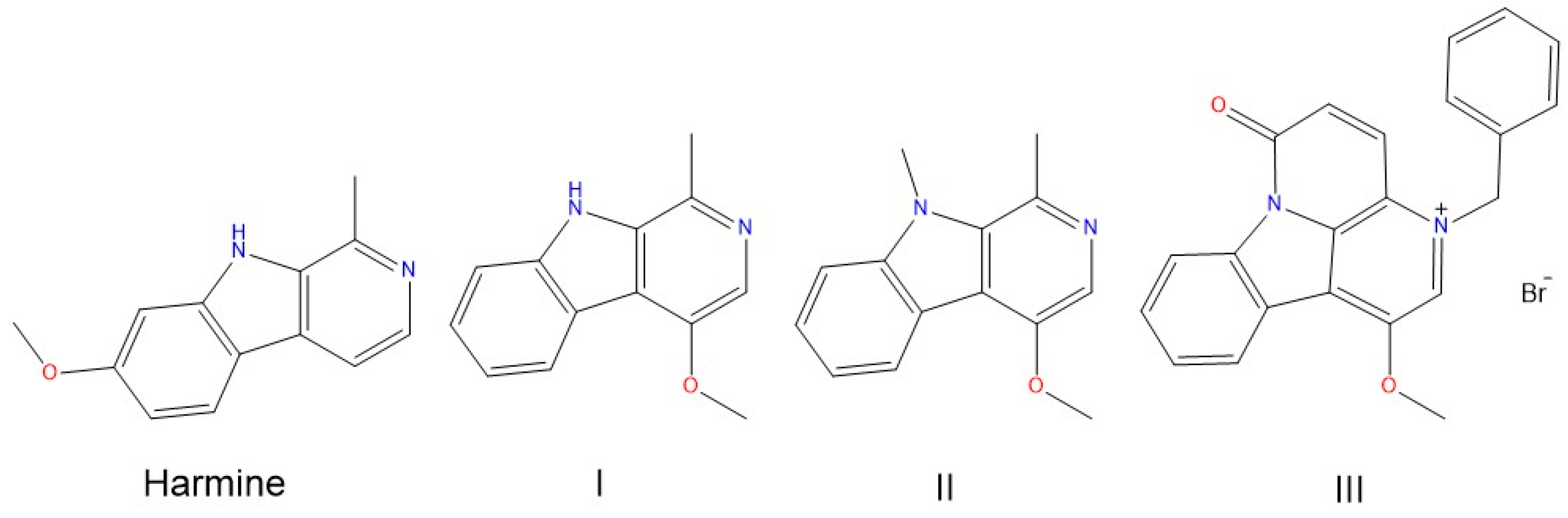
Binding of Harmine Derivatives to DNA: A Spectroscopic Investigation
Abstract
:1. Introduction
2. Results
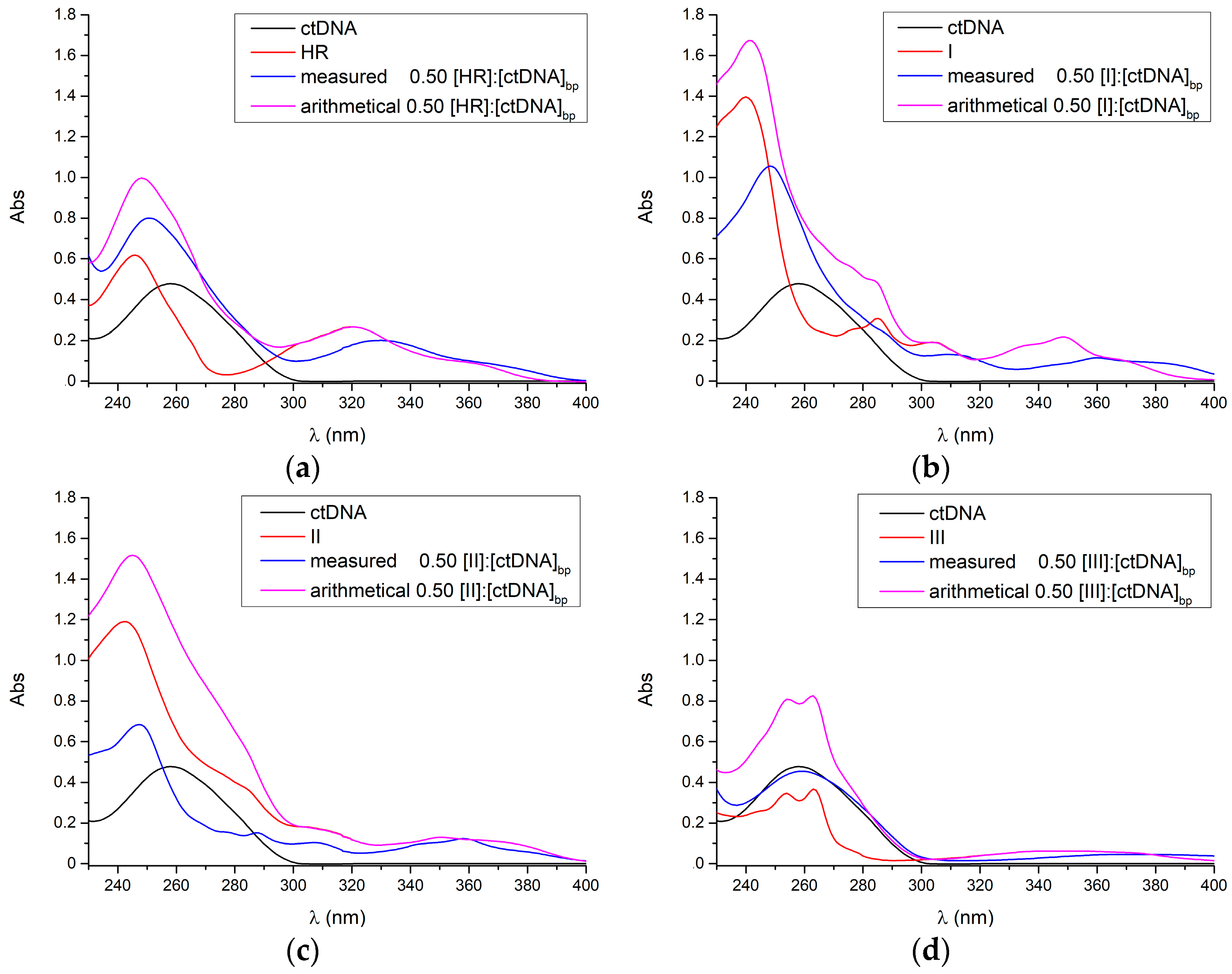
2.1. UV-Vis
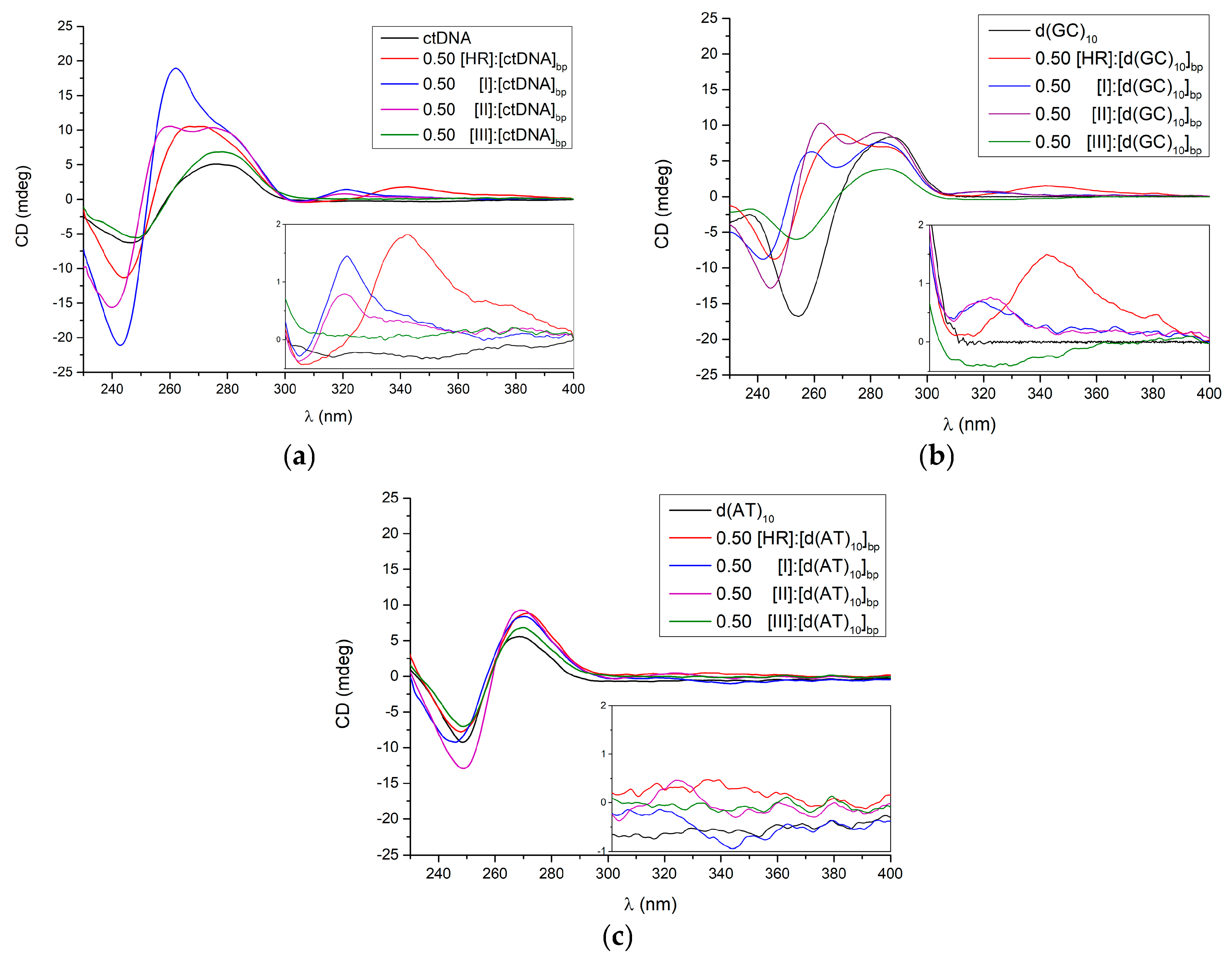
2.2. Circular Dichroism
3. Discussion
4. Materials and Methods
4.1. Chemicals
4.2. UV Measurements
4.3. CD Measurements
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Siddiqui, S.; Khan, O.Y.; Faizi, S.; Siddiqui, B.S. Studies on the chemical-constituents of the seeds of Peganum harmala. Isolation and structure of a new β-carboline alkaloid. Harmalicine. Heterocycles 1987, 26, 1563–1567. [Google Scholar] [CrossRef]
- Siddiqui, S.; Khan, O.Y.; Faizi, S.; Siddiqui, B.S. Studies on the chemical-constituents of the seeds of Peganum-harmala: Isolation and structure elucidation of two β-carbolines—harmalacinine and norharmine. Heterocycles 1989, 29, 521–527. [Google Scholar] [CrossRef]
- Moloudizargari, M.; Mikaili, P.; Aghajanshakeri, S.; Asghari, M.H.; Shayegh, J. Pharmacological and therapeutic effects of Peganum harmala and its main alkaloids. Pharmacogn. Rev. 2013, 7, 199–212. [Google Scholar] [CrossRef] [PubMed]
