Detection of Quantitative Trait Loci (QTLs) for Resistances to Small Brown Planthopper and Rice Stripe Virus in Rice Using Recombinant Inbred Lines
Abstract
:1. Introduction
2. Results
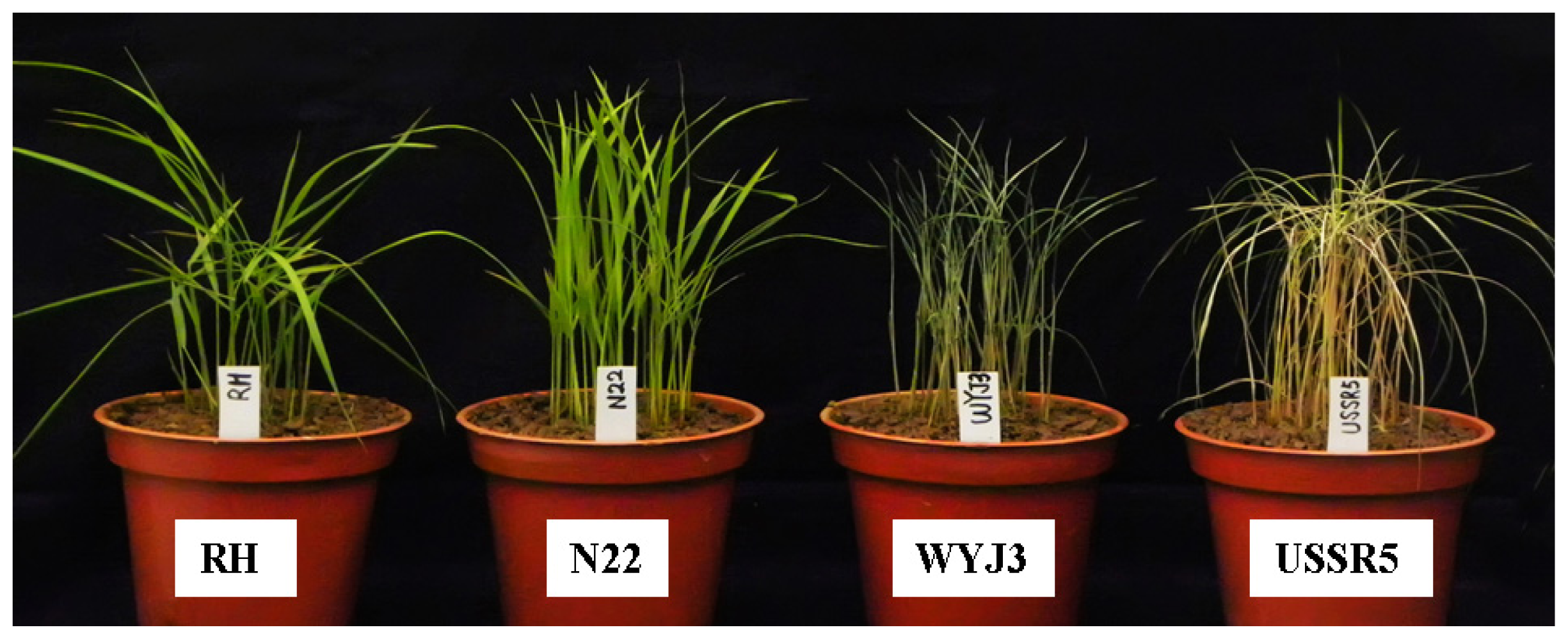
2.1. Screening Rice Varieties for Resistance to SBPH
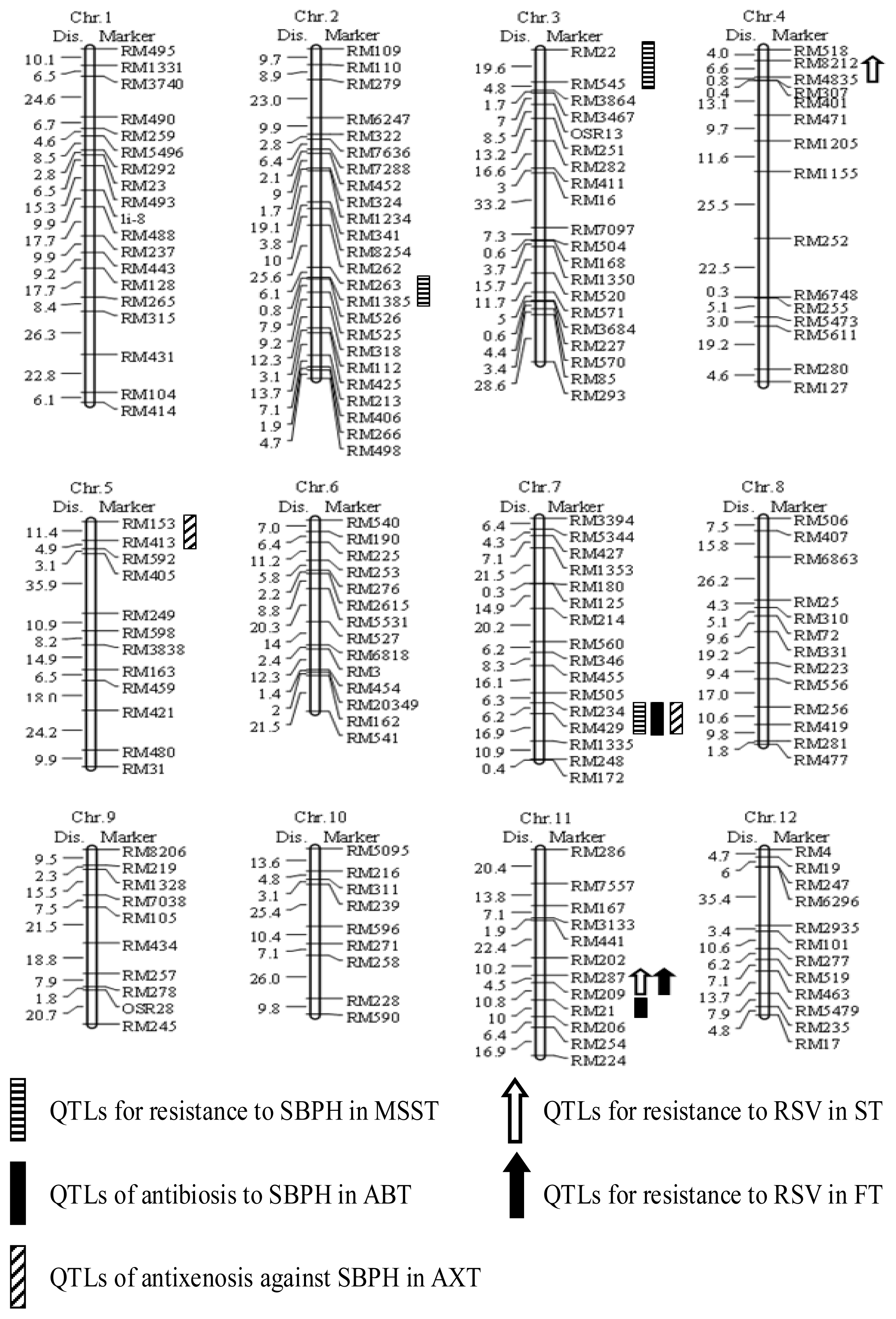
2.2. Construction of a Linkage Map with Simple Sequence Repeat (SSR) Markers
2.3. Evaluation of SBPH Reaction and QTL Analysis
2.4. Antibiosis Test and QTL Analysis
2.5. Antixenosis against SBPH and QTL Detection
2.6. QTL Analysis of RSV Resistance in the RIL Population
3. Discussion
3.1. Genetic Mechanisms of Resistance to SBPH in “N22”
3.2. A Reliable QTL for SBPH Resistance Detected on the Long Arm of Chromosome 7
3.3. The Inheritance of the RSV Resistance Present in “N22”
4. Experimental Section
4.1. Plant Materials
4.2. Insect Population
4.3. Inoculation Methods
- (1)
- Modified seedbox screening test (MSST): a modified seed-box screening test was applied to evaluate reactions of 312 rice accessions and control varieties, as well as the parents and 182 BILs, as described previously [20]. To evaluate each genotype, about 60 uniformly germinated seeds of each line were sown in an 8 cm diameter plastic pot with a hole in the base. Generally, 28 pots, together with one pot of each parent and the control variety, were placed in a 65 × 44 × 14 cm plastic seedbox. All seedlings under evaluation were incubated at 26 ± 2 °C in sunlight. About 2 cm of water was maintained in the bottom of the seedbox. At the 1.5- to 2.0-leaf stage, seedlings were infested with second to third instar SBPH nymphs at 15 insects per seedling. Scoring of all materials in each seedbox according to the standard evaluation systems [42] was conducted when more than 90% of Wuyujing 3 seedlings were dead at 14 ± 1 days after infestation. The score for each entry was then calculated based on the weighted average of the number of seedlings tested (Table 5).
