Molecular Cloning and Characterization of cDNA Encoding a Putative Stress-Induced Heat-Shock Protein from Camelus dromedarius
Abstract
:1. Introduction
2. Results
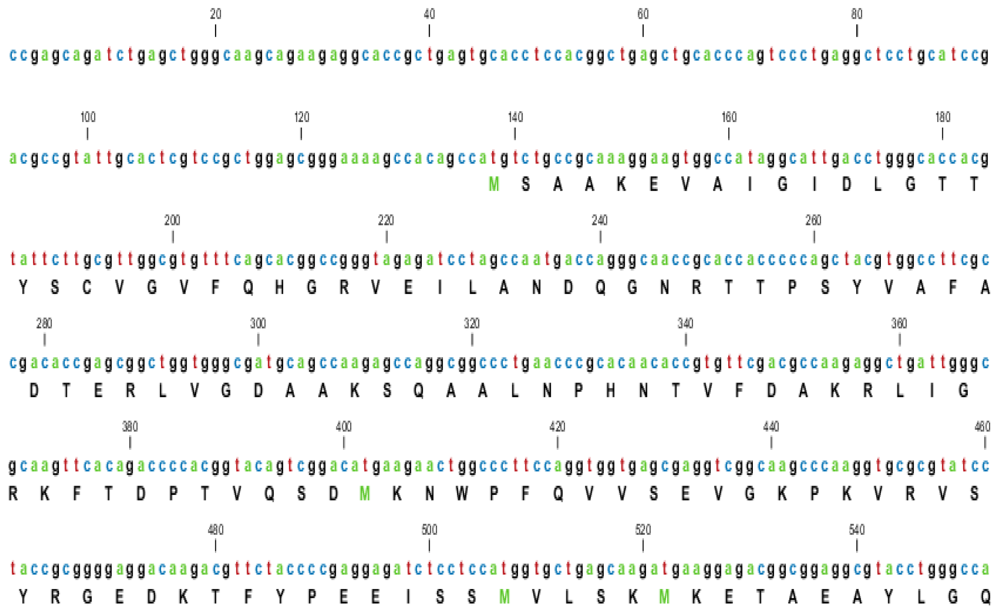
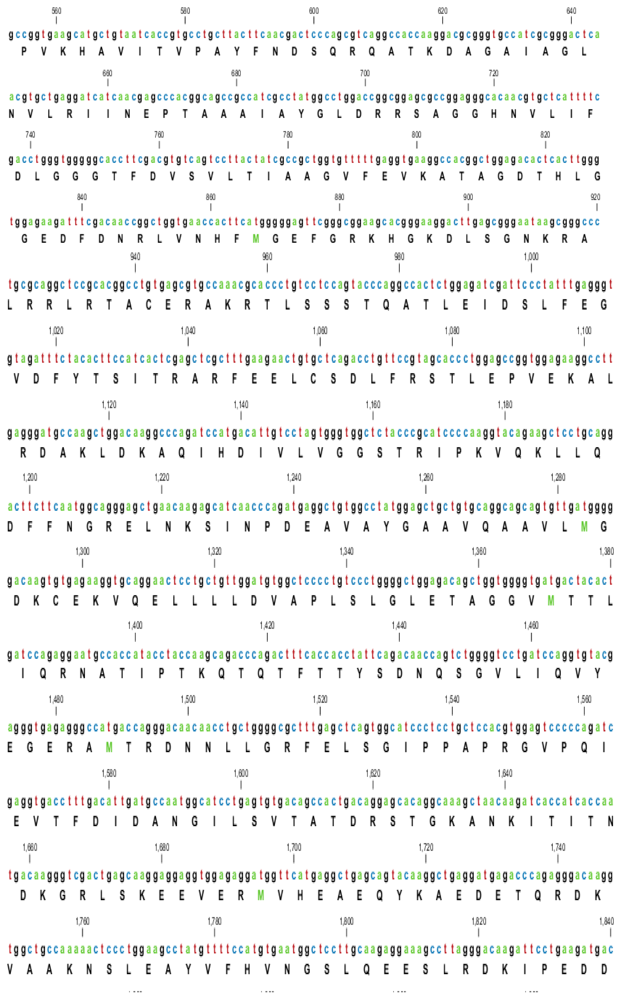
2.1. Characterization of HSPA6 Gene Full-Length cDNA
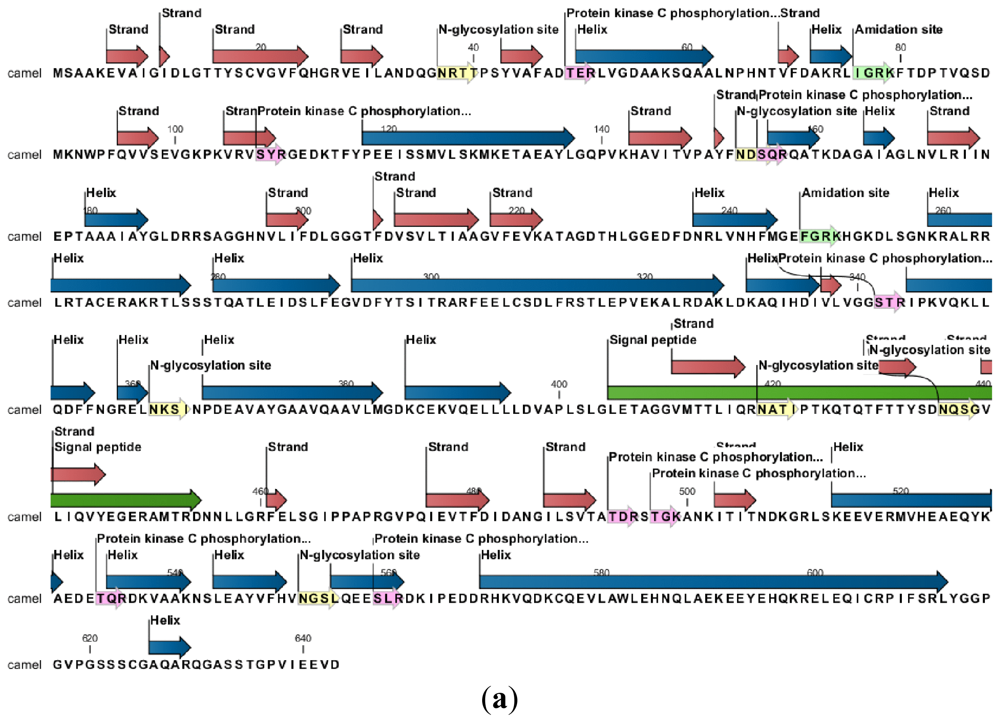
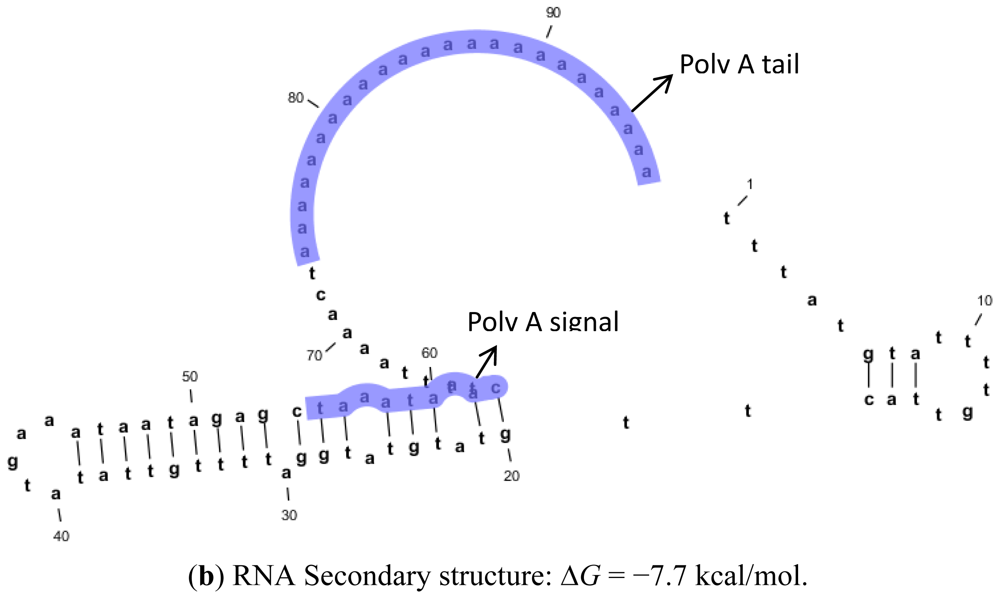
2.2. Protein and RNA Secondary Structure Prediction
2.3. Phylogenetic Analysis
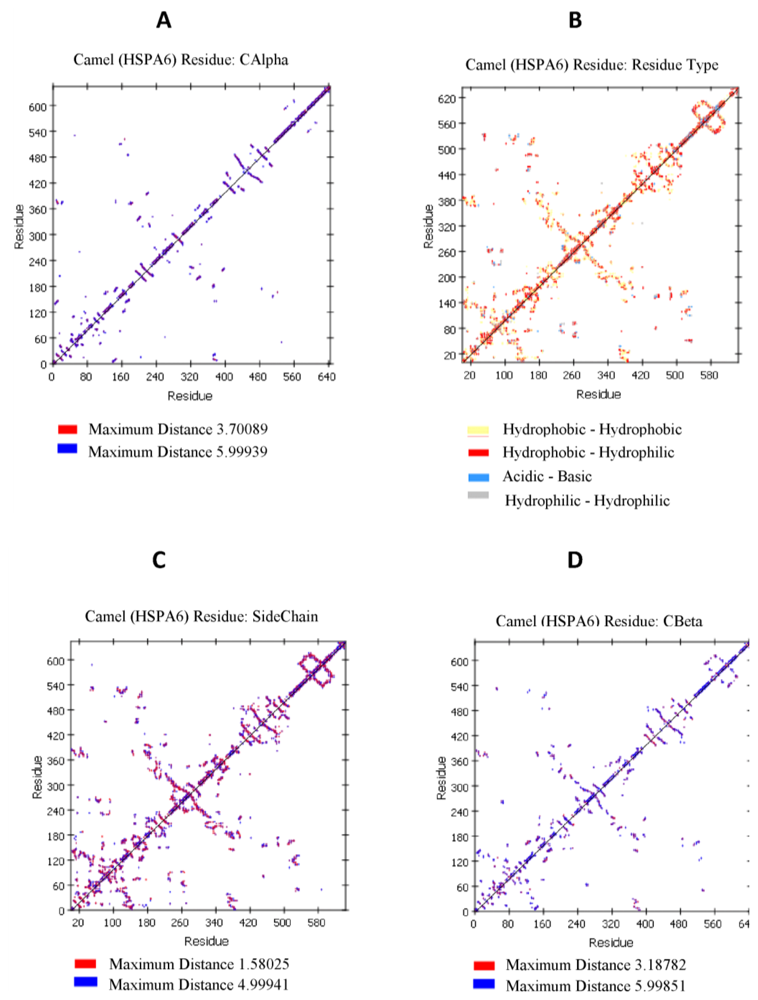
2.4. Camel HSPA6 Protein 3D Structure Prediction
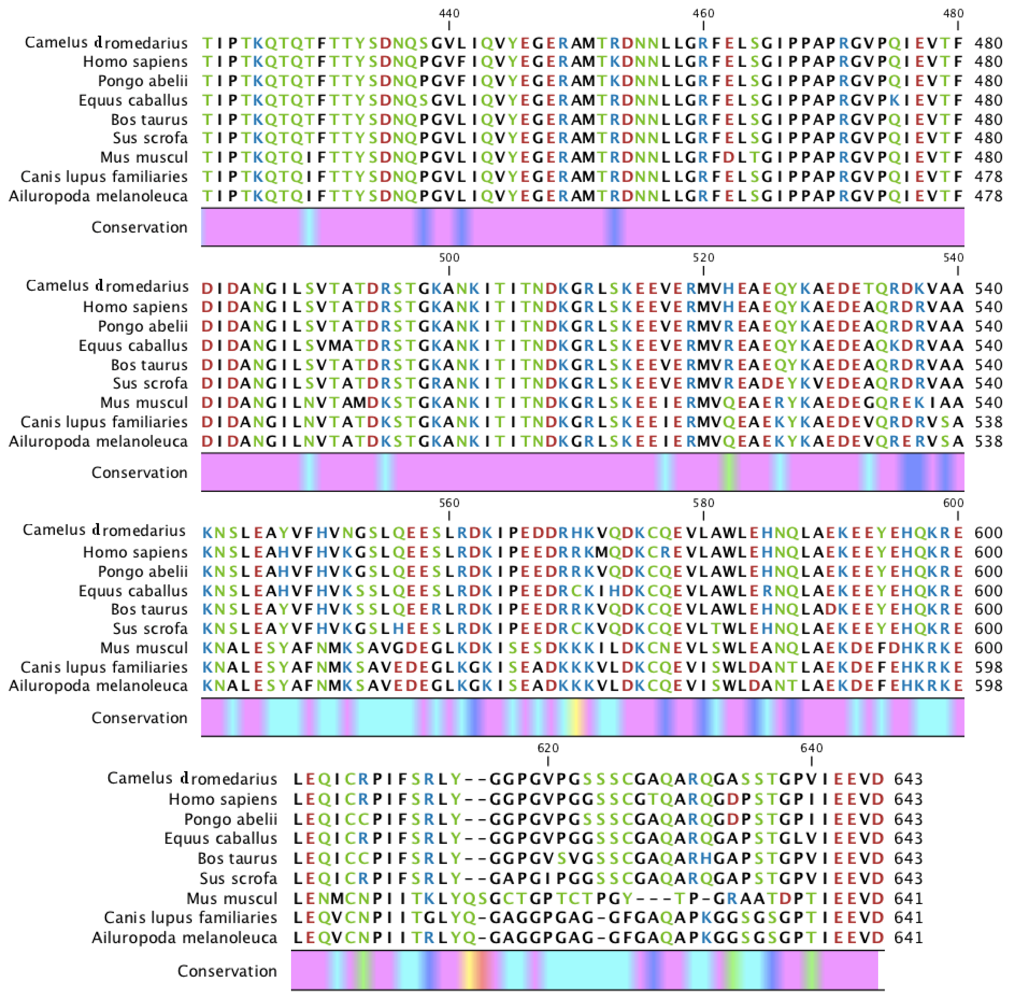
2.5. Camel HSPA6 3D Crystal Structures Alignment with Other Mammalian Species
3. Discussion
4. Experimental Section
4.1. Sample Collection
4.2. RNA Isolation and cDNA Synthesis
4.3. Synthesis and Confirmation of Partial cDNA of HSPA6 Gene
4.4. Rapid Amplification of cDNA Ends (RACE), Cloning and Sequencing
4.5. Analysis and Alignment of cDNA Sequence
4.6. Protein and mRNA Secondary Structure Prediction
4.7. Phylogenetic Analysis
4.8. Protein Structure and Sequence Analysis
4.9. Camel HSPA6 Protein 3D Structure Prediction
4.10. Camel HSPA6 3D Crystal Structures Alignment with Other Mammalian Species
5. Conclusions
Acknowledgments
References
- Pruski, AM; Dixon, DR. Heat shock protein expression pattern (HSP70) in the hydrothermal vent mussel Bathymodiolus azoricus. Mar. Environ. Res 2007, 64, 209–224. [Google Scholar]
- Lilja, K; Prevodnik, A; Gardeström, J; Elfwing, T; Tedengren, M; Bollner, T. Regional differences in mRNA responses in blue mussels within the Baltic proper. Comp. Biochem. Physiol 2008, 148, 101–106. [Google Scholar]