- Zaker, F.; Oody, A.; Arjmand, A. A study on the antitumoral and differentiation effects of Peganum harmala derivatives in combination with ATRA on leukaemic cells. Arch. Pharm. Res. 2007, 30, 844–849. [Google Scholar] [CrossRef] [PubMed]
- Egusa, H.; Doi, M.; Saeki, M.; Fukuyasu, S.; Akashi, Y.; Yokota, Y.; Yatani, H.; Kamisaki, Y. The small molecule harmine regulates nfatc1 and id2 expression in osteoclast progenitor cells. Bone 2011, 49, 264–274. [Google Scholar] [CrossRef] [PubMed]
- Frost, D.; Meechoovet, B.; Wang, T.; Gately, S.; Giorgetti, M.; Shcherbakova, I.; Dunckley, T. β-carboline compounds, including harmine, inhibit DYRK1A and tau phosphorylation at multiple Alzheimer’s disease-related sites. PLoS ONE 2011, 6, e19264. [Google Scholar] [CrossRef] [PubMed]
- Hamsa, T.P.; Kuttan, G. Harmine activates intrinsic and extrinsic pathways of apoptosis in b16f-10 melanoma. Chin. Med. 2011, 6, 11. [Google Scholar] [CrossRef] [PubMed]
- Yonezawa, T.; Hasegawa, S.; Asai, M.; Ninomiya, T.; Sasaki, T.; Cha, B.Y.; Teruya, T.; Ozawa, H.; Yagasaki, K.; Nagai, K.; et al. Harmine, a β-carboline alkaloid, inhibits osteoclast differentiation and bone resorption in vitro and in vivo. Eur. J. Pharmacol. 2011, 650, 511–518. [Google Scholar] [CrossRef] [PubMed]
- Onishi, Y.; Oishi, K.; Kawano, Y.; Yamazaki, Y. The harmala alkaloid harmine is a modulator of circadian Bmal1 transcription. Biosci. Rep. 2012, 32, 45–52. [Google Scholar] [CrossRef] [PubMed]