- (2)
- Antixenosis test (AXT): following Duan et al. [20], 15 germinated seeds of each entry were grown in a row in a 65 × 44 × 14 cm plastic seedbox at 26 ± 2 °C. At the 1.5- to 2.0-leaf stage, seedlings were transferred into cages covered with nylon nets and infested with second to third instar SBPH nymphs at a rate of five insects per seedling. The number of insects was counted on each seedling at 8:00 and 16:00 daily, and the insects were then dispersed in order to distribute them evenly among seedlings after counting every day [43]. The average number of insects on each entry was calculated and regarded as the score value of antixenosis after 5 days.
- (3)
- Antibiosis test (ABT): following Duan et al. [20], 5 germinated seeds for each entry (4 replicates) were grown in a 6 cm diameter × 15 cm high glass at 26 ± 2 °C. At the 1.5- to 2.0- leaf stage, seedlings were infested with 1–2 instar SBPH nymphs at a rate of 20 insects per glass. At 10 days after infestation, the survival percentage of insects on each variety was calculated and regarded as the antibiosis value.
- (1)
- A field test (FT) done in a paddy field at Nanjing. Field trials were conducted in randomized complete blocks with two replicates. Sixty seeds of each RIL were sown in a 40 × 60 cm area on 10 May 2009. Weak seedlings were eliminated until ~40 seedlings remained at the 2.5 leaf stage. Wheat surrounding the paddy field was harvested on 5 June, and imagoes of SBPH were transferred to the rice seedlings. No pesticide was used during the entire growth period.
- (2)
- A seedling test (ST) followed Sakurai et al. [44] with a few modifications: 30 germinated seeds of each line were sown in plastic dishes filled with soil. Weak seedlings were eliminated at the one leaf stage and 25 healthy seedlings of each line were kept for infestation. First to second instar SBPH nymphs were released into dishes covered with plastic cylinders at the rate of about five nymphs per seedling, when the seedlings were at the 1.5 leaf stage. During the infestation period, the insects in each dish were dispersed every day to avoid aggregation. Three days later, all SBPH nymphs were killed with pesticide, and seedlings were transferred to a greenhouse, where they produced symptoms after about one month. The experiments were performed with four replications. A relative disease rating index (RDRI = DRI × 100/the value of WYJ3) was calculated for each line, and QTL analysis was conducted, excluding the effect of the environment [45].
4.4. Genetic Linkage Map and QTL Analysis
5. Conclusions
Acknowledgments
References
- Gu, B.L.; Xue, P.X.; Shi, W.X.; Zhou, L.H. Observation on rice spikes infected by Laodelphax striatellus and rice yield loss. China Plant Prot 2005, 25, 7–8. (in Chinese). [Google Scholar]
- Tai, D.L.; Li, Y.; Mei, A.Z.; Ding, Z.K.; Wang, C.L.; Zhong, F.X. Factors for the outbreak of the small brown planthopper and its control measures in 2004. China Plant Prot 2005, 25, 33–35. (in Chinese). [Google Scholar]
- Zhang, J.F.; Gong, L.G.; Qu, Y.; Qu, H.L. The rice ears were damaged by the fifth and sixth generation of small brown planthopper in Changshu City in 2004. China Plant Prot 2005, 25, 39. (in Chinese). [Google Scholar]
