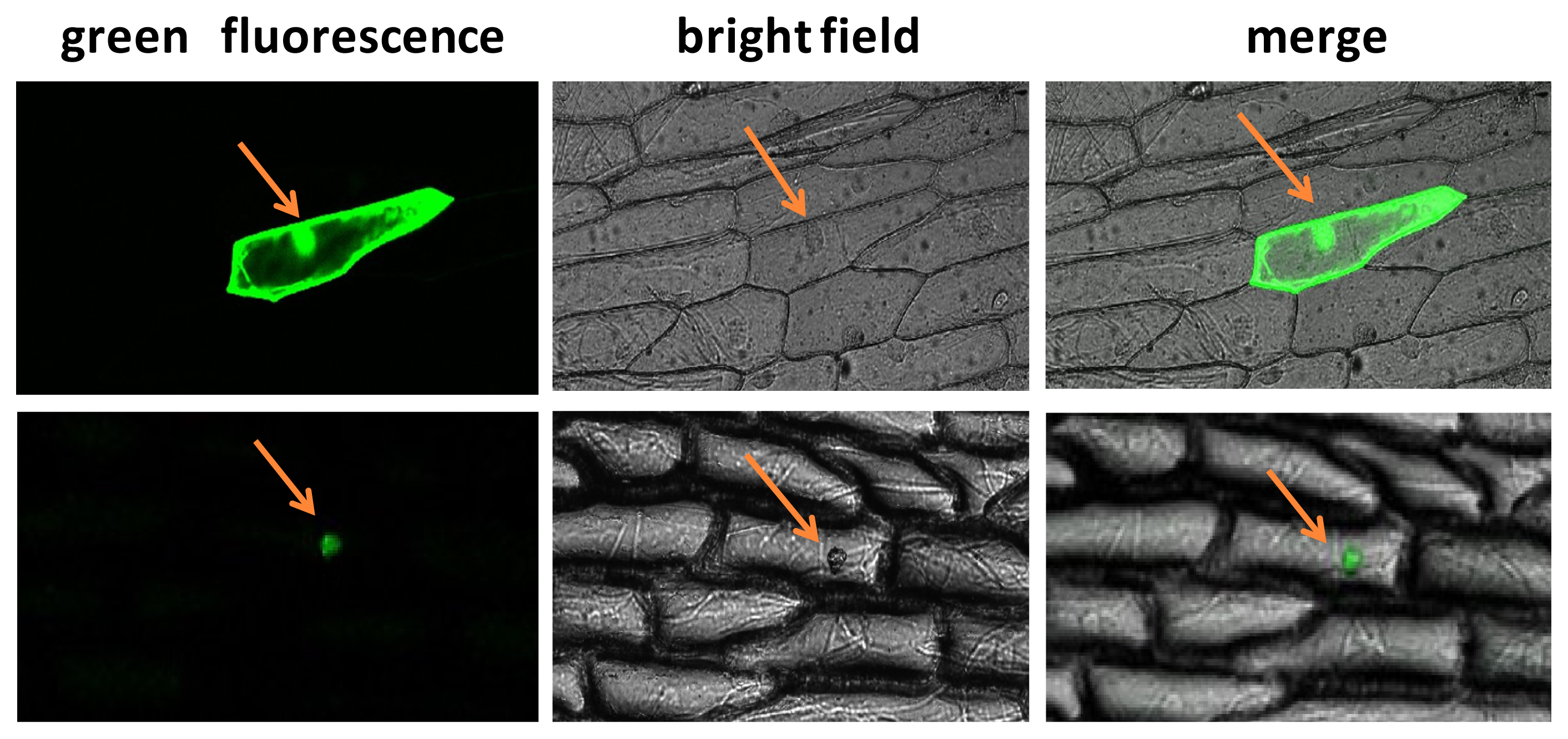
2.3. Subcellular Localization Analysis of BraYAB1-702
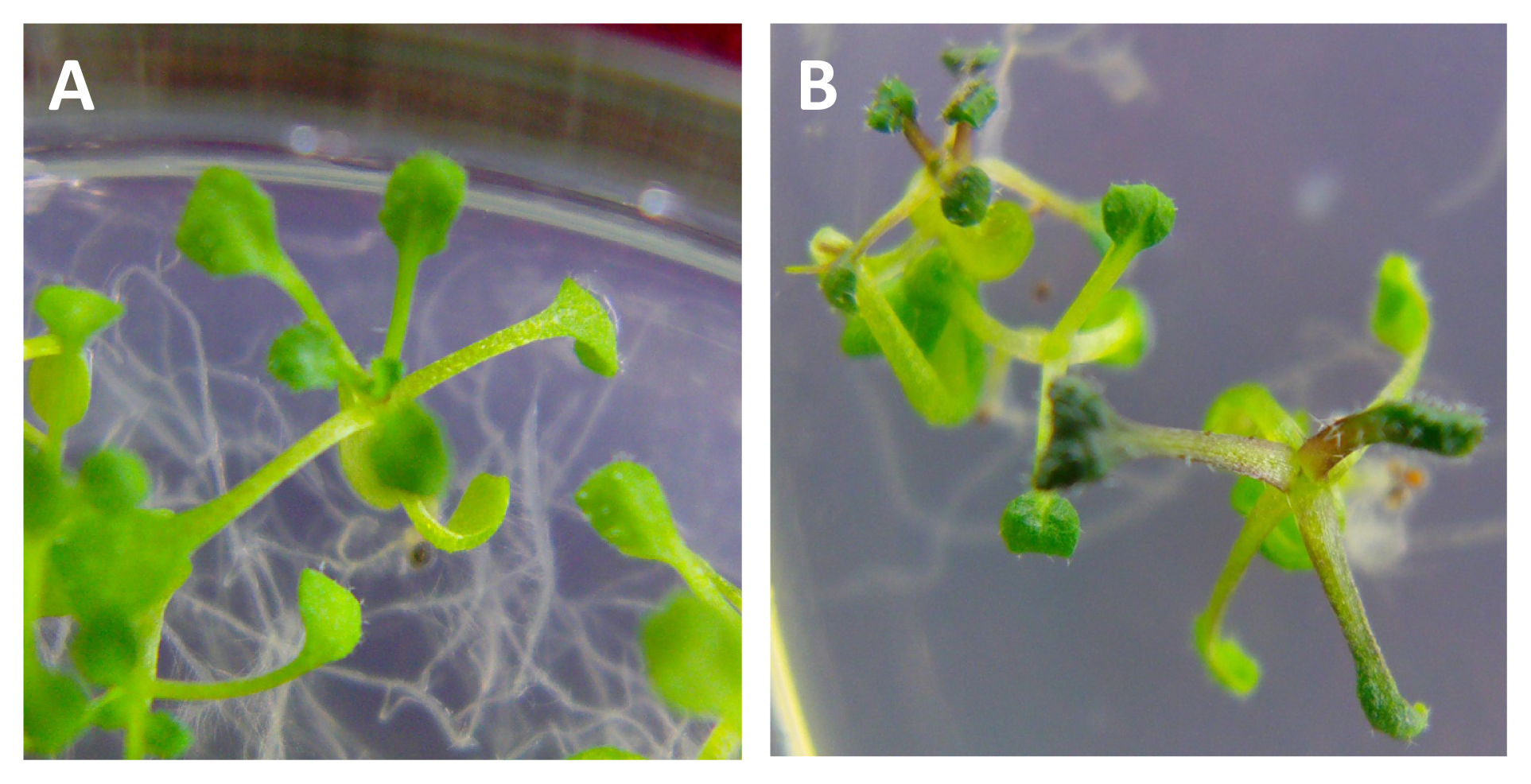
2.4. Phenotype Analysis of Arabidopsis thaliana Transformed with BraYAB1-702 Gene of Chinese Cabbage
2.5. Ectopic Expression of BraYAB1-702 in Arabidopsis thaliana Promotes Abaxialization of Adaxial Epidermises in Leaves
2.6. Real-Time Quantitative PCR of BraYAB1-702 Transgenic Arabidopsis thaliana
3. Experimental Section
3.1. Experimental Materials and Cultivation Management
3.2. Cloning and Sequence Analysis of BraYAB1-702 Gene
- Forward primer: 5′ CCCAAACACTCCAAAAAAGAGAGGA 3′,
- Reverse primer: 5′ CCCAAACACTCCAAAAAAGAGAGGA 3′.