- Monari, M; Foschi, J; Rosmini, R; Marin, MG; Serrazanetti, GP. Heat shock protein 70 response to physical and chemical stress in Chamelea gallina. J. Exp. Mar. Biol. Ecol 2011, 397, 71–78. [Google Scholar]
- Hageman, J; van Waarde, MAWH; Zylicz, A; Walerych, D; Kampinga, HH. The diverse members of the mammalian HSP70 machine show distinct chaperone-like activities. Biochem. J 2011, 435, 127–142. [Google Scholar]
- Hammerer-Lercher, A; Mair, J; Bonatti, J; Watzka, SB; Puschendorf, B; Dirnhofer, S. Hypoxia induces heat shock protein expression in human coronary artery bypass grafts. Cardiovasc. Res 2001, 50, 115–24. [Google Scholar]
- Hochachka, PW; Somero, GN. Biochemical Adaptation: Mechanism and Process in Physiological Evolution, 1st ed; Oxford University Press: Oxford, MS, USA, 2002; 290. [Google Scholar]
- Ackerman, PA; Forsyth, RB; Mazur, CF; Iwama, GK. Stress hormones and the cellular stress response in salmonids. Fish Physiol Biochem 2000, 23, 327–336. [Google Scholar]
- Bierkens, JGEA. Applications and pitfalls of stress proteins in biomonitoring. Toxicology 2000, 153, 61–72. [Google Scholar]
- Heldens, L; Dirks, RP; Hensen, SMM; Onnekink, C; van Genesen, ST; Rustenburg, F; Lubsen, NH. Co-chaperones are limiting in a depleted chaperone network. Cell. Mol. Life Sci 2010, 67, 4035–4048. [Google Scholar]
- Leung, TKC; Rajendran, MY; Monfries, C; Hall, C; Lim, L. The human heat-shock protein family: Expression of a novel heat-inducible HSP70 (HSP7OB′) and isolation of its cDNA and genomic DNA. Biochem. J 1990, 267, 125–132. [Google Scholar]