- Reus, G.Z.; Stringari, R.B.; Goncalves, C.L.; Scaini, G.; Carvalho-Silva, M.; Jeremias, G.C.; Jeremias, I.C.; Ferreira, G.K.; Streck, E.L.; Hallak, J.E.; et al. Administration of harmine and imipramine alters creatine kinase and mitochondrial respiratory chain activities in the rat brain. Depress. Res. Treat. 2012, 2012, 987397. [Google Scholar] [CrossRef] [PubMed]
- Yi, J.M.; Zhang, X.F.; Huan, X.J.; Song, S.S.; Wang, W.; Tian, Q.T.; Sun, Y.M.; Chen, Y.; Ding, J.; Wang, Y.Q.; et al. Dual targeting of microtubule and topoisomerase II by α-carboline derivative YCH337 for tumor proliferation and growth inhibition. Oncotarget 2015, 6, 8960–8973. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Li, M.; Tan, S.; Wang, C.; Fan, S.; Huang, C. Harmine is an inflammatory inhibitor through the suppression of NF-κB signaling. Biochem. Biophys. Res. Commun. 2017, 489, 332–338. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Chao, R.; Chen, H.; Hou, X.; Yan, H.; Zhou, S.; Peng, W.; Xu, A. Antitumor and neurotoxic effects of novel harmine derivatives and structure-activity relationship analysis. Int. J. Cancer 2005, 114, 675–682. [Google Scholar] [CrossRef] [PubMed]