3.3. Identification and Structure Analysis of Promoter Sequences
3.4. Subcellular Localization
- Forward primer: 5′ GGCTCTAGAATGTCTATGTCGTCTATG 3′,
- Reverse primer: 5′ GCCGGTACCTTAATAAGGAATCACACC 3′.
3.5. Construction and Transformation of the BraYAB1-702 Gene Expression Vector and Screening with Hygromycin
3.6. DNA Extraction and DNA-PCR Identification
- Forward primer: 5′ ATGTCTATGTCGTCTATGTCTTCTCC 3′,
- Reverse primer: 5′ TTAATAAGGAATCACACCAACATTAG 3′.
3.7. Analysis of BraYAB1-702, STM, BP and KNAT2 Expression by Real-Time PCR
3.8. Scanning Electron Microscope and Sample Preparation
3.9. Variance Analyses for Phenotype of Wild Type and Transgenic Arabidopsis
4. Conclusions
Acknowledgments
Conflict of Interest
References
- Bowman, J.L.; Eshed, Y.; Baum, S.F. Establishment of polarity in angiosperm lateral organs. Trends Genet 2002, 18, 134–141. [Google Scholar]
- Siegfried, K.R.; Eshed, Y.; Baum, S.F.; Otsuga, D.; Drews, G.N.; Bowman, J.L. Members of the YABBY gene family specify the abaxial cell fate in Arabidopsis. Development 1999, 126, 4117–4128. [Google Scholar]
- Bowman, J.L.; Smyth, D.R. CRABS CLAW, a gene that regulates carpel and nectary development in Arabidopsis, encodes a novel protein with zinc finger and helix-loop-helix domains. Development 1999, 126, 2387–2396. [Google Scholar]
- Villanueva, J.M.; Broadhvest, J.; Hauser, B.A.; Meister, R.J.; Schneitz, K.; Gasser, C.S. INNER NO OUTER regulates abaxial–adaxial patterning in Arabidopsis ovules. Gene Dev 1999, 13, 3160–3169. [Google Scholar]
- Golz, J.F.; Roccaro, M.; Kuzoff, R.; Hudson, A. GRAMINIFOLIA promotes growth and polarity of Antirrhinum leaves. Development 2004, 131, 3661–3670. [Google Scholar]
- Jang, S.; Hur, J.; Kim, S.J.; Han, M.J.; Kim, S.R.; An, G. Ectopic expression of OsYAB1 causes extra stamens and carpels in rice. Plant Mol. Biol 2004, 56, 133–143. [Google Scholar]
- Ku, L.; Zhang, J.; Guo, S.; Liu, H.; Zhao, R.; Chen, Y. Integrated multiple population analysis of leaf architecture traits in maize (Zea mays L.). J. Exp. Bot 2011, 63, 261–274. [Google Scholar]
- Zhao, W.; Su, H.Y.; Song, J.; Zhao, X.Y.; Zhang, X.S. Ectopic expression of TaYAB1, a member of YABBY gene family in wheat, causes the partial abaxialization of the adaxial epidermises of leaves and arrests the development of shoot apical meristem in Arabidopsis. Plant Sci 2006, 170, 364–371. [Google Scholar]
- Sawa, S.; Watanabe, K.; Goto, K.; Kanaya, E.; Morita, E.H.; Okada, K. FILAMENTOUS FLOWER, a meristem and organ identity gene of Arabidopsis, encodes a protein with a zinc finger and HMG-related domains. Gene Dev 1999, 13, 1079–1088. [Google Scholar]
- Eshed, Y.; Izhaki, A.; Baum, S.F.; Floyd, S.K.; Bowman, J.L. Asymmetric leaf development and blade expansion in Arabidopsis are mediated by KANADI and YABBY activities. Development 2004, 131, 2997–3006. [Google Scholar]
- Ori, N.; Eshed, Y.; Chuck, G.; Bowman, J.L.; Hake, S. Mechanisms that control knox gene expression in the Arabidopsis shoot. Development 2000, 127, 5523–5532. [Google Scholar]