- Wada, K; Taniguchi, A; Xu, L; Okano, T. Rapid and highly sensitive detection of cadmium chloride induced cytotoxicity using the HSP70B′ promoter in live cells. Biotechnol. Bioeng 2005, 92, 410–415. [Google Scholar]
- Wang, XY; Li, Y; Yang, G; Subjeck, JR. Current ideas about applications of heat shock proteins in vaccine design and immunotherapy. Int. J. Hyperthermia 2005, 8, 717–722. [Google Scholar]
- Xu, Q; Schett, G; Li, C; Hu, Y; Wick, G. Mechanical stress-induced heat shock protein 70 expression in vascular smooth muscle cells is regulated by Rac and Ras small G proteins but not mitogen-activated protein kinases. Circ. Res 2000, 86, 1122–1128. [Google Scholar]
- Romano, CC; Benedetto, N; Catania, MR; Rizzo, A; Gallè, F; Losi, E; Hasty, DL; Rossano, F. Commonly used antibiotics induce expression of Hsp 27 and Hsp 60 and protect human lymphocytes from apoptosis. Int. Immunopharmacol 2004, 8, 1067–1073. [Google Scholar]
- Noonan, EJ; Place, RF; Giardina, C; Hightower, LE. Hsp70B′ regulation and function. Cell Stress Chaperones 2007, 4, 393–402. [Google Scholar]
- Parsian, AJ; Sheren, JE; Tao, TY; Goswami, PC; Malyapa, R; Van Rheeden, R; Watson, MS; Hunt, CR. The human Hsp70B gene at the HSPA7 locus of chromosome 1 is transcribed but non-functional. Biochim. Biophys. Acta 2000, 1494, 201–205. [Google Scholar]
- Fowler, ME. Medicine and Surgery of Camelids, 3rd ed; Wiley-Blackwell: Ames, IA, USA, 2010; pp. 4–15. [Google Scholar]