- Peduto, A.; More, V.; de Caprariis, P.; Festa, M.; Capasso, A.; Piacente, S.; De Martino, L.; De Feo, V.; Filosa, R. Synthesis and cytotoxic activity of new β-carboline derivatives. Mini Rev. Med. Chem. 2011, 11, 486–491. [Google Scholar] [CrossRef] [PubMed]
- Funayama, Y.; Nishio, K.; Wakabayashi, K.; Nagao, M.; Shimoi, K.; Ohira, T.; Hasegawa, S.; Saijo, N. Effects of β- and γ-carboline derivatives on DNA topoisomerase activities. Mutat. Res-Fund. Mol. Mech. Mutagen. 1996, 349, 183–191. [Google Scholar] [CrossRef]
- Wink, M.; Schimmer, O. Molecular modes of action of defensive secondary metabolites. In Annual Plant Reviews Volume 39: Functions and Biotechnology of Plant Secondary Metabolites; Wiley-Blackwell: Oxford, UK, 2010; pp. 21–161. [Google Scholar]
- Zhao, L.; Wink, M. The β-carboline alkaloid harmine inhibits telomerase activity of MCF-7 cells by down-regulating hTERT mRNA expression accompanied by an accelerated senescent phenotype. Peerj 2013, 1, e174. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cao, R.; Peng, W.; Chen, H.; Ma, Y.; Liu, X.; Hou, X.; Guan, H.; Xu, A. DNA binding properties of 9-substituted harmine derivatives. Biochem. Biophys. Res. Commun. 2005, 338, 1557–1563. [Google Scholar] [CrossRef] [PubMed]