- Kumaran, M.K.; Bowman, J.L.; Sundaresan, V. YABBY polarity genes mediate the repression of KNOX homeobox genes in Arabidopsis. Plant Cell 2002, 14, 2761–2770. [Google Scholar]
- Juarez, M.T.; Twigg, R.W.; Timmermans, M.C. Specification of adaxial cell fate during maize leaf development. Development 2004, 131, 4533–4544. [Google Scholar]
- Zhao, X.Y.; Xie, H.T.; Chen, X.B.; Wang, S.S.; Zhang, X.S. Ectopic expression of TaYAB2, a member of YABBY gene family in wheat, causes partial abaxialization of adaxial epidermises of leaves in Arabidopsis. Acta Agron. Sin 2012, 38, 2042–2051. [Google Scholar]
- Bonaccorso, O.; Lee, J.E.; Puah, L.; Scutt, C.P.; Golz, J.F. FILAMENTOUS FLOWER controls lateral organ development by acting as both an activator and a repressor. BMC Plant Biol. 2012, 12. [Google Scholar] [CrossRef]
- Jackson, D.; Veit, B.; Hake, S. Expression of maize KNOTTED1 related homeobox genes in the shoot apical meristem predicts patterns of morphogenesis in the vegetative shoot. Development 1994, 120, 405–413. [Google Scholar]
- Long, J.A.; Moan, E.I.; Medford, J.I.; Barton, M.K. A member of the KNOTTED class of homeodomain proteins encoded by the STM gene of Arabidopsis. Nature 1996, 379, 66–69. [Google Scholar]
- Sentoku, N.; Sato, Y.; Kurata, N.; Ito, Y.; Kitano, H.; Matsuoka, M. Regional expression of the rice KN1-type homeobox gene family during embryo, shoot, and flower development. Plant Cell 1999, 11, 1651–1663. [Google Scholar]
- Bowman, J.L.; Eshed, Y. Formation maintenance of the shoot apical meristem. Trends Plant Sci 2000, 5, 110–115. [Google Scholar]
- Barton, M.K.; Poethig, R.S. Formation of the shoot apical meristem in Arabidopsis thaliana: An analysis of development in the wild type and in the shoot meristemless mutant. Development 1993, 119, 823–831. [Google Scholar]
- Endrizzi, K.; Moussian, B.; Haecker, A.; Levin, J.Z.; Laux, T. The SHOOT MERISTEMLESS gene is required for maintenance of undifferentiated cells in Arabidopsis shoot and floral meristems and acts at a different regulatory level than the meristem genes WUSCHEL and ZWILLE. Plant J 1996, 10, 967–979. [Google Scholar]
- Mele, G.; Ori, N.; Sato, Y.; Hake, S. The knotted1-like homeobox gene BREVIPEDICELLUS regulates cell differentiation by modulating metabolic pathways. Gene Dev 2003, 17, 2088–2093. [Google Scholar]
- Byrne, M.E.; Barley, R.; Curtis, M.; Arroyo, J.M.; Dunham, M.; Hudson, A.; Martienssen, R.A. Asymmetric leaves1 mediates leaf patterning and stem cell function in Arabidopsis. Lett. Nat 2000, 408, 967–971. [Google Scholar]
- Pautot, V.; Dockx, J.; Hamant, O.; Kronenberger, J.; Grandjean, O.; Jublot, D.; Traas, J. KNAT2 evidence for a link between Knotted-like genes and carpel development. Plant Cell 2001, 13, 1719–1734. [Google Scholar]
- Hamant, O.; Nogué, F.; Belles-Boix, E.; Jublot, D.; Grandjean, O.; Traas, J.; Pautot, V. The KNAT2 homeodomain protein interacts with ethylene and cytokinin signaling. Plant Physiol 2002, 130, 657–665. [Google Scholar]
- BLAST, Basic Local Alignment Search Tool. Available online: http://blast.ncbi.nlm.nih.gov/Blast.cgi (on accessed 30 March 2013).