- Sun, D.Z.; Jiang, L.; Zhang, Y.X.; Cheng, X.N.; Zhai, H.Q.; Wan, J.M. Quantitative trait loci for resistance to stripe disease in rice (Oryza sativa L.). Rice Sci 2007, 14, 157–160. [Google Scholar]
- Wang, B.X.; Jiang, L.; Chen, L.M.; Lu, B.G.; Wang, Q.; Le, Q.T.; Fan, J.W.; Chen, X.N.; Zhai, H.Q.; Xu, D.Y.; et al. Screening of rice resources against rice black-streaked dwarf virus and mapping of resistant QTL. Acta Agron. Sin 2010, 36, 1258–1264. (in Chinese). [Google Scholar]
- Xie, L.H. Research on rice virus diseases in China. Trop. Agric. Res. Ser 1986, 19, 45–50. [Google Scholar]
- Sun, D.Z.; Jiang, L.; Zhang, Y.X.; Cheng, X.N.; Wang, C.M.; Zhai, H.Q.; Wan, J.M. Resistance to rice stripe virus in eight rice varieties. Chin. J. Rice Sci 2006, 20, 219–222. (in Chinese). [Google Scholar]
- Hibino, H. Biology and epidemiology of rice viruses. Annu. Rev. Phytopathol 1996, 34, 249–274. [Google Scholar]
- Wang, H.D.; Chen, J.P.; Wang, A.G.; Jiang, X.H.; Adams, M.J. Studies on the epidemiology and yield losses from rice black-streaked dwarf disease in a recent epidemic in Zhejiang province, China. Plant Pathol. 2009, 58, 815–825. [Google Scholar]
- Xu, X.L.; Zhang, Y.G.; Yang, A.G. Study on the relationship between the numbers of Laodelphax striatellus and the incidences of rice stripe disease. China Plant Prot 2005, 3, 5–6. (in Chinese). [Google Scholar]
- Wang, H.D.; Zhu, Z.R.; Chen, J.P.; Wang, E.G.; Li, B.F. Epidemics, monitoring and key control techniques of the rice black-streaked dwarf viral disease. Acta Agric. Zhejiangensis 2007, 19, 141–146. [Google Scholar]
- Liao, X.G.; Wu, H.L.; Zhu, Z.R. Different rice varieties affect on the rice black-streaked virus disease occurrence. Plant Prot 1999, 25, 1–4. [Google Scholar]
- Sone, S.; Hattori, Y.; Tsuboi, S.; Otsu, Y. Difference in susceptibility to imidacloprid of the populations of the small brown planthopper, Laodelphax striatellus Fallén. J. Pestic. Sci 1995, 20, 541–543. [Google Scholar]
- Endo, S.; Tsurumachi, M. Insecticide resistance and insensitive acetylcholinesterase in small brown planthopper, Laodelphax striatellus. J. Pestic. Sci 2000, 25, 395–397. [Google Scholar]
- Endo, S.; Takahashi, A.; Tsurumachi, M. Insecticide susceptibility of the small brown planthopper, Laodelphax striatellus Fallén (Homoptera: Delphacidae), collected from East Asia. Appl. Entomol. Zool 2002, 37, 79–84. [Google Scholar]
- Hayano-Saito, Y.; Tsuji, T.; Fujii, K.; Saito, K.; Iwasaki, M.; Saito, A. Localization of the rice stripe disease resistance gene, Stvb-i, by graphical genotyping and linkage analyses with molecular markers. Theor. Appl. Genet 1998, 96, 1044–1049. [Google Scholar]
- Zhang, Y.X.; Wang, Q.; Jiang, L.; Liu, L.L.; Wang, B.X.; Shen, Y.Y.; Cheng, X.N.; Wan, J.M. Fine mapping of qSTV11KAS, a major QTL for rice stripe disease. Theor. Appl. Genet 2011, 122, 1591–1604. [Google Scholar]
- Wu, X.J.; Zuo, S.M.; Chen, Z.X.; Zhang, Y.F.; Zhu, J.K.; Ma, N.; Tang, J.Y.; Chu, C.C.; Pan, X.B. Fine mapping of qSTV11TQ, a major gene conferring resistance to rice stripe disease. Theor. Appl. Genet 2010, 122, 915–923. [Google Scholar]