- Khan, BB; Iqbal, A; Riaz, M. Production and Management of Camels, 1st ed; Pakistan T.M. Printers: Faisalabad, Pakistan, 2003. [Google Scholar]
- Elkhawad, AO. Selective brain cooling in desert animals: The Camel (Camelus dromedarius). Comp. Biochem. Physiol 1992, 101, 195–202. [Google Scholar]
- Thayyullathil, F; Chathoth, S; Hago, A; Wernery, U; Patel, M; Galadari, S. Investigation of heat stress response in the camel fibroblast cell line dubca. Ann. N.Y. Acad. Sci 2008, 1138, 376–384. [Google Scholar]
- Kregel, KC. Heat shock proteins: modifying factors in physiological stress responses and acquired thermotolerance. J. Appl. Physiol 2002, 92, 2177–2186. [Google Scholar]
- Leung, TKC; Hall, C; Rajendran, M; Spurr, NK; Lim, L. The human heat-shock genes Hspa6 and Hspa7 are both expressed and localize to chromosome-1. Genomics 1992, 12, 74–79. [Google Scholar]
- Gade, N; Mahapatra, RK; Sonawane, A; Singh, VK; Doreswamy, R; Saini, M. Molecular Characterization of Heat Shock Protein 70-1 Gene of Goat (Capra hircus). Mol Biol Int 2010. [Google Scholar] [CrossRef]
- Phelham, HRB. A regulatory upstream promoter element in the Drosophila Hsp 70 heat-shock gene. Cell 1982, 30, 517–528. [Google Scholar]
- Gutierrez, JA; Guerriero, V, Jr. Chemical modicfications of a recombinant bovine stress-inducible 70 kDa heat-shock protein (Hsp 70) mimics Hsp70 isoforms from tissues. Biochem. J 1995, 305, 197–203. [Google Scholar]
- Kano, R; Abe, K; Hasegawa, A. cDNA of canine heat shock protein 70 (HSP70). Vet. Res. Commun 2004, 28, 395–405. [Google Scholar]