- NCBI Conserved Domains. Available online: http://www.ncbi.nlm.nih.gov/cdd (on accessed 30 March 2013).
- Brassica Database. Available online: http://brassicadb.org/brad/ (on accessed 1 July 2013).
- NCBI, Genome. Available online: http://www.ncbi.nlm.nih.gov/genome?term=txid3702%5Borgn%5D (on accessed 1 July 2013).
- Plant Care, Plant Cis-Acting Regulatory Elements. Available online: http://bioinformatics.psb.ugent.be/webtools/plantcare/html/ (on accessed 1 July 2013).
- Rey, S.; Acab, M.; Gardy, J.L.; Laird, M.R.; Lambert, C.; Brinkman, F.S. PSORTdb: A protein subcellular localization database for bacteria. Nucleic Acids Res 2005, 33, D164–D168. [Google Scholar]
- Hui, M.X. Fine Mapping of Lobe-Leafed Gene and Cloning Functional Analysis of Boundary-Specific BcCUC1 and BuCUC3 Genes from Non-Heading Chinese Cabbage (Brassica campestris ssp. chinensis). Ph.D. Thesis, Northwest A&F University, Yangling, China, September 2011. [Google Scholar]











| Primer name | Forward primer(5′→3′) | Reverse primer(5′→3′) |
|---|---|---|
| BraYAB1-702 | TGATCCTCTCTCCCCTTCGG | CGGACGGTTACGGTCTTGAA |
| UBC | TAACTGCGACTCAGGGAATCTT | TCATCCTTTCTTAGGCATAGCG |
| STM | TGTCAGAAGGTTGGAGCACCA | TTTGTTGCTCCGAAGGGTAA |
| BP | TCATGGAAGCATACTGTGACA | TGACTCAGAAGGATATGGCCA |
| KNAT2 | GATTGCCAAAAGGTGGGAGC | TGTCGCCTTCAGTAGGGTA |
| ACTIN8 | ATGAAGATTAAGGTCGTGGCA | CCGAGTTTGAAGAGGCTAC |
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Zhang, X.-L.; Yang, Z.-P.; Zhang, J.; Zhang, L.-G. Ectopic Expression of BraYAB1-702, a Member of YABBY Gene Family in Chinese Cabbage, Causes Leaf Curling, Inhibition of Development of Shoot Apical Meristem and Flowering Stage Delaying in Arabidopsis thaliana. Int. J. Mol. Sci. 2013, 14, 14872-14891. https://doi.org/10.3390/ijms140714872
Zhang X-L, Yang Z-P, Zhang J, Zhang L-G. Ectopic Expression of BraYAB1-702, a Member of YABBY Gene Family in Chinese Cabbage, Causes Leaf Curling, Inhibition of Development of Shoot Apical Meristem and Flowering Stage Delaying in Arabidopsis thaliana. International Journal of Molecular Sciences. 2013; 14(7):14872-14891. https://doi.org/10.3390/ijms140714872
Chicago/Turabian StyleZhang, Xin-Ling, Ze-Ping Yang, Jing Zhang, and Lu-Gang Zhang. 2013. "Ectopic Expression of BraYAB1-702, a Member of YABBY Gene Family in Chinese Cabbage, Causes Leaf Curling, Inhibition of Development of Shoot Apical Meristem and Flowering Stage Delaying in Arabidopsis thaliana" International Journal of Molecular Sciences 14, no. 7: 14872-14891. https://doi.org/10.3390/ijms140714872