- Li, A.H.; Dai, Z.Y.; Ji, H.J.; Zhang, X.X.; Li, Y.H.; Pan, C.H.; Zhang, H.X.; Pan, X.B. Preliminary analysis on resistance of rice black-streaked dwarf viral disease for germplasms with different gene-types. J. Yangzhou Univ 2008, 29, 18–22. (in Chinese). [Google Scholar]
- Duan, C.X.; Wan, J.M.; Zhai, H.Q.; Chen, Q.; Wang, J.K.; Su, N.; Lei, C.L. Quantitative trait loci mapping of resistance to Laodelphax striatellus (Homoptera: Delphacidae) in rice using recombinant inbred lines. J. Econ. Entomol 2007, 100, 1450–1455. [Google Scholar]
- Liu, Y.Q.; Su, C.C.; Jiang, L.; He, J.; Wu, H.; Peng, C.; Wan, J.M. The distribution and identification of brown planthopper resistance genes in rice. Hereditas 2009, 146, 67–73. [Google Scholar]
- Sogawa, K.; Zhang, H.; Yang, X.J.; Liu, G.J. Whitebaceked planthopper resistance in Chinese rice varieties. Chin. J. Rice Sci 2003, 17, 47–52. (in Chinese). [Google Scholar]
- Kisimoto, R. Genetic Variation in the ability of a planthopper vector: Laodelphax striatellus (Fallén) to acquire the rice stripe virus. Virology 1967, 32, 144–152. [Google Scholar]
- Duan, C.X.; Su, N.; Cheng, Z.J.; Lei, C.L.; Wang, J.L.; Zhai, H.Q.; Wan, J.M. QTL analysis for the resistance to small brown planthopper (Laodelphax striatellus Fallén) in rice using backcross inbred lines. Plant Breed 2010, 129, 63–67. [Google Scholar]
- Zhang, Y.X.; Jiang, L.; Liu, L.L.; Wang, B.X.; Shen, Y.Y.; Wang, Q.; Cheng, X.N.; Wan, J.M. Quantitative trait loci associated with resistance to rice stripe virus and small brown planthopper infestation in rice. Crop Sci 2010, 50, 1854–1862. [Google Scholar]
- Duan, C.X.; Cheng, Z.J.; Lei, C.L.; Zhai, H.Q.; Wan, J.M. Analysis of QTLs for resistance to small brown planthopper in rice using an F2 population from a cross between Mudgo and Wuyujing 3. Acta Agron Sin 2009, 35, 388–394. [Google Scholar]
- Le, Q.T.; Liu, Y.Q.; Jiang, L.; Wang, B.X.; Wang, Q.; Hanh, T.T.T.; Wan, J.M. Identification of quantitative trait loci associated with small brown planthopper (Laodelphax striatellus Fallén) resistance in rice (Oryza sativa L.). Hereditas 2012, 149, 16–23. [Google Scholar]
- Sidhu, G.S.; Khush, G.S.; Medrano, F.G. A dominant gene in rice for resistance to white-backed planthopper and its relationship to other plant characteristics. Euphytica 1979, 28, 227–232. [Google Scholar]
- McCouch, S.R.; Khush, G.S.; Tanksley, S.D. Tagging Genes for Disease and Insect Resistance via Linkage to RFLP Markers. In Rice Genetics II: Proceedings of the Second International Rice Genetics Symposium; International Rice Research Institute (IRRI): Manila, Philippines, 1991; pp. 443–448. [Google Scholar]
- Soundararajan, R.P.; Kadirvel, P.; Gunathilagaraj, K.; Maheswaran, M. Mapping of QTLs associated with resistance to brown planthopper by means of a doubled-haploid population in rice. Crop Sci 2004, 44, 2214–2220. [Google Scholar]
- Geethanjali, S.; Kadirvel, P.; Maheswaran, M. Detection of quantitative trait loci (QTL) associated with resistance to whitebacked planthopper (Sogatella furcifera) in rice (Oryza sativa). Plant Breed 2009, 128, 130–136. [Google Scholar]