- Cao, R.; Peng, W.; Chen, H.; Hou, X.; Guan, H.; Chen, Q.; Ma, Y.; Xu, A. Synthesis and in vitro cytotoxic evaluation of 1,3-bisubstituted and 1,3,9-trisubstituted β-carboline derivatives. Eur. J. Med. Chem. 2005, 40, 249–257. [Google Scholar] [CrossRef] [PubMed]
- Airoldi, M.; Barone, G.; Gennaro, G.; Giuliani, A.M.; Giustini, M. Interaction of doxorubicin with polynucleotides. A spectroscopic study. Biochemistry 2014, 53, 2197–2207. [Google Scholar] [CrossRef] [PubMed]
- Bhadra, K.; Maiti, M.; Kumar, G.S. Molecular recognition of DNA by small molecules: At base pair specific intercalative binding of cytotoxic plant alkaloid palmatine. Biochim. Biophys. Acta 2007, 1770, 1071–1080. [Google Scholar] [CrossRef] [PubMed]
- Mergny, J.L.; Lacroix, L. Analysis of thermal melting curves. Oligonucleotides 2003, 13, 515–537. [Google Scholar] [CrossRef] [PubMed]
- Lyng, R.; Hard, T.; Norden, B. Induced CD of DNA intercalators: Electric dipole allowed transitions. Biopolymers 1987, 26, 1327–1345. [Google Scholar] [CrossRef] [PubMed]
- Monnot, M.; Mauffret, O.; Lescot, E.; Fermandjian, S. Probing intercalation and conformational effects of the anticancer drug 2-methyl-9-hydroxyellipticinium acetate in DNA fragments with circular-dichroism. Eur. J. Biochem. 1992, 204, 1035–1039. [Google Scholar] [CrossRef] [PubMed]