- Morimoto, RI; Hunt, C; Huang, SY; Berg, L; Banerji, S. Organization, nucleotide sequence and transcription of the chicken HSP70 gene. J. Biol. Chem 1986, 261, 12692–12699. [Google Scholar]
- Rensing, SA; Maier, UG. Phylogenetic analysis of the stress-70 protein family. J. Mol. Evol 1994, 39, 80–86. [Google Scholar]
- Allen, RL; O’Brien, DA; Jones, CC; Rockett, DL; Eddy, EM. Expression of heat shock proteins by isolated mouse spermatogenic cells. Mol. Cell. Biol 1988, 8, 3260–3266. [Google Scholar]
- White, CN; Hightower, LE; Schultz, RJ. Variation in heat-shock proteins among species of desert fishes (Poeciliidae, poecilliopsis). Mol. Biol. Evol. J 1994, 1, 106–119. [Google Scholar]
- Norris, CE; di Iorio, PJ; Schultz, RJ; Hightower, LE. Variation in heat shock proteins within tropical and desert species of Poeciliid fishes. Mol. Biol. Evol 1995, 6, 1048–1062. [Google Scholar]
- Tavaria, M; Gabriele, T; Kola, I; Anderson, RL. A hitchhiker’s guide to the human Hsp70 family. Cell Stress Chaperones 1996, 1, 23–28. [Google Scholar]
- Manzerra, P; Rush, SJ; Brown, IR. Tissue-specific differences in heat shock protein HSP70 and HSP70 in the control and hyperthermic rabbit. J. Cell. Physiol 1997, 170, 130–137. [Google Scholar]
- Place, SP; Hofmann, GE. Comparison of Hsc70 orthologs from polar and temperate notothenioid fishes: differences in prevention of aggregation and refolding of denatured proteins. Am. J. Physiol. Regul. Integr. Comp. Physiol 2005, 288, R1195–R11202. [Google Scholar]