- Xu, J.; Wang, J.; Ling, Z.; Zhu, L. Analysis of rice blast resistance genes by QTL mapping. Chin. Sci. Bull 2004, 49, 337–342. (in Chinese). [Google Scholar]
- Chen, H.L.; Wang, S.P.; Xing, Y.Z.; Xu, C.G.; Hayes, P.M.; Zhang, Q.F. Comparative analyses of genomic locations and race specificities of loci for quantitative resistance to Pyricularia grisea in rice and barley. Proc. Natl. Acad. Sci. USA 2003, 100, 2544–2549. [Google Scholar]
- Talukder, Z.I.; Tharreau, D.; Price, A.H. Quantitative trait loci analysis suggests that partial resistance to rice blast is mostly determined by race-specific interactions. New Phytol 2004, 162, 197–209. [Google Scholar]
- Ballini, E.; More, J.B.; Droc, G.; Price, A.; Courtois, B.; Notteghem, J.L.; Tharreau, D. A genome-wide meta-analysis of rice blast resistance genes and quantitative trait loci provides new insights into partial and complete resistance. Mol. Plant Microbe Interact 2008, 21, 859–868. [Google Scholar]
- Wang, Y.J.; Zhang, Z.G.; He, X.J.; Zhou, H.L.; Wen, Y.X.; Dai, J.X.; Zhang, J.S.; Chen, S.Y. A rice transcription factor OsbHLH1 is involved in cold stress response. Theor. Appl. Genet 2003, 107, 1402–1409. [Google Scholar]
- Fukuda, A.; Nakamura, A.; Tanaka, Y. Molecular cloning and expression of the Na+/H+ exchanger gene in Oryza sativa. Biochim. Biophys. Acta 1999, 1446, 149–155. [Google Scholar]
- Piao, H.L.; Xuan, Y.H.; Park, S.H.; Je, B.I.; Park, S.J.; Park, S.H.; Kim, C.M.; Huang, J.; Wang, G.K.; Kim, M.J. OsCIPK31, a CBL-interacting protein kinase is involved in germination and seedling growth under abiotic stress conditions in rice plants. Mol. Cells 2010, 30, 19–27. [Google Scholar]
- Huang, J.; Wang, J.F.; Wang, Q.H.; Zhang, H.S. Identification of a rice zinc finger protein whose expression is transiently induced by drought, cold but not by salinity and abscisic acid. DNA Seq 2005, 16, 130–136. [Google Scholar]
- Wang, Z.H.; Zhou, Y.J.; Fan, Y.J.; Xue, B.D.; Wu, S.H.; Cheng, Z.B.; Zhang, W.H. Detection of rice black-streaked dwarf Fijivirus by RT-PCR, dot-blot hybridization and SDS-PAGE. J. Nanjing Agric. Univ 2001, 24, 24–28. (in Chinese). [Google Scholar]
- Zhou, Y.J.; Liu, H.J.; Wang, G.Z.; Huang, Y.Z.; Cheng, B.; Chen, Z.X.; Zhou, X.P. Immunity detecting of rice stripe virus in Laodelphax striatellus. Jiangsu Agric. Sci 2004, 1, 50–51. (in Chinese). [Google Scholar]
- International Rice Research Institute (IRRI), Standard Evaluation Systems for RiceIRRI: Manila, Philippines, 1988.
- Nemoto, H.; Ishikawa, K.; Shimura, E. The resistances to rice stripe virus and small brown planthopper in rice variety IR50. Breed. Sci 1994, 44, 13, –18.. [Google Scholar]
- Sakurai, Y.; Ezuka, A.; Okamoro, H. The seedling test method of varietal resistance of rice plants to stripe virus disease (part 1) bull. Chugoku Agric. Expt. Station 1963, A9, 113–124. (in Japanese). [Google Scholar]
- Sun, D.Z.; Jiang, L.; Liu, S.J.; Zhang, Y.X.; Huang, P.H.; Cheng, X.N.; Zhai, H.Q.; Wan, J.M. Detection of QTLs for resistance to rice stripe virus and small brown planthopper in rice (Oryza sativa L.). Acta Agron. Sin 2006, 32, 805–810. (in Chinese). [Google Scholar]