- Sharma, S.; Yadav, M.; Gupta, S.P.; Pandav, K.; Kumar, S. Spectroscopic and structural studies on the interaction of an anticancer β-carboline alkaloid, harmine with GC and AT specific DNA oligonucleotides. Chem. Biol. Interact. 2016, 260, 256–262. [Google Scholar] [CrossRef] [PubMed]
- Saha, I.; Hossain, M.; Kumar, G.S. Sequence-selective binding of phenazinium dyes phenosafranin and safranin O to guanine-cytosine deoxyribopolynucleotides: Spectroscopic and thermodynamic studies. J. Phys. Chem. B 2010, 114, 15278–15287. [Google Scholar] [CrossRef] [PubMed]
- Herrera, J.E.; Chaires, J.B. A premelting conformational transition in poly(dA)-poly(dT) coupled to daunomycin binding. Biochemistry 1989, 28, 1993–2000. [Google Scholar] [CrossRef] [PubMed]
- Schwaller, M.A.; Aubard, J.; Auclair, C.; Paoletti, C.; Dodin, G. The G. C base-pair preference of 2-N-methyl 9-hydroxyellipticinium. Eur. J. Biochem. 1989, 181, 129–134. [Google Scholar] [CrossRef] [PubMed]
- Shehi, E.; Granata, V.; Del Vecchio, P.; Barone, G.; Fusi, P.; Tortora, P.; Graziano, G. Thermal stability and DNA binding activity of a variant form of the Sso7d protein from the archeon Sulfolobus solfataricus truncated at leucine 54. Biochemistry 2003, 42, 8362–8368. [Google Scholar] [CrossRef] [PubMed]