- Yamashita, M; Hirayoshi, K; Nagata, K. Characterization of multiple members of the HSP70 family in platyfish culture cells: Molecular evolution of stress protein HSP70 in vertebrates. Gene 2004, 336, 207–218. [Google Scholar]
- Hightower, LE; Norris, CE; di Iorio, PJ; Fielding, E. Heat shock responses of closely related species of tropical and desert fish. Integr. Comp. Biol 1999, 39, 877–888. [Google Scholar]
- Mahmood, T; Safdar, W; Abbasi, BH; Naqvi, SM. Overview on the small heat shock proteins. African J. Biotechnol 2010, 9, 927–949. [Google Scholar]
- Noonan, E; Giardinaa, C; Hightower, L. Hsp70B′ and Hsp72 form a complex in stressed human colon cells and each contributes to cytoprotection. Exp. Cell Res 2008, 314, 2468–2476. [Google Scholar]
- Chenchik, A; Moqadam, F; Siebert, P. A New Method for Full-Length cDNA Cloning by PCR. In A Laboratory Guide to RNA: Isolation, Analysis, and Synthesis, 1st ed; Krieg, PA, Ed.; Wiley-Liss: New York, NY, USA, 1996; pp. 273–321. [Google Scholar]
- Expasy Translate Tool. Available online: http://www.expasy.ch/tools/dna.html accessed on 24 June 2011.
- Bioedit. Available online: http://www.mbio.ncsu.edu/bioedit/bioedit.html accessed on 24 June 2011.
- NCBI BLAST program. Available online: http://www.ncbi.nlm.nih.gov/blast accessed on 24 June 2011.
- CLC Genome workbench version 6.0.1. Available online: http://www.clcbio.com accessed on 24 June 2011.
- Protein Calculator v3.3. Available online: http://www.scripps.edu/~cdputnam/protcalc.html accessed on 24 June 2011.