- Lander, E.S.; Green, P.; Abrahamson, J.; Barlow, A.; Daly, M.J.; Lincoln, S.E.; Newberg, L.A. MAPMAKER: An interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1987, 1, 174–181. [Google Scholar]
- Kosambi, D. The estimation of map distances from recombination values. Ann. Eugen 1994, 12, 172–175. [Google Scholar]
- Wang, S.C.; Basten, C.J.; Zeng, Z.B. Windows QTL Cartographer 2.5; Department of Statistics, North Carolina State University: Raleigh, NC, USA, 2006. [Google Scholar]
- McCouch, S.R.; Cho, Y.G.; Yano, M.; Paule, E.; Blinstrue, M.; Morishima, H.M.; Kinosita, T. Report on QTL nomenclature. Rice Genet. Newsl 1997, 14, 11–13. [Google Scholar]


| Origin Province/Country | Classification a | Total | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Japonica Type | Indica Type | ||||||||||||
| I | HR | R | MR | S | HS | I | HR | R | MR | S | HS a | ||
| Jilin | 8 | 8 | |||||||||||
| Heilongjiang | 1 | 1 | 4 | 6 | |||||||||
| Liaoning | 2 | 1 | 8 | 11 | |||||||||
| Shandong | 1 | 2 | 2 | 2 | 2 | 9 | |||||||
| Shanxi | 1 | 1 | 2 | ||||||||||
| Sichuan | 1 | 2 | 1 | 3 | 7 | ||||||||
| Guizhou | 1 | 1 | 1 | 1 | 4 | ||||||||
| Yunnan | 1 | 2 | 5 | 4 | 1 | 1 | 5 | 2 | 6 | 27 | |||
| Anhui | 1 | 3 | 6 | 4 | 1 | 5 | 20 | ||||||
| Jiangxi | 2 | 1 | 8 | 4 | 15 | ||||||||
| Hubei | 3 | 1 | 1 | 5 | |||||||||
| Hunan | 1 | 3 | 5 | 4 | 1 | 1 | 15 | ||||||
| Guangdong | 3 | 2 | 3 | 7 | 15 | ||||||||
| Guangxi | 2 | 2 | 5 | 3 | 12 | ||||||||
| Fujian | 5 | 1 | 2 | 3 | 11 | ||||||||
| Zhejiang | 2 | 1 | 3 | 4 | 2 | 1 | 1 | 1 | 15 | ||||
| Jiangsu | 3 | 2 | 4 | 1 | 2 | 12 | |||||||
| Taiwan | 1 | 1 | 1 | 5 | 2 | 2 | 12 | ||||||
| Taihu Valley | 6 | 3 | 9 | 15 | 1 | 1 | 35 | ||||||
| IRRI | 4 | 7 | 6 | 2 | 1 | 20 | |||||||
| India | 1 | 2 | 1 | 4 | |||||||||
| South Korea | 1 | 1 | 1 | 1 | 2 | 6 | |||||||
| Malaysia | 1 | 1 | 3 | 1 | 6 | ||||||||
| Indonesia | 2 | 2 | 1 | 5 | |||||||||
| Other | 1 | 1 | 2 | 4 | 11 | 4 | 2 | 3 | 2 | 30 | |||
| Total | 7 | 31 | 26 | 33 | 77 | 6 | 35 | 26 | 31 | 40 | 312 | ||
| Test Method | Control * | Variety | RILs Population | |||
|---|---|---|---|---|---|---|
| WYJ3 | RH | USSR5 | N22 | mean | range | |
| Evaluation of SBPH resistance | ||||||
| MSST | 9.5 ± 0.8 a | 0 c | 9.2 ± 0.4 a | 1.5 ± 0.2 b | 5.2 | 1.0–9.0 |
| ABT | 98.0 ± 0.5 a | 10.0 ± 0.7 c | 95.0 ± 0.6 a | 31.0 ± 1.3 b | 60.1 | 21.0–100.0 |
| AXT | 9.2 ± 0.6 a | 0.8 ± 0.2 c | 9.0 ± 0.2 a | 2.0 ± 0.3 b | 5.8 | 1.0–10.0 |
| Phenotyping Method | QTL | Marker Interval | Chromosome | LOD Score | PVE (%) a | Additive Effect b |
|---|---|---|---|---|---|---|
| Modified seedbox screening test | qSBPH2 | RM263-RM1385 | 2 | 3.03 | 10.0 | 0.81 |
| qSBPH3 | RM22-RM545 | 3 | 2.54 | 7.7 | −0.72 | |
| qSBPH7.1 | RM234-RM429 | 7 | 3.42 | 17.4 | −1.23 | |
| Antibiosis test | qSBPH7.2 | RM234-RM429 | 7 | 3.30 | 13.2 | −10.3 |
| qSBPH11 | RM209-RM21 | 11 | 2.60 | 7.5 | −5.4 | |
| Antixenosis test | qSBPH5 | RM153-RM413 | 5 | 2.51 | 8.2 | −0.37 |
| qSBPH7.3 | RM234-RM429 | 7 | 3.40 | 15.7 | −9.36 | |
| Seedling test | qSTV4 | RM8212-RM4835 | 4 | 5.20 | 13.4 | −3.19 |
| qSTV11.1 | RM287-RM209 | 11 | 8.58 | 28.9 | −7.77 | |
| Field test | qSTV11.2 | RM287-RM209 | 11 | 8.03 | 30.2 | −8.90 |
| Chromosome | QTL | Linked Marker | Population | Reference |