- Cantor, C.R.; Warshaw, M.M.; Shapiro, H. Oligonucleotide interactions. III. Circular dichroism studies of the conformation of deoxyoligonucleotides. Biopolymers 1970, 9, 1059–1077. [Google Scholar] [CrossRef] [PubMed]
Sample Avalaibility: Samples of the compounds I–III are available from authors. |
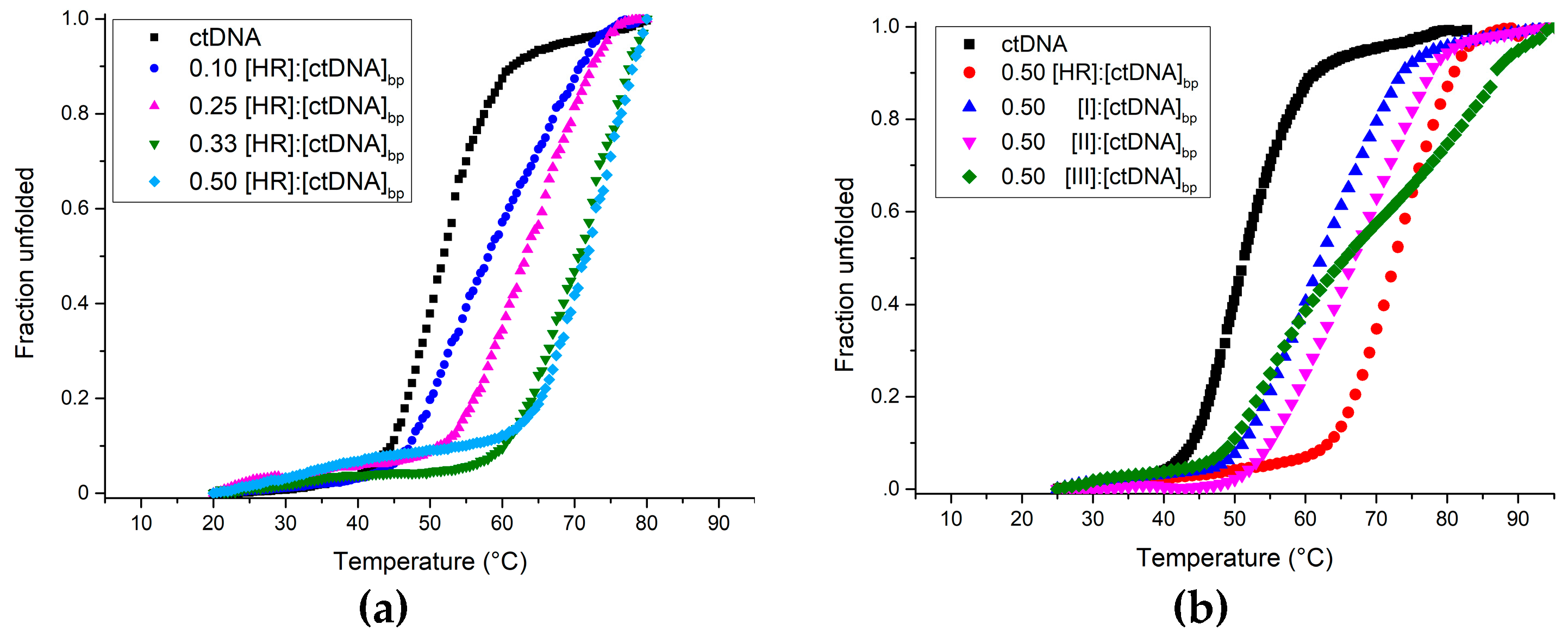
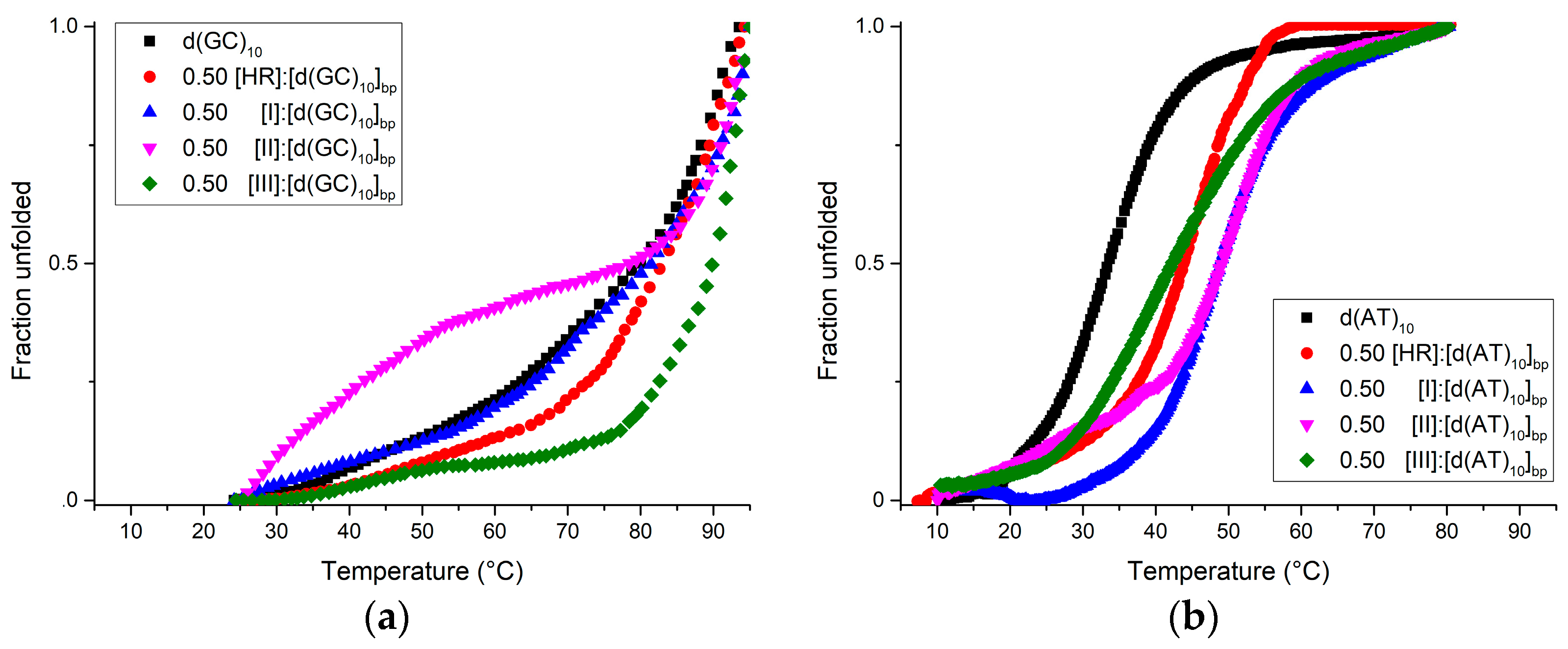
| Ligand | Tm (°C ± 1) | ||
|---|---|---|---|
| ctDNA | d(GC)10 | d(AT)10 | |
| No ligand | 55 | >85 | 35 |
| Harmine | 75 | >85 | 45 |
| I | 62 | >85 | 52 |
| II | 65 | >85 | 52 |
| III | ≈60–≈82 | >90 | 42 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pagano, B.; Caterino, M.; Filosa, R.; Giancola, C. Binding of Harmine Derivatives to DNA: A Spectroscopic Investigation. Molecules 2017, 22, 1831. https://doi.org/10.3390/molecules22111831
Pagano B, Caterino M, Filosa R, Giancola C. Binding of Harmine Derivatives to DNA: A Spectroscopic Investigation. Molecules. 2017; 22(11):1831. https://doi.org/10.3390/molecules22111831
Chicago/Turabian StylePagano, Bruno, Marco Caterino, Rosanna Filosa, and Concetta Giancola. 2017. "Binding of Harmine Derivatives to DNA: A Spectroscopic Investigation" Molecules 22, no. 11: 1831. https://doi.org/10.3390/molecules22111831