- Guindon, S; Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol 2003, 52, 696–704. [Google Scholar]
- Dereeper, A; Audic, S; Claverie, JM; Blanc, G. BLAST-EXPLORER helps you building datasets for phylogenetic analysis. BMC Evol. Biol 2010, 10, 8. [Google Scholar]
- Dereeper, A; Guignon, V; Blanc, G; Audic, S; Buffet, S; Chevenet, F; Dufayard, JF; Guindon, S; Lefort, V; Lescot, M; et al. Phylogeny.fr: Robust phylogenetic analysis for the nonspecialist. Nucleic Acids Res 2008, 36, W465–W469. [Google Scholar]
- Anisimova, M; Gascuel, O. Approximate likelihood ratio test for branches: A fast, accurate and powerful alternative. Syst. Biol 2006, 55, 539–552. [Google Scholar]
- Eisenberg, D; Schwarz, E; Komaromy, M; Wall, R. Analysis of membrane and surface protein sequences with the hydrophobic moment plot. J. Mol. Biol 1984, 179, 125–142. [Google Scholar]
- Eswar, N; Eramian, D; Webb, B; Shen, M; Sali, A. Protein Structure Modeling With MODELLER. Methods Mol. Biol 2006, 426, 145–159. [Google Scholar]
- Marti-Renom, MA; Madhusudhan, MS; Sali, A. Alignment of protein sequences by their profiles. Protein Sci 2004, 13, 1071–1087. [Google Scholar]
- Edgar, RC. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 2004, 32, 1792–1797. [Google Scholar]
- Roy, A; Kucukural, A; Zhang, Y. I-TASSER: A unified platform for automated protein structure and function prediction. Nat. Protoc 2010, 5, 725–738. [Google Scholar]
- Zhang, Y. I-TASSER server for protein 3D structure prediction. BMC Bioinfma 2008, 9, 40. [Google Scholar]
- Benkert, P; Künzli, M; Schwede, T. QMEAN server for protein model quality estimation. Nucleic Acids Res 2009, 37, W510–W514. [Google Scholar]
- Benkert, P; Biasini, M; Schwede, T. Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics 2011, 27, 343–350. [Google Scholar]
- Holm, L; Park, J. DaliLite workbench for protein structure comparison. Bioinformatics 2000, 16, 566–567. [Google Scholar]






| Name | GeneBank Accession Number | cDNA Length (bp) | UTR | Total Score | Coverage % | Identity % | |
|---|---|---|---|---|---|---|---|
| 5′ bp | 3′ bp | ||||||
| Arabian Camel (Camelus dromedarius) | HQ214118 | 2417 | 136 | 349 | 2529 | 100 | 100 |
| Human (Homo sapiens) | NM_002155 | 2664 | 413 | 319 | 2523 | 81 | 89 |
| Sumatran orangutan (Pongo abelii) | XM_002809884 | 2398 | 133 | 334 | 2529 | 81 | 89 |
| Horse (Equus caballus) | XM_001488139 | 2124 | 130 | 62 | 2682 | 86 | 89 |
| Cow (Bos Taurus) | XM_589747 | 2210 | 193 | 85 | 2915 | 88 | 91 |
| Pig (Sus scrofa) | NM_001123127 | 2518 | 105 | 481 | 3025 | 93 | 91 |
| Mouse (Mus musculus) | M32218 | 2298 | 173 | 371 | 1518 | 75 | 77 |
| Dog (Canis lupus familiaris) | NM_001003067 | 2026 | 86 | 284 | 1906 | 79 | 80 |
| Giant panda (Ailuropoda melanoleuca) | XM_002931069 | 2356 | 121 | 309 | 1480 | 79 | 81 |
| Name | Protein Accession Number | Amino Acid Residues | Total Score | Coverage % | Identity % | pI | Molecular Weight | Estimate Charge at pH |
|---|---|---|---|---|---|---|---|---|
| Arabian Camel (Camelus dromedarius) | ADO12067 | 643 | 1252 | 100 | 100 | 6.00 | 70.5 | −9.0 |