|---|---|---|---|---|
| 1 | qSBPH1 | C949–GA506 | ZYQ8/JX17 DH a lines | Zhang et al. [25] |
| 2 | qSBPH2 | RG322–CT41 | ZYQ8/JX17 DH lines | Zhang et al. [25] |
| Qsbph2 | R1843–R712 | Nipponbare/Kasalath//Nipponbare BILs b | Duan et al. [24] | |
| Qsbph2b | RM5791-RM29 | Mudgo/Wuyujing 3 F2:3 lines | Duan et al. [26] | |
| 3 | Qsbph3b | XNpb74-XNpb144 | Kinmaze/DV85 RILs | Duan et al. [20] |
| Qsbph3b | C80-C1677 | Nipponbare/Kasalath//Nipponbare BILs | Duan et al. [24] | |
| Qsbph3c | R2170–C1135 | Nipponbare/Kasalath//Nipponbare BILs | Duan et al. [24] | |
| Qsbph3d | RM3199–RM5442 | Mudgo/Wuyujing 3 F2:3 lines | Duan et al. [26] | |
| 4 | qSBPH4 | RM451–RM5473 | 02428/Rathu Heenati F2 population | Le et al. [27] |
| 8 | Qsbph8 | C390-R1943 | Nipponbare/Kasalath//Nipponbare BILs | Duan et al. [24] |
| 11 | Qsbph11a | R2918-C410 | Kinmaze/DV85 RILs c | Duan et al. [20] |
| Qsbph11b | XNpb202-C1172 | Kinmaze/DV85 RILs | Duan et al. [20] | |
| Qsbph11c | XNpb202-C1172 | Kinmaze/DV85 RILs | Duan et al. [20] | |
| Qsbph11d | XNpb202-C1172 | Kinmaze/DV85 RILs | Duan et al. [20] | |
| Qsbph11d | R1506–C950 | Nipponbare/Kasalath//Nipponbare BILs | Duan et al. [24] | |
| Qsbph11e | S2260–G257 | Nipponbare/Kasalath//Nipponbare BILs | Duan et al. [24] | |
| Qsbph11f | S2260–G257 | Nipponbare/Kasalath//Nipponbare BILs | Duan et al. [24] | |
| Qsbph11g | S2260–G257 | Nipponbare/Kasalath//Nipponbare BILs | Duan et al. [24] | |
| qSBPH11h | RG211–PTA818 | ZYQ8/JX17 DH lines | Zhang et al. [25] | |
| 12 | Qsbph12a | I12-17–RM3331 | Mudgo/Wuyujing 3 F2:3 lines | Duan et al. [26] |
| qSBPH12 | RM519–RM3331 | 02428/Rathu Heenati F2 population | Le et al. [27] | |
| Symptoms | Score | Reaction a |
|---|---|---|
| No visible damage | 0 | I |
| Very slightly damage | 1 | HR |
| Partial yellowing of the first and the second leaves | 3 | R |
| Pronounced yellowing and some seedlings slight stunting | 5 | MR |
| Seedlings showing signs of wilting and severe stunting | 7 | S |
| Seedlings dead | 9 | HS |
© 2013 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Wang, Q.; Liu, Y.; Hu, J.; Zhang, Y.; Xie, K.; Wang, B.; Tuyen, L.Q.; Song, Z.; Wu, H.; Liu, Y.; et al. Detection of Quantitative Trait Loci (QTLs) for Resistances to Small Brown Planthopper and Rice Stripe Virus in Rice Using Recombinant Inbred Lines. Int. J. Mol. Sci. 2013, 14, 8406-8421. https://doi.org/10.3390/ijms14048406
Wang Q, Liu Y, Hu J, Zhang Y, Xie K, Wang B, Tuyen LQ, Song Z, Wu H, Liu Y, et al. Detection of Quantitative Trait Loci (QTLs) for Resistances to Small Brown Planthopper and Rice Stripe Virus in Rice Using Recombinant Inbred Lines. International Journal of Molecular Sciences. 2013; 14(4):8406-8421. https://doi.org/10.3390/ijms14048406
Chicago/Turabian StyleWang, Qi, Yuqiang Liu, Jinlong Hu, Yingxin Zhang, Kun Xie, Baoxiang Wang, Le Quang Tuyen, Zhaoqiang Song, Han Wu, Yanling Liu, and et al. 2013. "Detection of Quantitative Trait Loci (QTLs) for Resistances to Small Brown Planthopper and Rice Stripe Virus in Rice Using Recombinant Inbred Lines" International Journal of Molecular Sciences 14, no. 4: 8406-8421. https://doi.org/10.3390/ijms14048406