| Human (Homo sapiens) | NP_002146 | 643 | 1227 | 100 | 94 | 6.13 | 71.0 | −7.3 |
| Sumatran orangutan (Pongo abelii) | XP_002809930 | 643 | 1252 | 100 | 94 | 5.86 | 70.9 | −9.5 |
| Horse (Equus caballus) | XP_001488189 | 643 | 1217 | 100 | 93 | 6.20 | 70.8 | −7.0 |
| Cow (Bos Taurus) | XP_589747 | 643 | 1244 | 100 | 94 | 6.04 | 71.0 | −8.3 |
| Pig (Sus scrofa) | NP_001116599.1 | 643 | 1251 | 100 | 94 | 6.06 | 71.1 | −7.5 |
| Mouse (Mus musculus) | AAA74906 | 641 | 1077 | 100 | 80 | 6.12 | 70.7 | −6.0 |
| Dog (Canis lupus familiaris) | BAC79353 | 641 | 1092 | 99 | 83 | 6.30 | 70.5 | −4.2 |
| Giant panda (Ailuropoda melanoleuca) | XP_002931115 | 641 | 1093 | 99 | 83 | 5.73 | 70.2 | −8.4 |
| Name | Beta Interaction Energy (Z-score) | Pairwise Energy (Z-score) | Solvent Energy (Z-score) | Torsion Angle Energy (Z-score) | Secondary Structure Alignment (Z-score) | Solvent Accessibility Agreement (Z-score) | Total QMEANScore; Estimated Model Reliability between 0–1) (Z-score) |
|---|---|---|---|---|---|---|---|
| Camelus dromedarius | −263.13 (0.64) | −18120.78 (0.75) | −70.23 (0.63) | −61.42 (−2.84) | 77.1% (−0.33) | 76.7% (−0.81) | 0.601 (1.79 |
| Bos Taurus | −335.84 (1.25) | −21082.24 (1.32) | −87.35 (1.46) | −121.57 (−1.41) | 78.1% (−0.13) | 74.3% (−1.25) | 0.606 (−1.74) |
| Homo sapiens | −206.76 (1.05) | −13292.65 (1.21) | −42.48 (0.51) | −119.06 (0.65) | 83.4% (0.47) | 90.5% (1.87) | 0.939 (1.96) |
| Mus musculus | −594.81 (2.02) | −36711.18 (2.12) | −109.35 (0.58) | −262.22 (0.21) | 85.1% (0.65) | 84.5% (0.77) | 0.847 (1.02) |
| Canis lupus | −720.31 (1.67) | −46429.75 (1.87) | −155.98 (1.06) | −267.45 (−0.59) | 81.6% (−0.01) | 74.8% (−0.92) | 0.658 (−0.99) |
| No. | Name | Chain | Z-Score | Aligned Residues | RMSD Å | Sequence Identity% |
|---|---|---|---|---|---|---|
| 1 | Camel + Homo sapiens | A | 49.9 | 380 | 2.6 | 87 |
| 2 | Camel + Mus musculus | A | 47.3 | 376 | 3.0 | 82 |
| 3 | Camel + Canis lupus | A | 1.7 | 52 | 3.3 | 8 |
| 4 | Camel + Bos taurus | A | 61.2 | 606 | 2.2 | 27 |
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Elrobh, M.S.; Alanazi, M.S.; Khan, W.; Abduljaleel, Z.; Al-Amri, A.; Bazzi, M.D. Molecular Cloning and Characterization of cDNA Encoding a Putative Stress-Induced Heat-Shock Protein from Camelus dromedarius. Int. J. Mol. Sci. 2011, 12, 4214-4236. https://doi.org/10.3390/ijms12074214
Elrobh MS, Alanazi MS, Khan W, Abduljaleel Z, Al-Amri A, Bazzi MD. Molecular Cloning and Characterization of cDNA Encoding a Putative Stress-Induced Heat-Shock Protein from Camelus dromedarius. International Journal of Molecular Sciences. 2011; 12(7):4214-4236. https://doi.org/10.3390/ijms12074214
Chicago/Turabian StyleElrobh, Mohamed S., Mohammad S. Alanazi, Wajahatullah Khan, Zainularifeen Abduljaleel, Abdullah Al-Amri, and Mohammad D. Bazzi. 2011. "Molecular Cloning and Characterization of cDNA Encoding a Putative Stress-Induced Heat-Shock Protein from Camelus dromedarius" International Journal of Molecular Sciences 12, no. 7: 4214-4236. https://doi.org/10.3390/ijms12074214




